Lus10041607 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10041607 pacid=23146600 polypeptide=Lus10041607 locus=Lus10041607.g ID=Lus10041607.BGIv1.0 annot-version=v1.0
ATGGATGACATCAGCGACCTCTTTTTCTCCGGTCCTCCCCCCGACGACGATCACCACCACCTCCTCAGTCGCCGTCAGGATGTCAACACCGATTCCTCCT
ACTCGTCCCCTAAAATCTTCGACCGCTACTTCTCCTCCTCTTCCGACGACGAGGCCGAGGGCGATTACTCACGCCCGTCTAATTCCTCTTCTATGGAAGC
TACCAGCAAGCGTCTTGATTACATGATCCAGTTTCTTGACCGGAAAATCTCCTCTGATAACGACCCCAACAACTGTTCACTCCCGCTTGACCTTGATGTC
GGCAATAACAACAACAACAACAAGTCGCCCTCCTCATCTTCTTTAACGCCACTGCCGGAATTCGTCGGGAGAGGAGGCGGGACCGGGATTTTCAGGCTTC
CAGTTCGCGCTGCGCTTCATCCTTCTCGTCCTCCCAGTCTAGAAGTTAGGCCGCATCCTCTGAGAGAAACTCAGATTGGATGTTCCGTGCGGAACATTGT
CACATCCAGCTCTCAGCTGTGGGCAGGGAAGGGAGATGGGGGACTCCAGGTGTGGCAATTTCGCGATTTGTATGGAGGAACTGAGGATATAGCGCCGTTT
CAAGAGTCTGCGGCCTCTGGATCTGGGGTTCTTTGTTTGGTGGCAGACGAAGGGAGTCGGGTTATTTGGAGTGGGCATAAGGATGGGAGGATCAGGTGCT
GGAAAATGGATTCAGTTTCCGATGGCTTCAAGGAATGCCTTTCATGGATAGCTCACCGAGGCCCTGTTCTCTCCATGGTCATTACTAGCTACGGGGACTT
ATGGTCTGGTTCGGAGAGTGGTGCTATGAAGATATGGCCATGGGAAGCTATAGAAAAATCATTTTCTTATACACCGGAAGAAAGACATGTGGCTGCAATG
TTGGTGGAGAGATCATATATTGACCTTCGGGGCCAATTCAATGTGAATGGGTTGTCGAATGCATTAACATCTGATGTCATGTACTTGCTATCTGACAATT
TTAAGGGGAAAGTTTGGAGTGCTAGTTATCTGTCCTTTGCTCTATGGGATGCTCGTTCAAAGGAGTTGCTAAAGTTGTTCAATATAGATGGCCAGTTAGA
AAGAATGGATATGGTATCAGGACAAGACTTTACTCTTGAGGAAGATATCAAGGTGAAAATTGTTGCTGGATCCAAGAAAGATAAGGCACAAAACTCATTT
GGTTTCTTCCAGCGTTCGAGGAATGCTATAACTGATGCTGTTCGCCGAGTTGCTGCAAAAGGTGGATTTGGAGACGATCGCAGAACTGAGGCAATAACAC
TAACATCAGACGGAATAATATGGACTGGCTGTACGAATGGAGCAATTGTTCAGTGGGATGGGAATGGGAATCGTTTACAGGAAATTCAATACCAGTTTGC
TGCTGTTCGATGCTTTTGTTCCTTCGGGTCACGTGTATGGGTAGGCTATGTCAGTGGTATGATCCAGGTTTTTGACCTTGAGGGTAATCTGCTTGGAGAG
TGGGTAGCTCACAGTAGTGTGGTGACTGAAATGGCTGCGGGTTCTGGTTATGTTTTCTCATTGGCTAATCATGGTGGTATCCGGGGCTGGAATGCCATGG
CTCCTGGACCTATTGATGGAATTCTGCGATCAGAATTGGCAGGAAAAGAATTCTTGTATACAAGAATAGAAAATTTGAAAGTTTTGACTGGTACATGGAA
TGTAGCACAAGAAAGAGCTAGTCGTGACTCACTTGTGTCCTGGCTAGGTTCTGCAGCAGCAGATGTTGGAATTGTTGTTGTTGGGTTGCAAGAGGTTGAA
ATGGGAGCTGGAGTCCTCGCAATGTCGGCAGCAAAAGAAACTGTTGGACTTGAAGGTAGCGCTCTTGGGCAGTGGTGGCTAGATATGATAGGCAAAACAT
TAGATGAAGGGTCAACATTTGAGCGTGTTGGTTCTCGGCAGCTAGCGGGGTTGCTCATCTCTATATGGGTTAGGAGTAATTTAAAACCCCATGTTGTGTG
TGTTGATGCTGCTGCTGTCCCATGTGGATTTGGGCGTGCAATTGGTAACAAGGGAGCTGTTGGTCTGAGGATCAGAGTTTTTGACCGGATAATATGTTTC
ATTAATTGTCACTTTGCGGCACATTTAGAAGCAGTTGGGCGACGCAATGATGACTTTGATCATGTTTATCGAACCATGTCATTTGGCCGATCATCAAGCC
ACTTTGCTGGTGCAGCTGGTATGGTGCCGTTCCTGGTCTTCTCTTGCACGCTTGCCTGCTCAATGTGTTTATTTTGGCTTGTTTATAGATCTGGATTGCT
ATTGCTCCTTTCTGTTGCAGTTGGGACTTCATCTGCTGGTCAAATGCTTCGTGGTACAACCGTTTTCAATGGAAGCTCGGCAAATACAATTCCAGAGCTA
TCTGAGGCAGATATGGTCGTATTTCTTGGTGACTTTAATTATCGGCTGGGTAATATTTCCTATGACGAAGCAAGGGATTTTATTTCTCAAAGATGCTTTG
ATTGGCTTAAAGAAAGGGATCAGCTACACTCAGAAATGAAAGCTGGGAATGTTTTCCAAGGAATGCGTGAAGCAGTCATTAGATTTCCTCCTACATACAA
GTTTGACAAGCACCAGCATGGTTTAGCAGGATACGATTCTGGTGAAAAGAAGCGTATTCCGGCCTGGTGTGACCGAATACTGTACCGTGACAGCCGGTCA
GCTGGGGTGTCTGAGTGCAGCTTACAGTGCCCTGTTGTAGGTTTGATTTCGCAGTACGATGCTTGCATGGATGTGACTGACAGTGATCACAAGCCTGTAC
GTTGCATCTTTAGTGTCGATATTGCAAAAGTAGATGAGTCTGTAAGAAGACAAGAACTTGGAGACATTATGAACTCAAATGTGGAAATCAGATCCATACT
AAAAGAGCTATGCAAGATTCCGGAAACTATTGTTAGCACAAATAACATAATCCTGCAAAACCTGGACAGAACCATTTTACGGATGACAAATAAATCAGAT
CAATTTGATGCTCTGTATCAAATCGTCTGCGAGGGCGAGTCAACTGTTGATGAGAATGGACAAGCTGGTGATCATCATCCAAGGGGTTCCTTTGGGTTTC
CTCAGTGGCTTCAGGTAATTCCTGCGGCAGGCGTGATTAAACCGGGAAACATTGCAGAGATTGGGGTCCACCACGAGGATTTCTCAACCGTCGAGGAGTT
TGTGGACGGTGTCCCGCAGAACTCCTGGTGTGAAGACATCAGAGACAAGGAAGTTATATTGGTCGTGAAAGTACGAGGCACTTACAACACGACCGAGACT
AGAAATCATAGGATTCGTGTGCGCCATTGTCGTTCATCACGAGTTGATCCAAATGTTGACAGCAATAAGTCGCAGGGGAGTCTTCTCCACCGGGCTGATT
ATCAAAACCTTAGCAGCTCGTACGATGTGGTGAACCGCCTTCGGAAGTTGAATAGTCCGTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10041607 pacid=23146600 polypeptide=Lus10041607 locus=Lus10041607.g ID=Lus10041607.BGIv1.0 annot-version=v1.0
MDDISDLFFSGPPPDDDHHHLLSRRQDVNTDSSYSSPKIFDRYFSSSSDDEAEGDYSRPSNSSSMEATSKRLDYMIQFLDRKISSDNDPNNCSLPLDLDV
GNNNNNNKSPSSSSLTPLPEFVGRGGGTGIFRLPVRAALHPSRPPSLEVRPHPLRETQIGCSVRNIVTSSSQLWAGKGDGGLQVWQFRDLYGGTEDIAPF
QESAASGSGVLCLVADEGSRVIWSGHKDGRIRCWKMDSVSDGFKECLSWIAHRGPVLSMVITSYGDLWSGSESGAMKIWPWEAIEKSFSYTPEERHVAAM
LVERSYIDLRGQFNVNGLSNALTSDVMYLLSDNFKGKVWSASYLSFALWDARSKELLKLFNIDGQLERMDMVSGQDFTLEEDIKVKIVAGSKKDKAQNSF
GFFQRSRNAITDAVRRVAAKGGFGDDRRTEAITLTSDGIIWTGCTNGAIVQWDGNGNRLQEIQYQFAAVRCFCSFGSRVWVGYVSGMIQVFDLEGNLLGE
WVAHSSVVTEMAAGSGYVFSLANHGGIRGWNAMAPGPIDGILRSELAGKEFLYTRIENLKVLTGTWNVAQERASRDSLVSWLGSAAADVGIVVVGLQEVE
MGAGVLAMSAAKETVGLEGSALGQWWLDMIGKTLDEGSTFERVGSRQLAGLLISIWVRSNLKPHVVCVDAAAVPCGFGRAIGNKGAVGLRIRVFDRIICF
INCHFAAHLEAVGRRNDDFDHVYRTMSFGRSSSHFAGAAGMVPFLVFSCTLACSMCLFWLVYRSGLLLLLSVAVGTSSAGQMLRGTTVFNGSSANTIPEL
SEADMVVFLGDFNYRLGNISYDEARDFISQRCFDWLKERDQLHSEMKAGNVFQGMREAVIRFPPTYKFDKHQHGLAGYDSGEKKRIPAWCDRILYRDSRS
AGVSECSLQCPVVGLISQYDACMDVTDSDHKPVRCIFSVDIAKVDESVRRQELGDIMNSNVEIRSILKELCKIPETIVSTNNIILQNLDRTILRMTNKSD
QFDALYQIVCEGESTVDENGQAGDHHPRGSFGFPQWLQVIPAAGVIKPGNIAEIGVHHEDFSTVEEFVDGVPQNSWCEDIRDKEVILVVKVRGTYNTTET
RNHRIRVRHCRSSRVDPNVDSNKSQGSLLHRADYQNLSSSYDVVNRLRKLNSP
|
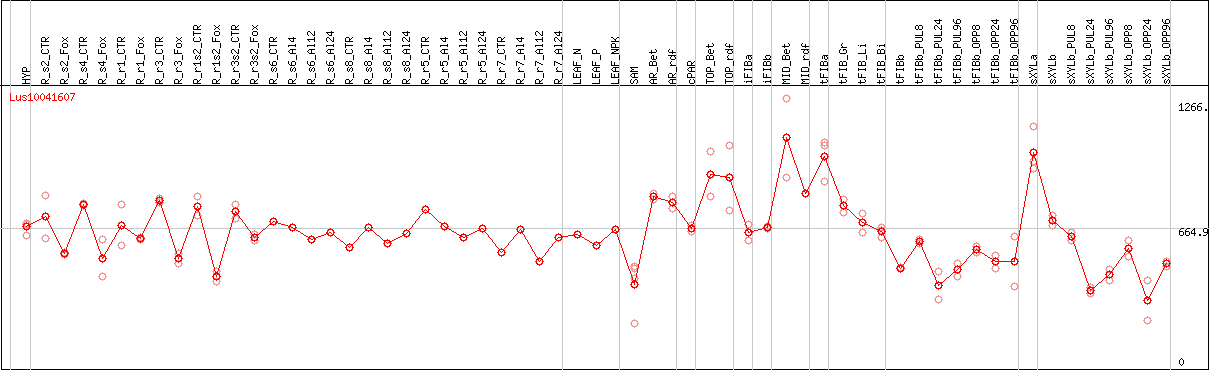
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10041607 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
