External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G36380 634 / 0
ROT3
ROTUNDIFOLIA 3, Cytochrome P450 superfamily protein (.1)
AT3G13730 477 / 2e-165
CYP90D1
"cytochrome P450, family 90, subfamily D, polypeptide 1", cytochrome P450, family 90, subfamily D, polypeptide 1 (.1)
AT5G05690 325 / 4e-106
CBB3, DWF3, CYP90A1, CYP90A, CPD
DWARF 3, CYTOCHROME P450 90A1, CONSTITUTIVE PHOTOMORPHOGENIC DWARF, CABBAGE 3, Cytochrome P450 superfamily protein (.1.2.3)
AT3G50660 306 / 1e-98
PSC1, CYP90B1, CLM, SNP2, DWF4, SAV1
SUPPRESSOR OF NPH4 2, SHADE AVOIDANCE 1, PARTIALLY SUPPRESSING COI1 INSENSITIVITY TO JA 1, DWARF 4, CYTOCHROME P450 90B1, CLOMAZONE-RESISTANT, Cytochrome P450 superfamily protein (.1)
AT5G38970 261 / 1e-81
ATBR6OX, CYP85A1, BR6OX1
brassinosteroid-6-oxidase 1 (.1.2.3)
AT1G73340 259 / 3e-80
Cytochrome P450 superfamily protein (.1)
AT1G12740 257 / 6e-80
CYP87A2
"cytochrome P450, family 87, subfamily A, polypeptide 2", cytochrome P450, family 87, subfamily A, polypeptide 2 (.1.2)
AT3G30180 251 / 9e-78
CYP85A2, BR6OX2
brassinosteroid-6-oxidase 2 (.1)
AT4G19230 249 / 4e-77
CYP707A1
"cytochrome P450, family 707, subfamily A, polypeptide 1", cytochrome P450, family 707, subfamily A, polypeptide 1 (.1.2)
AT5G45340 236 / 4e-72
CYP707A3
"cytochrome P450, family 707, subfamily A, polypeptide 3", cytochrome P450, family 707, subfamily A, polypeptide 3 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10028345
944 / 0
AT4G36380 652 / 0.0
ROTUNDIFOLIA 3, Cytochrome P450 superfamily protein (.1)
Lus10040193
335 / 5e-110
AT5G05690 687 / 0.0
DWARF 3, CYTOCHROME P450 90A1, CONSTITUTIVE PHOTOMORPHOGENIC DWARF, CABBAGE 3, Cytochrome P450 superfamily protein (.1.2.3)
Lus10014850
328 / 2e-107
AT5G05690 679 / 0.0
DWARF 3, CYTOCHROME P450 90A1, CONSTITUTIVE PHOTOMORPHOGENIC DWARF, CABBAGE 3, Cytochrome P450 superfamily protein (.1.2.3)
Lus10006152
281 / 2e-90
AT3G13730 378 / 2e-128
"cytochrome P450, family 90, subfamily D, polypeptide 1", cytochrome P450, family 90, subfamily D, polypeptide 1 (.1)
Lus10024495
271 / 2e-85
AT1G12740 751 / 0.0
"cytochrome P450, family 87, subfamily A, polypeptide 2", cytochrome P450, family 87, subfamily A, polypeptide 2 (.1.2)
Lus10016065
268 / 6e-84
AT3G50660 399 / 3e-134
SUPPRESSOR OF NPH4 2, SHADE AVOIDANCE 1, PARTIALLY SUPPRESSING COI1 INSENSITIVITY TO JA 1, DWARF 4, CYTOCHROME P450 90B1, CLOMAZONE-RESISTANT, Cytochrome P450 superfamily protein (.1)
Lus10030404
254 / 3e-80
AT5G05690 531 / 0.0
DWARF 3, CYTOCHROME P450 90A1, CONSTITUTIVE PHOTOMORPHOGENIC DWARF, CABBAGE 3, Cytochrome P450 superfamily protein (.1.2.3)
Lus10014240
252 / 3e-78
AT3G30180 531 / 0.0
brassinosteroid-6-oxidase 2 (.1)
Lus10000647
252 / 4e-78
AT3G30180 540 / 0.0
brassinosteroid-6-oxidase 2 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.007G018400
654 / 0
AT4G36380 626 / 0.0
ROTUNDIFOLIA 3, Cytochrome P450 superfamily protein (.1)
Potri.001G200100
516 / 0
AT3G13730 663 / 0.0
"cytochrome P450, family 90, subfamily D, polypeptide 1", cytochrome P450, family 90, subfamily D, polypeptide 1 (.1)
Potri.003G038200
516 / 1e-180
AT3G13730 662 / 0.0
"cytochrome P450, family 90, subfamily D, polypeptide 1", cytochrome P450, family 90, subfamily D, polypeptide 1 (.1)
Potri.010G189800
339 / 9e-112
AT5G05690 717 / 0.0
DWARF 3, CYTOCHROME P450 90A1, CONSTITUTIVE PHOTOMORPHOGENIC DWARF, CABBAGE 3, Cytochrome P450 superfamily protein (.1.2.3)
Potri.008G067500
338 / 2e-111
AT5G05690 710 / 0.0
DWARF 3, CYTOCHROME P450 90A1, CONSTITUTIVE PHOTOMORPHOGENIC DWARF, CABBAGE 3, Cytochrome P450 superfamily protein (.1.2.3)
Potri.007G026500
303 / 1e-97
AT3G50660 757 / 0.0
SUPPRESSOR OF NPH4 2, SHADE AVOIDANCE 1, PARTIALLY SUPPRESSING COI1 INSENSITIVITY TO JA 1, DWARF 4, CYTOCHROME P450 90B1, CLOMAZONE-RESISTANT, Cytochrome P450 superfamily protein (.1)
Potri.005G124000
298 / 1e-95
AT3G50660 738 / 0.0
SUPPRESSOR OF NPH4 2, SHADE AVOIDANCE 1, PARTIALLY SUPPRESSING COI1 INSENSITIVITY TO JA 1, DWARF 4, CYTOCHROME P450 90B1, CLOMAZONE-RESISTANT, Cytochrome P450 superfamily protein (.1)
Potri.010G156800
278 / 5e-88
AT3G30180 561 / 0.0
brassinosteroid-6-oxidase 2 (.1)
Potri.001G109100
266 / 2e-83
AT1G12740 770 / 0.0
"cytochrome P450, family 87, subfamily A, polypeptide 2", cytochrome P450, family 87, subfamily A, polypeptide 2 (.1.2)
Potri.003G122500
263 / 2e-82
AT1G12740 770 / 0.0
"cytochrome P450, family 87, subfamily A, polypeptide 2", cytochrome P450, family 87, subfamily A, polypeptide 2 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00067
p450
Cytochrome P450
Representative CDS sequence
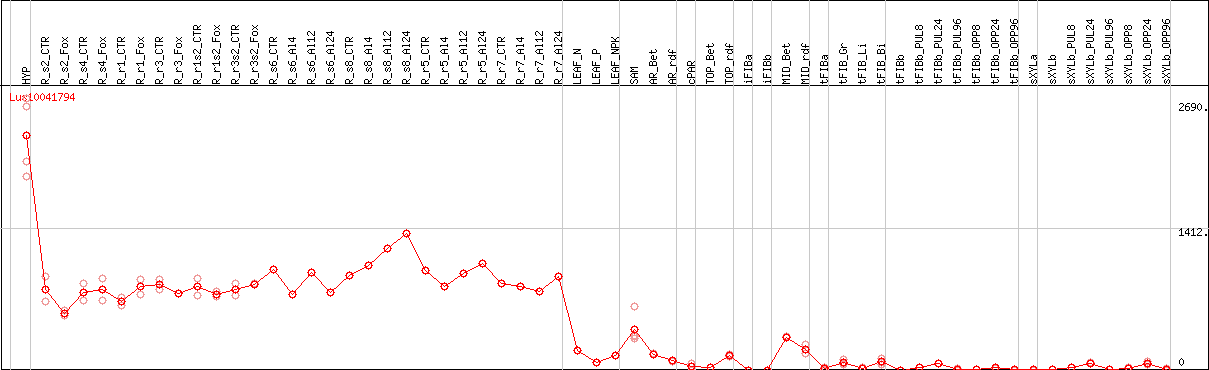
>Lus10041794 pacid=23146831 polypeptide=Lus10041794 locus=Lus10041794.g ID=Lus10041794.BGIv1.0 annot-version=v1.0
ATGCTGAATTTGGAGATGTCGTCGGAAGGGATGTGGTGGTCGGTAGGATCAGTGATTCTCGCCGTCTTCTTTGGAGCAGCTCTTCCGCTCTACTGGTGGC
GGCGCCGGCTACCCACGGCGGCGTCGTCGTCGGAGGCCGTTGTGCCGAGAGGGAATTTGGGATGGCCTTTGATCGGAGAGACTCTTGACTTTATCGCCTG
CGGATACACTTCTCGCCCTGTTTCTTTCATGGACACTCGCAAATCTCTATATGGGAATGTATTCAAATCGAACATACTGGGAACTCCGATAATCGTATCC
ACCGATCCGGAAGTGAACAAAGTAGTGTTACAGAACCATGGAAACAACTCATTCATCCCGGCTTACCCTAAATCGATCCGCGAAATATTCGGCAAAGAGT
CCATTCTACAAACCAATGGTCCTATGCAGAAGAAAGTCCACGGCTTGATCGGTGGGTTCTTGAGATCCCCTCAGTTCAAAGCCAGAATCGCCAAGGACAT
CGAGGACTCTGTGAAGTTCACTTTGGACACTTGGAAGGACTTACCCGTCGTTTATGTCCAGGATGAAATAAAAAAGGTCTTGATGACGTTCAAAGTGCTG
GTGAAAGCTCTGATTAGCGTCGGGCCAGGGGCAGATTTGGAATTCATGAAATCTGAATTTGAGGAGCTTATCAAAGGCTTAATCTGCTTCCCCATCAAAG
TCCCTGGAACCACTCTTTACAAGTCATTGAAGGCGAGAGAGAGGTTGTTGAAGTTTGTGAGGAGAAAGGTGGACGAAAGGAAAAGCAAAACGACGACGGA
CGAGGGTCCACGGGATGCAGTGGACGTGCTGCTAGGGGAAAAAGAGGAGAAACAACAACAACAAATGCTGTCGTTGGATTTCATAAGCGGGAACGTGTTG
GAGATGATGATCCCAGGGGAAGAGACGGTGCCGACGGCGATGACGTTAGCTGTCAAGTTCTTGAGTGATTATCCACGCGCTTTAGCCCAACTAACGGAGG
AGAACTTGGTTTTGAAGAGGCAGAAGGAAGAGTTGGGTGAGGAGTATACGTGGACTGACTACGTGTCCTTGCCATTTACTCAGAATGTGATCAGTGAGAC
ACTAAGAATGGCAAATATCATTAATGGGGTTTGGAGGAAAGCTCTAAAAAATGTGGAAATTAAAGGACATTTGATTCCGAAAGGATGGTGTGTTTTGGCA
TCTTTCACCTCTGTTCATATGAATGAAGATAACTACCAACACCCTTACGAATTCAATCCATTGAGATGGGAGCAAAAGGGAGCTTCAGTAAACAACAATT
GCTACACACCATTTGGTGGAGGGCAAAGGCTATGCCCTGGTTTGGAGCTGTCCAGGCTTGAAATCTCTATTTTCCTTCACCACCTAGTCACCACTTACAA
ATGGAGTGCTGAGAAGGATGACATCGTCTACTTTCCAACTGTGAAGATGAAGAGGAAGATGCCCATCACTGTCACACCTTTTAATACATAA
AA sequence
>Lus10041794 pacid=23146831 polypeptide=Lus10041794 locus=Lus10041794.g ID=Lus10041794.BGIv1.0 annot-version=v1.0
MLNLEMSSEGMWWSVGSVILAVFFGAALPLYWWRRRLPTAASSSEAVVPRGNLGWPLIGETLDFIACGYTSRPVSFMDTRKSLYGNVFKSNILGTPIIVS
TDPEVNKVVLQNHGNNSFIPAYPKSIREIFGKESILQTNGPMQKKVHGLIGGFLRSPQFKARIAKDIEDSVKFTLDTWKDLPVVYVQDEIKKVLMTFKVL
VKALISVGPGADLEFMKSEFEELIKGLICFPIKVPGTTLYKSLKARERLLKFVRRKVDERKSKTTTDEGPRDAVDVLLGEKEEKQQQQMLSLDFISGNVL
EMMIPGEETVPTAMTLAVKFLSDYPRALAQLTEENLVLKRQKEELGEEYTWTDYVSLPFTQNVISETLRMANIINGVWRKALKNVEIKGHLIPKGWCVLA
SFTSVHMNEDNYQHPYEFNPLRWEQKGASVNNNCYTPFGGGQRLCPGLELSRLEISIFLHHLVTTYKWSAEKDDIVYFPTVKMKRKMPITVTPFNT