External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G24670 189 / 9e-58
TAR2
tryptophan aminotransferase related 2 (.1.2)
AT1G23320 169 / 2e-50
TAR1
tryptophan aminotransferase related 1 (.1)
AT1G70560 160 / 6e-47
CKRC1, WEI8, TAA1, sav3
WEAK ETHYLENE INSENSITIVE 8, SHADE AVOIDANCE 3, cytokinin induced root curling 1, tryptophan aminotransferase of Arabidopsis 1 (.1)
AT1G34040 146 / 2e-41
ATMEPCT
Pyridoxal phosphate (PLP)-dependent transferases superfamily protein (.1)
AT1G34060 142 / 8e-40
Pyridoxal phosphate (PLP)-dependent transferases superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10018088
299 / 1e-96
AT1G60780 632 / 0.0
HAPLESS 13, Clathrin adaptor complexes medium subunit family protein (.1)
Lus10039944
249 / 7e-81
AT4G24670 489 / 5e-172
tryptophan aminotransferase related 2 (.1.2)
Lus10027678
241 / 5e-79
AT4G24670 449 / 7e-158
tryptophan aminotransferase related 2 (.1.2)
Lus10006199
169 / 2e-50
AT1G70560 405 / 6e-141
WEAK ETHYLENE INSENSITIVE 8, SHADE AVOIDANCE 3, cytokinin induced root curling 1, tryptophan aminotransferase of Arabidopsis 1 (.1)
Lus10036846
162 / 1e-47
AT1G70560 415 / 4e-144
WEAK ETHYLENE INSENSITIVE 8, SHADE AVOIDANCE 3, cytokinin induced root curling 1, tryptophan aminotransferase of Arabidopsis 1 (.1)
Lus10028695
140 / 5e-39
AT1G34060 474 / 4e-165
Pyridoxal phosphate (PLP)-dependent transferases superfamily protein (.1)
Lus10007085
138 / 5e-38
AT1G34060 456 / 5e-158
Pyridoxal phosphate (PLP)-dependent transferases superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.012G083300
202 / 9e-63
AT4G24670 489 / 1e-171
tryptophan aminotransferase related 2 (.1.2)
Potri.T125108
202 / 1e-62
AT4G24670 487 / 7e-171
tryptophan aminotransferase related 2 (.1.2)
Potri.010G044500
164 / 9e-49
AT1G70560 504 / 4e-179
WEAK ETHYLENE INSENSITIVE 8, SHADE AVOIDANCE 3, cytokinin induced root curling 1, tryptophan aminotransferase of Arabidopsis 1 (.1)
Potri.008G187800
164 / 1e-48
AT1G70560 477 / 1e-168
WEAK ETHYLENE INSENSITIVE 8, SHADE AVOIDANCE 3, cytokinin induced root curling 1, tryptophan aminotransferase of Arabidopsis 1 (.1)
Potri.002G063800
139 / 2e-38
AT1G34060 472 / 1e-164
Pyridoxal phosphate (PLP)-dependent transferases superfamily protein (.1)
Potri.002G064000
135 / 4e-37
AT1G34060 514 / 0.0
Pyridoxal phosphate (PLP)-dependent transferases superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0061
PLP_aminotran
PF04864
Alliinase_C
Allinase
Representative CDS sequence
>Lus10042084 pacid=23154020 polypeptide=Lus10042084 locus=Lus10042084.g ID=Lus10042084.BGIv1.0 annot-version=v1.0
ATGGGCCGGCGACGCCTCCAAATTCCAAGCGCACGGGCCCTTCGTGGAATTCGTAACTTCGCCCAACAACCCAGACGGGTTTACTCGACACCCGGTCGTG
AACCGAACGGAGGGTTTCTGGTTCACGATCTCGCTTACTACGGGCCGCAATACACCCCCGTGACTTCCCCTTCCGATCACGACATCACCCTTTTCACCGT
CTCCAAATCCACCGGCCACGCTGGCATCCGGATCGGATGGGCGCTGGTGAAGGATCCTGAGGTGGCGAGGAAAATGACGAAATTCATAGAGCTGAGCAGC
CTGGGGGTGTCCAAGGACTCGCAGCTGAGAGCAACCAAGATAATGGAGGTCGTTTCGGCGGATGCTGAGGAGGAATCCAAGCGGGTCTCATCGCTGTTCG
AGTTCGGGTTGAAGAACATGAAACGGCGGTGGAAGATGTTGAGAGCGGCGGTGAAAAAGAGCGGAGGAATGTTCAGCTTGCCGGATTACAGATCGGAGTT
CTGCAGTTTCAACGGCCAGTCCATGTCAAACCAACCCGCGTTTGCGTGGGTGAAGTGCGAGGGAGGAGACGTTGAAGACTGTGAAGAGGCGTTCAAGAAG
AATAAGATTGAGTCACTTTTAATATCTACATGGGATAATCATGCATTGTGGTTGATAAATCAGTACTACGTGTATTGTTTACTTTTAGTAAAATTTGGTG
ACTTACTGGTTAATTGA
AA sequence
>Lus10042084 pacid=23154020 polypeptide=Lus10042084 locus=Lus10042084.g ID=Lus10042084.BGIv1.0 annot-version=v1.0
MGRRRLQIPSARALRGIRNFAQQPRRVYSTPGREPNGGFLVHDLAYYGPQYTPVTSPSDHDITLFTVSKSTGHAGIRIGWALVKDPEVARKMTKFIELSS
LGVSKDSQLRATKIMEVVSADAEEESKRVSSLFEFGLKNMKRRWKMLRAAVKKSGGMFSLPDYRSEFCSFNGQSMSNQPAFAWVKCEGGDVEDCEEAFKK
NKIESLLISTWDNHALWLINQYYVYCLLLVKFGDLLVN
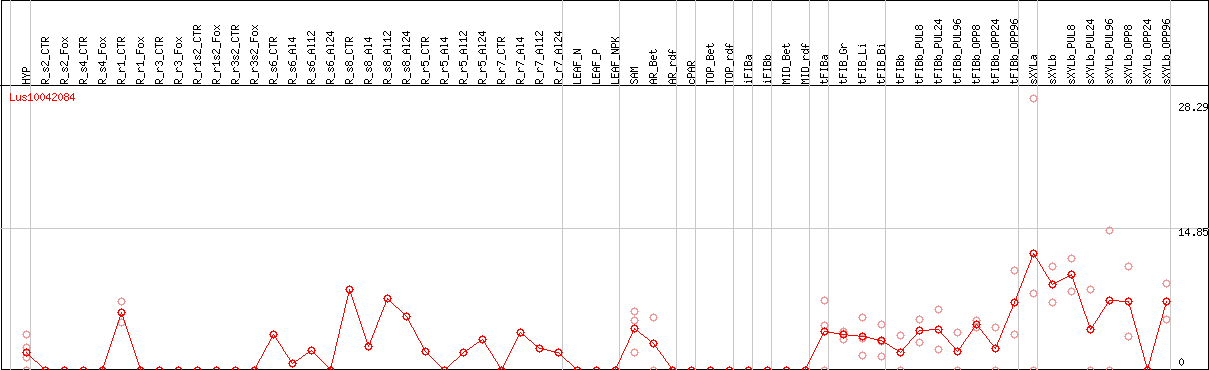
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10042084 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.