External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G22630 362 / 2e-129
PRCGB, PBD1
20S proteasome beta subunit D1 (.1)
AT4G14800 357 / 2e-127
PBD2
20S proteasome beta subunit D2 (.1.2)
AT3G14290 66 / 6e-13
PAE2
20S proteasome alpha subunit E2 (.1)
AT1G53850 65 / 1e-12
PAE1, ATPAE1
ARABIDOPSIS 20S PROTEASOME ALPHA SUBUNIT E1, 20S proteasome alpha subunit E1 (.1.2)
AT1G13060 51 / 7e-08
PBE1
20S proteasome beta subunit E1 (.1.2)
AT3G26340 51 / 8e-08
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1)
AT3G60820 51 / 9e-08
PBF1
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
AT5G40580 48 / 1e-06
PBB2
20S proteasome beta subunit PBB2 (.1.2.3)
AT3G27430 48 / 1e-06
PBB1
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
AT1G56450 43 / 5e-05
PBG1
20S proteasome beta subunit G1 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10021909
414 / 4e-150
AT3G22630 360 / 2e-128
20S proteasome beta subunit D1 (.1)
Lus10039351
392 / 2e-141
AT3G22630 371 / 5e-133
20S proteasome beta subunit D1 (.1)
Lus10006596
184 / 1e-60
AT3G22630 168 / 2e-54
20S proteasome beta subunit D1 (.1)
Lus10020180
67 / 2e-13
AT4G31300 429 / 7e-155
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
Lus10026984
67 / 3e-13
AT4G31300 357 / 2e-124
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
Lus10003936
64 / 1e-11
AT1G53850 463 / 9e-162
ARABIDOPSIS 20S PROTEASOME ALPHA SUBUNIT E1, 20S proteasome alpha subunit E1 (.1.2)
Lus10037454
63 / 1e-11
AT1G53850 464 / 8e-163
ARABIDOPSIS 20S PROTEASOME ALPHA SUBUNIT E1, 20S proteasome alpha subunit E1 (.1.2)
Lus10006426
52 / 4e-08
AT3G26340 462 / 7e-167
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1)
Lus10011369
51 / 1e-07
AT3G26340 465 / 1e-167
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G084800
369 / 7e-132
AT3G22630 366 / 6e-131
20S proteasome beta subunit D1 (.1)
Potri.008G155500
368 / 8e-132
AT3G22630 364 / 6e-130
20S proteasome beta subunit D1 (.1)
Potri.006G077900
66 / 3e-13
AT4G31300 416 / 2e-149
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
Potri.018G145900
64 / 3e-12
AT4G31300 405 / 3e-145
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
Potri.001G162900
64 / 3e-12
AT3G14290 471 / 4e-171
20S proteasome alpha subunit E2 (.1)
Potri.003G072500
60 / 6e-11
AT3G14290 464 / 1e-168
20S proteasome alpha subunit E2 (.1)
Potri.014G069800
55 / 3e-09
AT3G60820 362 / 6e-129
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
Potri.008G177000
53 / 2e-08
AT3G26340 473 / 7e-171
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1)
Potri.002G148300
52 / 3e-08
AT3G60820 387 / 2e-138
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
Potri.010G058100
49 / 4e-07
AT3G26340 463 / 6e-167
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0052
NTN
PF00227
Proteasome
Proteasome subunit
Representative CDS sequence
>Lus10041194 pacid=23180022 polypeptide=Lus10041194 locus=Lus10041194.g ID=Lus10041194.BGIv1.0 annot-version=v1.0
ATGGAGTGCGTATTTGGGTTAGTAGGAAACGATTTCGCAATAGTTGCGGCAGATACGTCGGCCGTCCACAGCATTCTGGTCCACAAATCCAACGAGGACA
AGATCATGCTCCTCGACTCCCACAAGCTCGTCGCCGCCAGCGGCGAGCCCGGCGACAGAGTTCAATTCACGGAGTACATCCAGAAGAACGTGACTTTGTA
TCAGTTCAGGAATGGAATTCCTCTTACCACGCCTGCTGCAGCCAACTTTACTCGAGGGGAGCTGGCTACTGCTTTGAGAAAGAATCCTTATCAAGTTAAC
CTCTTGATGGCTGGTTATGACAAAGACATAGGCCCATCTTTGTATTACATCGACTACATTGCTACCCTTCACAAGGTTGACAAGGGTGCATTCGGCTATG
GCTCCTATTTCTGTCTTTCCATGATGGATCGACTGTACCACAGTGGCATGACTGTGGAGGAAGCTATTGATCTCGTAGACAAGTGCATAGCGGAGATCCG
GTTGAGGTTGGTAGTGGCACCGCCAAATTTCGTCATTAAAATCGTCGACAAGGATGGGGCAAGGGAGTACGCCTGGCGCAAGTCTGTCAAGGATGATGAA
GCAGCTTAA
AA sequence
>Lus10041194 pacid=23180022 polypeptide=Lus10041194 locus=Lus10041194.g ID=Lus10041194.BGIv1.0 annot-version=v1.0
MECVFGLVGNDFAIVAADTSAVHSILVHKSNEDKIMLLDSHKLVAASGEPGDRVQFTEYIQKNVTLYQFRNGIPLTTPAAANFTRGELATALRKNPYQVN
LLMAGYDKDIGPSLYYIDYIATLHKVDKGAFGYGSYFCLSMMDRLYHSGMTVEEAIDLVDKCIAEIRLRLVVAPPNFVIKIVDKDGAREYAWRKSVKDDE
AA
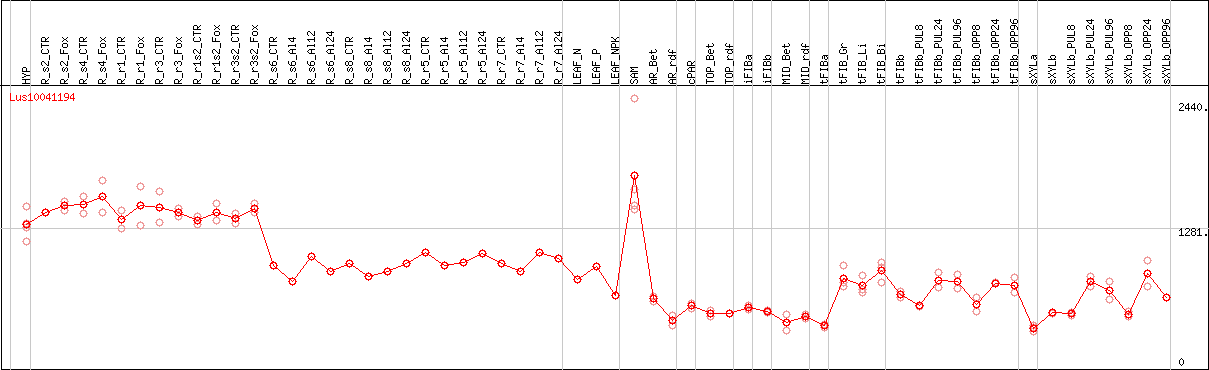
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10041194 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.