External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G22430 186 / 4e-57
unknown protein
AT5G23570 52 / 2e-08
SGS3, ATSGS3
SUPPRESSOR OF GENE SILENCING 3, XS domain-containing protein / XS zinc finger domain-containing protein-related (.1)
AT4G01180 42 / 8e-05
XH/XS domain-containing protein (.1)
AT5G59390 41 / 0.0002
XH/XS domain-containing protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10021917
237 / 2e-81
AT3G22430 125 / 2e-34
unknown protein
Lus10021918
78 / 2e-17
AT3G22430 298 / 6e-97
unknown protein
Lus10020567
72 / 2e-15
AT3G22430 129 / 8e-33
unknown protein
Lus10004302
72 / 4e-15
AT5G23570 517 / 4e-180
SUPPRESSOR OF GENE SILENCING 3, XS domain-containing protein / XS zinc finger domain-containing protein-related (.1)
Lus10006278
64 / 2e-12
AT3G22430 132 / 2e-32
unknown protein
Lus10019214
56 / 2e-09
AT5G23570 515 / 2e-179
SUPPRESSOR OF GENE SILENCING 3, XS domain-containing protein / XS zinc finger domain-containing protein-related (.1)
Lus10011024
52 / 4e-08
AT5G41150 1233 / 0.0
ULTRAVIOLET HYPERSENSITIVE 1, Restriction endonuclease, type II-like superfamily protein (.1.2)
Lus10000949
47 / 1e-06
AT3G48670 319 / 2e-100
RNA-DIRECTED DNA METHYLATION 12, INVOLVED IN DE NOVO 2, XH/XS domain-containing protein (.1.2)
Lus10002703
46 / 4e-06
AT3G48670 495 / 3e-166
RNA-DIRECTED DNA METHYLATION 12, INVOLVED IN DE NOVO 2, XH/XS domain-containing protein (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.008G152900
211 / 7e-67
AT3G22430 409 / 2e-138
unknown protein
Potri.010G087400
209 / 3e-66
AT3G22430 418 / 3e-142
unknown protein
Potri.010G235700
79 / 1e-17
AT3G22430 125 / 4e-30
unknown protein
Potri.009G147201
71 / 8e-15
AT3G22430 148 / 3e-37
unknown protein
Potri.T045700
70 / 2e-14
AT5G23570 604 / 0.0
SUPPRESSOR OF GENE SILENCING 3, XS domain-containing protein / XS zinc finger domain-containing protein-related (.1)
Potri.018G137400
70 / 2e-14
AT5G23570 603 / 0.0
SUPPRESSOR OF GENE SILENCING 3, XS domain-containing protein / XS zinc finger domain-containing protein-related (.1)
Potri.003G021000
68 / 9e-14
AT5G23570 439 / 3e-147
SUPPRESSOR OF GENE SILENCING 3, XS domain-containing protein / XS zinc finger domain-containing protein-related (.1)
Potri.008G024500
66 / 3e-13
AT3G22430 127 / 1e-30
unknown protein
Potri.003G187800
57 / 7e-10
AT5G23570 226 / 4e-67
SUPPRESSOR OF GENE SILENCING 3, XS domain-containing protein / XS zinc finger domain-containing protein-related (.1)
Potri.003G188350
56 / 2e-09
AT5G23570 269 / 7e-78
SUPPRESSOR OF GENE SILENCING 3, XS domain-containing protein / XS zinc finger domain-containing protein-related (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0221
RRM
PF03468
XS
XS domain
Representative CDS sequence
>Lus10041203 pacid=23179586 polypeptide=Lus10041203 locus=Lus10041203.g ID=Lus10041203.BGIv1.0 annot-version=v1.0
ATGATCTGGCCACCTGTGGTGATTATTCATAACACCGCTACAGGAAAGAGTAAGGAGGGACGCATAGAAGGTTTGGGAAACAAAGCTATGGATAGCAAAA
TAAGAGCTCTTGGATTCCCAGGAGGCAAGTCGAAATCACTGTATGGTAGAGAAGGGCATCAAGGAACAACTCATGTAAAGTTTGCGGGCGACACGCCTGG
CCTAAGAGAAGCTGTGGGGCTGTCAGATCATTTTGCGAAACAAAATCGCGGACGAAATGCATGGGAACGTGTACAAGCAGCAACGGGAGGGAAAAGCGAC
GATAGGAATCCAGACCTTGTGAAGTTGGATCGCAGCGGGGATAAAACGAAGGTTCTCTACGGTTACCTTGCCACAGCTGCTGATCTTCCCAGGTTTGATT
TCGATACGAAGAAGAAGCTCACACTTGAGAGTGTGCGAGAGTATGAAGGATTCCAAGTTGGTCGTCCTCCCAGATAA
AA sequence
>Lus10041203 pacid=23179586 polypeptide=Lus10041203 locus=Lus10041203.g ID=Lus10041203.BGIv1.0 annot-version=v1.0
MIWPPVVIIHNTATGKSKEGRIEGLGNKAMDSKIRALGFPGGKSKSLYGREGHQGTTHVKFAGDTPGLREAVGLSDHFAKQNRGRNAWERVQAATGGKSD
DRNPDLVKLDRSGDKTKVLYGYLATAADLPRFDFDTKKKLTLESVREYEGFQVGRPPR
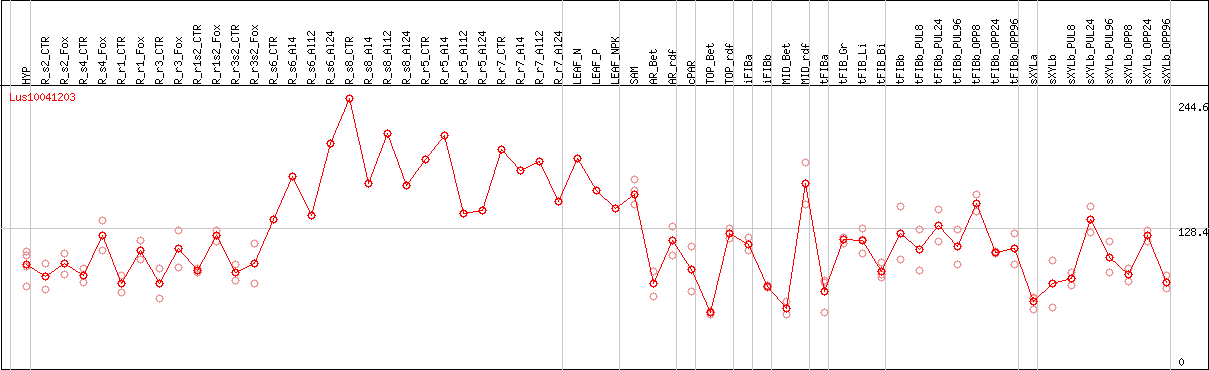
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10041203 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.