External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G53950 281 / 6e-95
NAC
ATCUC2, CUC2, ANAC098
CUP-SHAPED COTYLEDON 2, Arabidopsis NAC domain containing protein 98, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
AT3G15170 258 / 1e-86
NAC
ATNAC1, CUC1, ANAC054
CUP-SHAPED COTYLEDON1, Arabidopsis NAC domain containing protein 54, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
AT5G61430 247 / 3e-82
NAC
ANAC100, ATNAC5
NAC domain containing protein 100 (.1)
AT5G07680 243 / 1e-80
NAC
ANAC079, ATNAC4, ANAC080
Arabidopsis NAC domain containing protein 79, NAC domain containing protein 80 (.1.2)
AT5G18270 239 / 5e-79
NAC
ANAC087
Arabidopsis NAC domain containing protein 87 (.1.2)
AT3G18400 236 / 6e-78
NAC
ANAC058
NAC domain containing protein 58 (.1)
AT2G24430 234 / 2e-77
NAC
ANAC039, ANAC038
Arabidopsis NAC domain containing protein 39, NAC domain containing protein 38 (.1.2)
AT3G29035 234 / 5e-77
NAC
ORS1, AtNAC3, ANAC059
ORE1 SISTER1, Arabidopsis NAC domain containing protein 59, NAC domain containing protein 3 (.1)
AT5G39610 227 / 5e-75
NAC
ORE1, AtNAC2, ANAC092, ATNAC6
ORESARA 1, NAC domain containing protein 2, Arabidopsis NAC domain containing protein 92, NAC domain containing protein 6 (.1)
AT3G04060 226 / 9e-74
NAC
ANAC046
NAC domain containing protein 46 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10005537
390 / 1e-138
AT5G53950 317 / 3e-107
CUP-SHAPED COTYLEDON 2, Arabidopsis NAC domain containing protein 98, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Lus10021659
242 / 6e-80
AT5G61430 361 / 8e-125
NAC domain containing protein 100 (.1)
Lus10001648
242 / 7e-80
AT5G61430 359 / 4e-124
NAC domain containing protein 100 (.1)
Lus10009669
232 / 5e-76
AT3G18400 308 / 3e-104
NAC domain containing protein 58 (.1)
Lus10026879
232 / 8e-76
AT5G18270 325 / 3e-110
Arabidopsis NAC domain containing protein 87 (.1.2)
Lus10020165
233 / 2e-75
AT2G24430 306 / 2e-102
Arabidopsis NAC domain containing protein 39, NAC domain containing protein 38 (.1.2)
Lus10009029
231 / 2e-75
AT3G18400 311 / 3e-105
NAC domain containing protein 58 (.1)
Lus10003435
229 / 1e-74
AT5G18270 315 / 2e-106
Arabidopsis NAC domain containing protein 87 (.1.2)
Lus10026966
228 / 5e-74
AT2G24430 311 / 6e-105
Arabidopsis NAC domain containing protein 39, NAC domain containing protein 38 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.011G115400
281 / 3e-95
AT5G53950 312 / 1e-104
CUP-SHAPED COTYLEDON 2, Arabidopsis NAC domain containing protein 98, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Potri.001G396300
272 / 4e-92
AT5G53950 299 / 5e-100
CUP-SHAPED COTYLEDON 2, Arabidopsis NAC domain containing protein 98, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Potri.012G001400
247 / 9e-82
AT5G61430 448 / 1e-158
NAC domain containing protein 100 (.1)
Potri.019G031600
243 / 1e-79
AT5G18270 328 / 6e-111
Arabidopsis NAC domain containing protein 87 (.1.2)
Potri.015G020000
240 / 6e-79
AT5G61430 446 / 1e-157
NAC domain containing protein 100 (.1)
Potri.013G054200
240 / 1e-78
AT5G18270 339 / 4e-115
Arabidopsis NAC domain containing protein 87 (.1.2)
Potri.005G255900
239 / 4e-78
AT1G76420 295 / 4e-98
CUP SHAPED COTYLEDON3, Arabidopsis NAC domain containing protein 31, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Potri.002G005800
239 / 5e-78
AT1G76420 326 / 7e-110
CUP SHAPED COTYLEDON3, Arabidopsis NAC domain containing protein 31, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
Potri.012G056300
232 / 1e-76
AT3G18400 317 / 6e-108
NAC domain containing protein 58 (.1)
Potri.018G003800
233 / 2e-76
AT2G24430 350 / 1e-120
Arabidopsis NAC domain containing protein 39, NAC domain containing protein 38 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF02365
NAM
No apical meristem (NAM) protein
Representative CDS sequence
>Lus10041924 pacid=23153621 polypeptide=Lus10041924 locus=Lus10041924.g ID=Lus10041924.BGIv1.0 annot-version=v1.0
ATGGATCCACACCCTTTCCACCCTTGCTTCCCATACTCCAACCCTCCTCCATCAACCGACTACACCACCACCGCCAATTTCAACTACACTTCCTCCGCCA
ACCACCACCAACAAAACCAAACCACCAACCTCCCACCAGGCTTCAGATTCCACCCGACCGACGAGGAGCTCATAACCTACTACCTCCTCAAAAAGGTCAT
GGACTCTTCCTTCTCAGGGCGCGCCATAGCCGAGGTGGACCTCAACAAGTGCGAGCCGTGGCACTTGCCGGAGAAGGCCAAGATGGGGGAGAAAGAGTGG
TACTTCTTCTCCCTCCGAGACCGCAAGTACCCGACCGGGCTCCGGACCAACCGAGCCACCGAGGCAGGGTACTGGAAGGCTACGGGGAAAGACAGGGAGA
TCTACAGTAGCAAAAGCTGTGCCCTTGTTGGGATGAAGAAGACTCTTGTTTTCTACCGTGGGAGGGCTCCAAAGGGTGAGAAGAGCAATTGGGTCATGCA
TGAGTATCGTTTGGAAGGGAAGTTCTCCTACCACTTCCTCTCCCGCTCGTCCAAGGTTAGTACGACGTCGTATTTAAGTTTTACTTGTGAACGTGGAAAT
GCATGGTTGTAG
AA sequence
>Lus10041924 pacid=23153621 polypeptide=Lus10041924 locus=Lus10041924.g ID=Lus10041924.BGIv1.0 annot-version=v1.0
MDPHPFHPCFPYSNPPPSTDYTTTANFNYTSSANHHQQNQTTNLPPGFRFHPTDEELITYYLLKKVMDSSFSGRAIAEVDLNKCEPWHLPEKAKMGEKEW
YFFSLRDRKYPTGLRTNRATEAGYWKATGKDREIYSSKSCALVGMKKTLVFYRGRAPKGEKSNWVMHEYRLEGKFSYHFLSRSSKVSTTSYLSFTCERGN
AWL
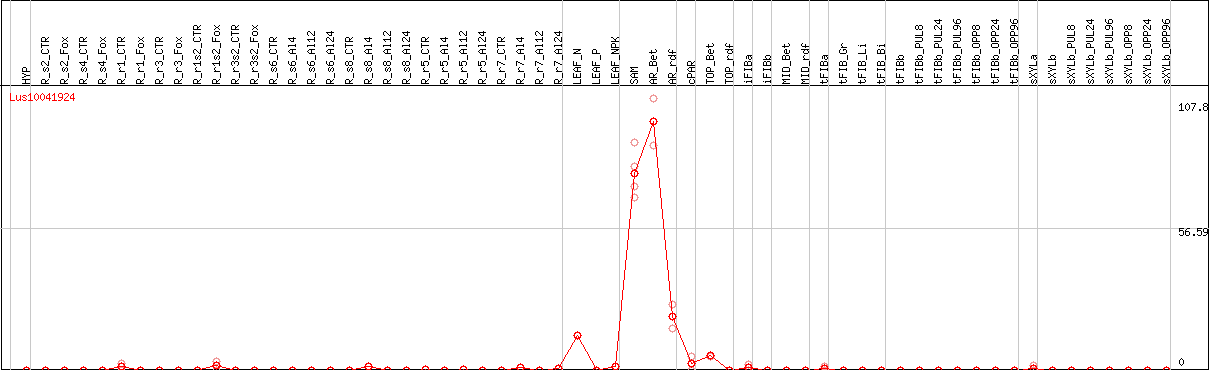
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10041924 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.