External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G40390 71 / 1e-14
RS5, SIP1
seed imbibition 1-like, raffinose synthase 5, Raffinose synthase family protein (.1)
AT3G57520 52 / 3e-08
RS2, ATSIP2
raffinose synthase 2, seed imbibition 2 (.1.2.3)
AT5G20250 49 / 3e-07
RS6, DIN10
raffinose synthase 6, DARK INDUCIBLE 10, Raffinose synthase family protein (.1.2.3.4)
AT1G55740 48 / 5e-07
RS1, ATSIP1
raffinose synthase 1, seed imbibition 1 (.1)
AT4G01970 48 / 8e-07
RS4, ATSTS
raffinose synthase 4, stachyose synthase (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10039945
129 / 7e-38
AT5G40390 164 / 8e-58
seed imbibition 1-like, raffinose synthase 5, Raffinose synthase family protein (.1)
Lus10027679
127 / 1e-34
AT5G40390 1040 / 0.0
seed imbibition 1-like, raffinose synthase 5, Raffinose synthase family protein (.1)
Lus10022240
69 / 4e-14
AT5G40390 744 / 0.0
seed imbibition 1-like, raffinose synthase 5, Raffinose synthase family protein (.1)
Lus10022241
65 / 6e-13
AT5G40390 342 / 2e-112
seed imbibition 1-like, raffinose synthase 5, Raffinose synthase family protein (.1)
Lus10017516
56 / 2e-09
AT5G20250 1145 / 0.0
raffinose synthase 6, DARK INDUCIBLE 10, Raffinose synthase family protein (.1.2.3.4)
Lus10007537
54 / 1e-08
AT3G57520 1235 / 0.0
raffinose synthase 2, seed imbibition 2 (.1.2.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.T124908
78 / 3e-17
AT5G40390 1055 / 0.0
seed imbibition 1-like, raffinose synthase 5, Raffinose synthase family protein (.1)
Potri.007G123400
74 / 6e-16
AT5G40390 1147 / 0.0
seed imbibition 1-like, raffinose synthase 5, Raffinose synthase family protein (.1)
Potri.004G207900
71 / 1e-14
AT5G40390 1145 / 0.0
seed imbibition 1-like, raffinose synthase 5, Raffinose synthase family protein (.1)
Potri.017G036700
70 / 1e-14
AT5G40390 1118 / 0.0
seed imbibition 1-like, raffinose synthase 5, Raffinose synthase family protein (.1)
Potri.014G118400
58 / 2e-10
AT4G01970 1087 / 0.0
raffinose synthase 4, stachyose synthase (.1)
Potri.002G193700
56 / 2e-09
AT4G01970 1040 / 0.0
raffinose synthase 4, stachyose synthase (.1)
Potri.018G126400
51 / 7e-08
AT5G20250 1218 / 0.0
raffinose synthase 6, DARK INDUCIBLE 10, Raffinose synthase family protein (.1.2.3.4)
Potri.006G065700
51 / 9e-08
AT5G20250 1154 / 0.0
raffinose synthase 6, DARK INDUCIBLE 10, Raffinose synthase family protein (.1.2.3.4)
Potri.011G166700
50 / 1e-07
AT1G55740 1154 / 0.0
raffinose synthase 1, seed imbibition 1 (.1)
Potri.015G086400
50 / 1e-07
AT3G57520 827 / 0.0
raffinose synthase 2, seed imbibition 2 (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0058
Glyco_hydro_tim
PF05691
Raffinose_syn
Raffinose synthase or seed imbibition protein Sip1
Representative CDS sequence
>Lus10041370 pacid=23179982 polypeptide=Lus10041370 locus=Lus10041370.g ID=Lus10041370.BGIv1.0 annot-version=v1.0
ATGGCCACCCAATCCTCACCCAAGTCCCCTCCAACATCACCACCTCTTTCATACCTTCCCATCAATCCACTTCCGCCGGTTGCTTCGTCGGGTTCCATAG
GGGTGCTGATAGGGATCCGGTTCACGAGCATTTTCCGGTTCAAAGTCTGGTGGACTACCCACTGGGTCGGGAACAGGGGCAGGGAAGTCCAACTCGAGAC
CCAAGTGATAATCCTCAACCGAAACGACGACGGATTGACCAGCATGCTCAACTCCGGCGGTGCGATTCAGTTGGTTGAGATGAAGGAAGAGGTCGGACAC
TTGATGATGCGCGTTGGAGTGAAGGGATGTGGAGAAATGAAGGCTTCTGCTTCGGAGAAGCCGTCAAGTTGTTGGATGGATGGGAAGGAATTTGAGTTTC
AGTATGATGATGATCAGATGGTCACTATTGGAATCCCTTGGCCGGGATCTTGTAGGTTTACTGAAATCGAATACAGATTTTGA
AA sequence
>Lus10041370 pacid=23179982 polypeptide=Lus10041370 locus=Lus10041370.g ID=Lus10041370.BGIv1.0 annot-version=v1.0
MATQSSPKSPPTSPPLSYLPINPLPPVASSGSIGVLIGIRFTSIFRFKVWWTTHWVGNRGREVQLETQVIILNRNDDGLTSMLNSGGAIQLVEMKEEVGH
LMMRVGVKGCGEMKASASEKPSSCWMDGKEFEFQYDDDQMVTIGIPWPGSCRFTEIEYRF
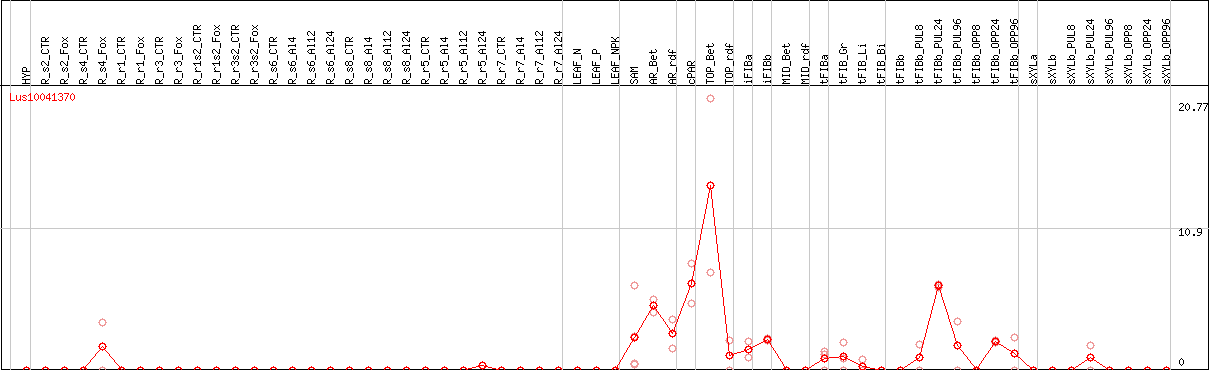
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10041370 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.