Lus10042045 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10042045 pacid=23153676 polypeptide=Lus10042045 locus=Lus10042045.g ID=Lus10042045.BGIv1.0 annot-version=v1.0
ATGGAGAGGTCCACCACGTCTGGGTTTTACAACGTCTTGGCTTCACGAAAAGCCGAGGAAGCTGTTTGGAGGCGTTACCAAGCTACTGAGTGGCTCGAAA
GCCTAGTGGGTCCGTTGAGTATACCGAGCCATCCGTCGGAGAAAGAATTCATTTCTCGATTGAGAAATGGATTGATTCTTTGCAATGCCATTAACAAAGT
CCATCCAGGATCAGTAACTAAGGTTGTGGAGAATCACGTGCCTGTGCAATCCCTAACTCGAGAATCCCAACCATTACCAGCGTATCAGCACTTTGAGAAT
GTAAAGAATTTCCTAGTAGCTATGGGAGATTTGAAACTTCCGGCATTTGAAGCTTCTGATCTAGAAAGGGATACATTGGAACTGGAAGCAGGATCAACTG
CTAAGGTGGTTGATTCAATCTTGGCCTTGAAGTCATACCATGAGCACAAGAATATGAATGGTAGTAATGGGGCTTTCAAGTCTACAAAATCCCCTATGGT
TTCTCACAGTGCGTCGAGGAGAAATCTCCAGGGGTTGTCGTCAGATTCTTGTCGGCGGTTGGAGATGAATGTAGCATGCAAGGGACATCTATCAGGCAAT
GCTGACAGTCAGAAACTGGAAGATGAACTGATCAAGGTACTTGCAGAATACATGGCTGATACAAAAGAAAACATGGGTGACTACTTTCTCACCTCTTTCC
GAGCTGGGAATGTGGATCCAATGAAGCTATTTGGTAGGATCGTGTCAAGTTGTCTCAAGGACAATCTACATAAGTTTCCTGAGGGCCAGACAGTGGTCAT
GGACACCTTGAAAGAGAGTAGTAATTTACCTGCTCATTTGGAAGCGACTCCATTGGATGACATATCTGATCGCAGAAATAGTAAGTGTTGTCGAACATGC
TTAAGGAAAGGCAATTGCAATCACAGGCGTTTACTGCAAGCCCAGGAAAGAGAACTTCAGAACCTGAAGACCTTGTTAACTACAACAAGGAAAGAGTTTG
AAGGCTTGCAATTCCAGTTGCAGAGCGATCTGACACAGTGCAGCATTCAGGTTCAGGAGATGTCTAGAGCTGCTCAAGGATATCACAAAGTGTTGAAAGA
AAACAGAATTCTGTACAACATGGTTCAGGATTTGAAAGGAAATATTCGAGTTTATTGTAGAGTAAGACCATCTTTCACCGCTGGAGCAAGCAATGCAATG
GAATTTGTTGGAGAGGATGGCTCACTAGTGATTGTAGATCCATCAAAGCCCACGAAAGATGGAAGGAGGCATTTTCAGTTCAACCAGGTTTTTGGTCCTA
CCGCCACCCAAGAAAATGTGTTTGAACATGTCCAACCCCTAATCAGATCAGTGATGGATGGCCACAATGTATGCATTTTTGCTTATGGCCAAACCGGCTC
TGGTAAAACATACACAATGAGTGGTCAAGGTGGATCTACCAAGGAGCTGGGAATAAATTACCTGGCCCTCACTGACCTTTTCCAGATGTCTGATAGAAGA
AAAGATGTCGTAAACTATGATATACAAGTTCAGATGGTGGAGATATACAATGAACAAATACGGGACCTTCTTACTGAAGATTCTTCAGCCACCAAATATC
CTTTTATGCTTTCAATTACTAGTTGTGCTGGGGATGATGGTCTAACTCTTCCGGATGCTACGTTGCTCCCTGTGACATCTACTTCTGATGTTCTTAATTT
GATGAAACTTGGTGAGGTGAATCGTGTTTTCAGTTCTACTGCAATGAATAATAGAAGCAGTCGTTCCCACAGTGTCCTGACAGTTCACGTGCATGGCAAA
GAAGAGTCTGGAGCTAATCTCCGCGGCTGCCTGCATTTGGTGGACTTAGCCGGAAGCGAAAGAGTTGACAAATCGGAAGTTACCGGGGATAGGCTCAAGG
AGGCAACATATATCAACAAATCTCTTTCATGCCTAGGGGACGTGGTCACAGCATTGGCACATAAGAATTCTTACATTCCATACAGAAACAGCAAACTCAC
ACTTCTGCTCCAAGATTCCTTAGGTGGACATGCTAAAACACTGATGTTTGCTCATGTGAGCCCAGAAGAAGACTTCTTATCAGAGACTATCACCACCTTG
AAGTTTGCTCAGCGGGTATCCAGCGTTGAACTTGGCGCGGCTAGTGCAAACAAAGAGAGCAGCGGGATTATGCAATTGAAAGAACAGGTTGAGAACCTCA
AAAGGGCCATGACTAACAAACAGTCATCTAATGAAGCAAGATCTCCAATTGGGAAATCAAATGCAGGTGCTGGAAGAACACCGCCACGGTCAAGGAGACT
CAGTATTGAGAACATCAGCAGCAACAATAAGCTGACCAAGACCACAAACCATGATGACAGGAAGGCATTGAAGACTCCTCCCTTGCCTACCCGTTCTCGA
AGGCTAAGTTTGGAAGCTCCAACTTACAGTAAAAGGGATGAGGATTTCTTGACAAAACAAGCAGATGACGAAACAATAAGGAAGCCCCTTCGGTTTGAAC
GGACACCAAAGCAGAACAACCATTCACAGGATGTAGAAGCCACACCAAAGAAGAACAACTATTCTGCAACGATTAGCAGCAATTCGTCCGCAAAGATTTA
CCAACGTACTCCTCCTAGAAGTCCTACTAGTTCCTCATATCAAAGGCAAACACAAATTCTTCACCTTCATTTGCCATTAGCCACAGAAAACCAAGTGGTT
GCGAGAAATGAAGTTCAGAATCTGATGCAAAGTCAAGTTGTCAGTTCCGCAACGGGCAGTGTTAATGGGAAAGGGTCTCAGATAAGGAAGTCCCTTCGGA
CCATAGGAAAGCTGATTAACGGATCTGAAAAGAGGAAACAACGGAAAGAACAGTCCCAAGCAATTTTGAAGAGTTTGAATGAAGAAATCAAGTCACCAGT
ACGGAGTAGCATCAAAAGGGCAGCAAGGAGGCAATCACTCACTGGAGTTTACAAGTCCTCAGTTGACTTAGGTGAGAGTTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10042045 pacid=23153676 polypeptide=Lus10042045 locus=Lus10042045.g ID=Lus10042045.BGIv1.0 annot-version=v1.0
MERSTTSGFYNVLASRKAEEAVWRRYQATEWLESLVGPLSIPSHPSEKEFISRLRNGLILCNAINKVHPGSVTKVVENHVPVQSLTRESQPLPAYQHFEN
VKNFLVAMGDLKLPAFEASDLERDTLELEAGSTAKVVDSILALKSYHEHKNMNGSNGAFKSTKSPMVSHSASRRNLQGLSSDSCRRLEMNVACKGHLSGN
ADSQKLEDELIKVLAEYMADTKENMGDYFLTSFRAGNVDPMKLFGRIVSSCLKDNLHKFPEGQTVVMDTLKESSNLPAHLEATPLDDISDRRNSKCCRTC
LRKGNCNHRRLLQAQERELQNLKTLLTTTRKEFEGLQFQLQSDLTQCSIQVQEMSRAAQGYHKVLKENRILYNMVQDLKGNIRVYCRVRPSFTAGASNAM
EFVGEDGSLVIVDPSKPTKDGRRHFQFNQVFGPTATQENVFEHVQPLIRSVMDGHNVCIFAYGQTGSGKTYTMSGQGGSTKELGINYLALTDLFQMSDRR
KDVVNYDIQVQMVEIYNEQIRDLLTEDSSATKYPFMLSITSCAGDDGLTLPDATLLPVTSTSDVLNLMKLGEVNRVFSSTAMNNRSSRSHSVLTVHVHGK
EESGANLRGCLHLVDLAGSERVDKSEVTGDRLKEATYINKSLSCLGDVVTALAHKNSYIPYRNSKLTLLLQDSLGGHAKTLMFAHVSPEEDFLSETITTL
KFAQRVSSVELGAASANKESSGIMQLKEQVENLKRAMTNKQSSNEARSPIGKSNAGAGRTPPRSRRLSIENISSNNKLTKTTNHDDRKALKTPPLPTRSR
RLSLEAPTYSKRDEDFLTKQADDETIRKPLRFERTPKQNNHSQDVEATPKKNNYSATISSNSSAKIYQRTPPRSPTSSSYQRQTQILHLHLPLATENQVV
ARNEVQNLMQSQVVSSATGSVNGKGSQIRKSLRTIGKLINGSEKRKQRKEQSQAILKSLNEEIKSPVRSSIKRAARRQSLTGVYKSSVDLGES
|
DESeq2's median of ratios [FLAX]
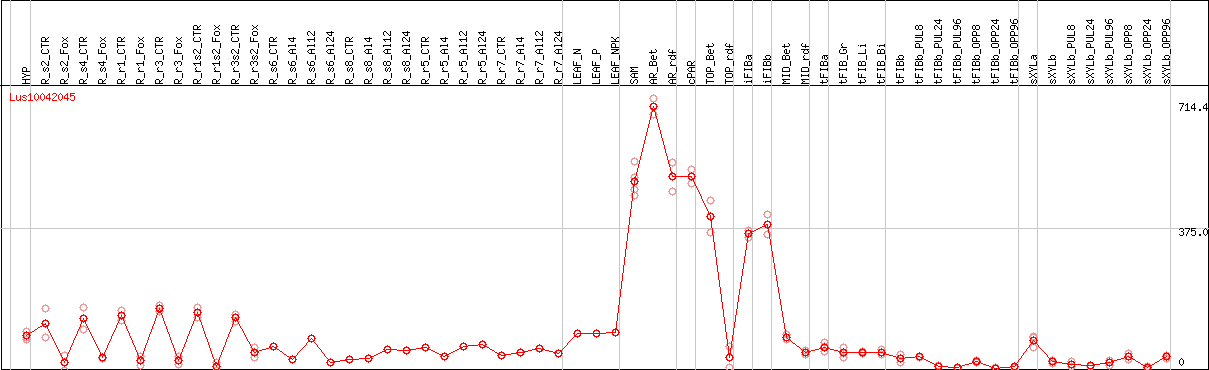
Coexpressed genes
Lus10042045 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
