External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G41660 169 / 5e-51
MIZ1
mizu-kussei 1, Protein of unknown function, DUF617 (.1)
AT3G25640 144 / 2e-41
Protein of unknown function, DUF617 (.1)
AT5G42680 134 / 3e-38
Protein of unknown function, DUF617 (.1.2)
AT5G06990 128 / 2e-35
Protein of unknown function, DUF617 (.1)
AT2G37880 123 / 1e-33
Protein of unknown function, DUF617 (.1)
AT4G39610 123 / 2e-33
Protein of unknown function, DUF617 (.1)
AT5G23100 122 / 3e-33
Protein of unknown function, DUF617 (.1)
AT1G21050 119 / 4e-32
Protein of unknown function, DUF617 (.1)
AT2G21990 118 / 8e-32
Protein of unknown function, DUF617 (.1)
AT1G76610 114 / 1e-30
Protein of unknown function, DUF617 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10018050
418 / 6e-149
AT2G41660 171 / 1e-51
mizu-kussei 1, Protein of unknown function, DUF617 (.1)
Lus10031059
182 / 1e-55
AT2G41660 258 / 4e-85
mizu-kussei 1, Protein of unknown function, DUF617 (.1)
Lus10035443
179 / 2e-54
AT2G41660 248 / 4e-81
mizu-kussei 1, Protein of unknown function, DUF617 (.1)
Lus10021759
136 / 1e-38
AT5G42680 254 / 3e-85
Protein of unknown function, DUF617 (.1.2)
Lus10000863
134 / 8e-38
AT5G23100 276 / 3e-93
Protein of unknown function, DUF617 (.1)
Lus10027241
133 / 2e-37
AT5G23100 270 / 1e-90
Protein of unknown function, DUF617 (.1)
Lus10019166
131 / 1e-36
AT3G25640 258 / 2e-86
Protein of unknown function, DUF617 (.1)
Lus10034386
128 / 3e-35
AT3G25640 268 / 3e-90
Protein of unknown function, DUF617 (.1)
Lus10036630
127 / 1e-34
AT4G39610 308 / 5e-105
Protein of unknown function, DUF617 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G048800
175 / 1e-53
AT2G41660 241 / 3e-79
mizu-kussei 1, Protein of unknown function, DUF617 (.1)
Potri.016G056800
165 / 8e-50
AT2G41660 202 / 6e-64
mizu-kussei 1, Protein of unknown function, DUF617 (.1)
Potri.001G031900
151 / 3e-44
AT5G06990 275 / 6e-93
Protein of unknown function, DUF617 (.1)
Potri.014G035000
139 / 3e-40
AT5G42680 286 / 2e-98
Protein of unknown function, DUF617 (.1.2)
Potri.002G128800
139 / 4e-40
AT5G42680 269 / 2e-91
Protein of unknown function, DUF617 (.1.2)
Potri.012G058300
139 / 5e-40
AT5G23100 261 / 7e-88
Protein of unknown function, DUF617 (.1)
Potri.015G053000
139 / 6e-40
AT5G23100 293 / 3e-100
Protein of unknown function, DUF617 (.1)
Potri.003G193700
139 / 9e-40
AT5G06990 328 / 6e-114
Protein of unknown function, DUF617 (.1)
Potri.010G131600
139 / 2e-39
AT3G25640 249 / 1e-82
Protein of unknown function, DUF617 (.1)
Potri.008G114500
133 / 2e-37
AT3G25640 228 / 2e-74
Protein of unknown function, DUF617 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF04759
DUF617
Protein of unknown function, DUF617
Representative CDS sequence
>Lus10042046 pacid=23153897 polypeptide=Lus10042046 locus=Lus10042046.g ID=Lus10042046.BGIv1.0 annot-version=v1.0
ATGTATACATACTGGCCGGCGGAGAGAGACTGCAGCCGGAGATCATCAACATCAAGCGGAAGGATCACCCCATCATCCCCATATTCAAACTACTACTACA
ATAGTACCACGCTGACTACCACCAGAACCTCCTTCTCCGACCAACAATATTATGACCACCACCGCCGCCACCTCATCCCATCTGACCAAAACAACAGGAA
AGTCAAAGTCAACATATACATGAAATCCGTCTCCTTTCTCCGCACAATATTCCTCAACTTCCTTTCTCCACCTAACTCCATCATTCAACAACAACAACAA
CAACAACAAATCATGATCCTGCCCCGCAAGAAGGTGACGACGACGTGTACTCTCTTCGGCTACCGTAGAGGACACGTCAGCTTCGCCGTCCACGTCTACG
ATCCCAAGACCACCACTTCTGACCCGATCCTCCTCCTCCAGCTCCCCATCTCCACCGCCGCCCTCCTCGACGACATGGCTTCCGGTCTCGTCCGGATCTC
CCTCGAGTGCCACACCAAGTCTGATGATGTGCTGATGAAGGAGCCAGCGTGGAGTATGTACTGCAACGGGAGGAAATGCGGCGGGAAAGCGGTGGCGCGG
TGGAGCTACACTGTTCTTGACTCGCACGTGCTAAACACCGTCCGCAGCGTCTCGGTGGGGGCCGGGGTTATTCCTCCTCCTCCTCGTTCCACGTGCGGGG
GTGGTGGTGAGTTGTTGTATATGAGGGCTAAGTTTGAGCGGGTTGTTGCGGGGAGTCGTGACTCGGAGGCTTTTCACATGTTGAACCCTGACAGTAGGAT
CTGTACCGGGTCGGGACCTGAACTCAGCCTCTTCTTCCTCCGCTTCTGA
AA sequence
>Lus10042046 pacid=23153897 polypeptide=Lus10042046 locus=Lus10042046.g ID=Lus10042046.BGIv1.0 annot-version=v1.0
MYTYWPAERDCSRRSSTSSGRITPSSPYSNYYYNSTTLTTTRTSFSDQQYYDHHRRHLIPSDQNNRKVKVNIYMKSVSFLRTIFLNFLSPPNSIIQQQQQ
QQQIMILPRKKVTTTCTLFGYRRGHVSFAVHVYDPKTTTSDPILLLQLPISTAALLDDMASGLVRISLECHTKSDDVLMKEPAWSMYCNGRKCGGKAVAR
WSYTVLDSHVLNTVRSVSVGAGVIPPPPRSTCGGGGELLYMRAKFERVVAGSRDSEAFHMLNPDSRICTGSGPELSLFFLRF
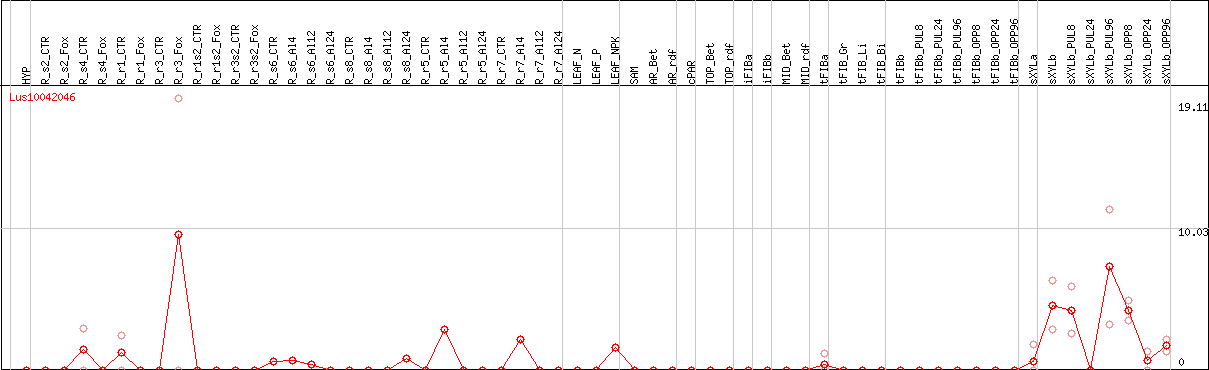
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10042046 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.