External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G35770 181 / 1e-58
ATSEN1, DIN1, SEN1
SENESCENCE ASSOCIATED GENE 1, DARK INDUCIBLE 1, ARABIDOPSIS THALIANA SENESCENCE 1, Rhodanese/Cell cycle control phosphatase superfamily protein (.1.2.3)
AT5G66040 166 / 1e-53
STR16
sulfurtransferase protein 16 (.1.2)
AT5G66170 112 / 4e-32
STR18
sulfurtransferase 18 (.1.2.3)
AT2G21045 99 / 1e-26
Rhodanese/Cell cycle control phosphatase superfamily protein (.1)
AT2G17850 89 / 1e-22
Rhodanese/Cell cycle control phosphatase superfamily protein (.1)
AT4G27700 60 / 3e-11
Rhodanese/Cell cycle control phosphatase superfamily protein (.1)
AT4G24750 44 / 3e-05
Rhodanese/Cell cycle control phosphatase superfamily protein (.1)
AT2G42220 39 / 0.0008
Rhodanese/Cell cycle control phosphatase superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10028390
268 / 8e-93
AT4G35770 182 / 3e-59
SENESCENCE ASSOCIATED GENE 1, DARK INDUCIBLE 1, ARABIDOPSIS THALIANA SENESCENCE 1, Rhodanese/Cell cycle control phosphatase superfamily protein (.1.2.3)
Lus10005635
157 / 4e-49
AT5G66040 144 / 1e-44
sulfurtransferase protein 16 (.1.2)
Lus10041525
146 / 5e-43
AT5G14030 216 / 8e-70
translocon-associated protein beta (TRAPB) family protein (.1), translocon-associated protein beta (TRAPB) family protein (.2), translocon-associated protein beta (TRAPB) family protein (.3), translocon-associated protein beta (TRAPB) family protein (.4)
Lus10012566
140 / 9e-43
AT5G66040 129 / 1e-39
sulfurtransferase protein 16 (.1.2)
Lus10041895
91 / 5e-23
AT5G66170 116 / 3e-33
sulfurtransferase 18 (.1.2.3)
Lus10028442
89 / 3e-22
AT5G66170 119 / 2e-34
sulfurtransferase 18 (.1.2.3)
Lus10041894
88 / 3e-22
AT5G66170 120 / 1e-35
sulfurtransferase 18 (.1.2.3)
Lus10021227
84 / 1e-20
AT5G66040 73 / 4e-17
sulfurtransferase protein 16 (.1.2)
Lus10028441
68 / 1e-14
AT5G66170 89 / 9e-24
sulfurtransferase 18 (.1.2.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G106400
210 / 3e-69
AT4G35770 199 / 1e-64
SENESCENCE ASSOCIATED GENE 1, DARK INDUCIBLE 1, ARABIDOPSIS THALIANA SENESCENCE 1, Rhodanese/Cell cycle control phosphatase superfamily protein (.1.2.3)
Potri.002G014900
163 / 2e-51
AT4G35770 151 / 8e-47
SENESCENCE ASSOCIATED GENE 1, DARK INDUCIBLE 1, ARABIDOPSIS THALIANA SENESCENCE 1, Rhodanese/Cell cycle control phosphatase superfamily protein (.1.2.3)
Potri.014G131300
113 / 3e-32
AT2G21045 204 / 2e-68
Rhodanese/Cell cycle control phosphatase superfamily protein (.1)
Potri.005G111200
106 / 2e-29
AT2G17850 162 / 5e-52
Rhodanese/Cell cycle control phosphatase superfamily protein (.1)
Potri.012G020700
58 / 3e-10
AT4G27700 262 / 4e-89
Rhodanese/Cell cycle control phosphatase superfamily protein (.1)
Potri.015G008000
54 / 8e-09
AT4G27700 268 / 2e-91
Rhodanese/Cell cycle control phosphatase superfamily protein (.1)
Potri.006G100600
50 / 1e-07
AT3G08920 238 / 6e-80
Rhodanese/Cell cycle control phosphatase superfamily protein (.1)
Potri.006G059200
45 / 9e-06
AT2G42220 318 / 4e-111
Rhodanese/Cell cycle control phosphatase superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0031
Phosphatase
PF00581
Rhodanese
Rhodanese-like domain
Representative CDS sequence
>Lus10041843 pacid=23146951 polypeptide=Lus10041843 locus=Lus10041843.g ID=Lus10041843.BGIv1.0 annot-version=v1.0
ATGGAATCCTGCAGATCAATTGTTTCGTCCAGTACTGCTGGTTTGACCGCCGCTTCGCTTGCTCCGCTTCTTCACCTTCCTTCAAATGTGAAAGTCTTCT
CCAGGGTACAATCTCATTCCCAGTTGAGGAAAATACCAATTACTAACAGCAGGAGGAGGATCTCAAGATTCTGCACGAAGGCGGTGGTGGGGGAGAACTT
GAATGCGGCAGCCGTTCCGACATCAGTGCCAGTAAGAGTAGCGTATGAACTTCTGCTAGCTGGGCATCGCTACCTTGATGTCAGGACACCGGAAGAGTTC
AGCGCAGGACACCCAGAAGGAGCAGTCAATGTTCCTTACATGTATAGAGTTGGTTCAGGTTTGGCAAAGAACCCAAAATTTGTAGAGCAAGTTTCATCAC
AGTTTGGCAAGTACAATGAGATCATTGTTGGTTGTCAGCTGGGGAAAAGGTCTATGATGGCGGCAACTGATCTTTTAGCTGCCGGTTTTATGGCAGTGAC
AGACATAGCGGGCGGGTATGCAGCTTGGACGCAGAATGACCTCCCGACTGAGAAGTAA
AA sequence
>Lus10041843 pacid=23146951 polypeptide=Lus10041843 locus=Lus10041843.g ID=Lus10041843.BGIv1.0 annot-version=v1.0
MESCRSIVSSSTAGLTAASLAPLLHLPSNVKVFSRVQSHSQLRKIPITNSRRRISRFCTKAVVGENLNAAAVPTSVPVRVAYELLLAGHRYLDVRTPEEF
SAGHPEGAVNVPYMYRVGSGLAKNPKFVEQVSSQFGKYNEIIVGCQLGKRSMMAATDLLAAGFMAVTDIAGGYAAWTQNDLPTEK
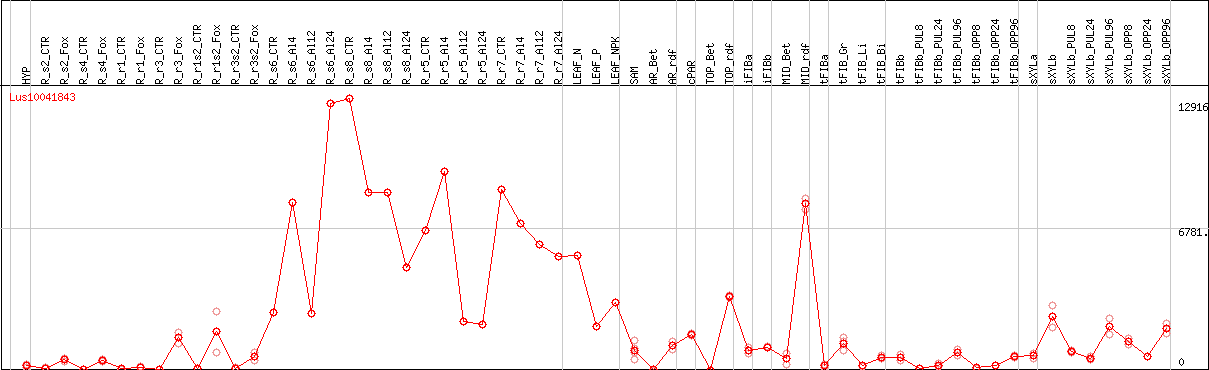
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10041843 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.