External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G38130 287 / 3e-100
ATMAK3
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2)
AT5G11340 62 / 3e-12
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1)
AT1G03150 39 / 0.0007
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1)
AT2G06025 39 / 0.001
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10012579
384 / 1e-138
AT2G38130 286 / 7e-100
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2)
Lus10008088
56 / 2e-09
AT5G11340 249 / 2e-85
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1)
Lus10013120
51 / 5e-08
AT5G11340 256 / 8e-89
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1)
Lus10042595
41 / 0.0002
AT1G03150 348 / 6e-125
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1)
Lus10037996
41 / 0.0002
AT2G39000 335 / 2e-115
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.3)
Lus10009229
39 / 0.0007
AT2G39000 320 / 5e-112
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.3)
Lus10021559
39 / 0.001
AT2G06025 244 / 2e-80
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G435300
316 / 2e-111
AT2G38130 299 / 6e-105
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2)
Potri.001G432400
315 / 2e-111
AT2G38130 296 / 4e-104
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2)
Potri.001G429200
315 / 3e-111
AT2G38130 299 / 6e-105
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2)
Potri.011G139300
308 / 3e-108
AT2G38130 294 / 1e-102
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2)
Potri.001G425800
222 / 5e-75
AT2G38130 207 / 2e-69
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2)
Potri.001G433800
121 / 8e-36
AT2G38130 117 / 3e-34
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2)
Potri.006G248900
61 / 1e-11
AT5G11340 280 / 3e-98
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1)
Potri.018G032400
57 / 2e-10
AT5G11340 270 / 4e-94
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1)
Potri.006G142100
40 / 0.0007
AT2G06025 347 / 1e-120
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1)
Potri.008G039300
39 / 0.0008
AT2G39000 387 / 1e-135
Acyl-CoA N-acyltransferases (NAT) superfamily protein (.1), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.2), Acyl-CoA N-acyltransferases (NAT) superfamily protein (.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0257
Acetyltrans
PF00583
Acetyltransf_1
Acetyltransferase (GNAT) family
Representative CDS sequence
>Lus10041514 pacid=23146704 polypeptide=Lus10041514 locus=Lus10041514.g ID=Lus10041514.BGIv1.0 annot-version=v1.0
ATGGACGGTGAGCGAGGTGTAGAAGGAGACACCCCAAAAGAAGAGAGAGCTACATTCGATGAATCGGAGATAGAGTACGTGAGCTACGGCGGCGAGCATC
ACCTACCGCTAATAATGGGTCTGGTTGATCAGGAGCTGAGTGAACCCTACTCCATCTTCACTTATCGCTACTTCGTATACCTATGGCCCCAGCTTTCCTT
TCTGGCGTTCCATAGAGGGAGATGTGTTGGGACGGTGGTATGCAAAATGGGGGAGCATCGTAATACTACTTTCAGAGGGTACATAGCAATGCTGGTTGTC
ATCAAGCCTTATCGGGGCAAAGGCATTGCTACTGAACTTGTAACCAGATCGATCAGAGCAATGATGGACTCCGGATGTGAAGAGGTAACACTGGAAGCAG
AAGTAACAAATAAAGGTGCTCTTGCACTTTATGGCAGACTGGGGTTCATCAGGGCGAAACGACTGTTTCATTACTACTTGAATGGAGTAGACGCTTTTCG
TCTGAAATTGCGGTTCCCTCAACTTGAGATGAACCCATCTGCCATTAATTTTCTGGACGAGACACAGAGATGA
AA sequence
>Lus10041514 pacid=23146704 polypeptide=Lus10041514 locus=Lus10041514.g ID=Lus10041514.BGIv1.0 annot-version=v1.0
MDGERGVEGDTPKEERATFDESEIEYVSYGGEHHLPLIMGLVDQELSEPYSIFTYRYFVYLWPQLSFLAFHRGRCVGTVVCKMGEHRNTTFRGYIAMLVV
IKPYRGKGIATELVTRSIRAMMDSGCEEVTLEAEVTNKGALALYGRLGFIRAKRLFHYYLNGVDAFRLKLRFPQLEMNPSAINFLDETQR
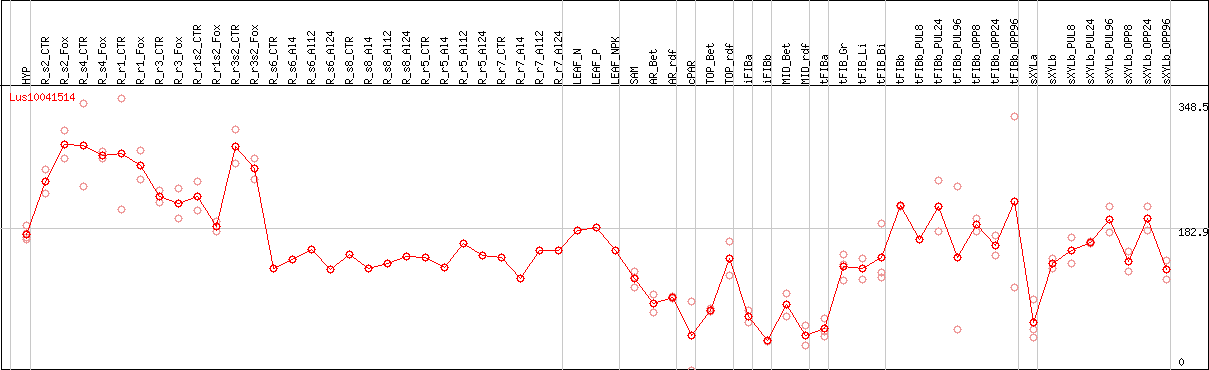
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10041514 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.