External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G01510 183 / 8e-55
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G53360 169 / 8e-49
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G29230 167 / 2e-48
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G28690 164 / 4e-48
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT1G77170 163 / 4e-48
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT5G66520 165 / 7e-48
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT2G36730 163 / 9e-48
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT4G33990 166 / 1e-47
EMB2758
embryo defective 2758, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G21065 160 / 4e-47
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
AT3G49142 164 / 5e-47
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10008618
360 / 8e-123
AT4G33990 367 / 4e-117
embryo defective 2758, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10002024
175 / 3e-52
AT5G56310 344 / 6e-113
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10013319
167 / 8e-49
AT4G21065 744 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Lus10022890
169 / 1e-48
AT3G61170 865 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10005207
165 / 1e-47
AT4G21065 741 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Lus10030124
163 / 1e-47
AT3G15130 613 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10024876
164 / 1e-46
AT3G12770 886 / 0.0
mitochondrial editing factor 22 (.1)
Lus10000706
163 / 2e-46
AT3G12770 878 / 0.0
mitochondrial editing factor 22 (.1)
Lus10029344
161 / 2e-46
AT5G06540 803 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G252400
181 / 1e-54
AT4G21065 496 / 8e-172
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Potri.001G258500
181 / 5e-53
AT2G40720 481 / 4e-157
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.017G001000
176 / 5e-52
AT4G21065 793 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Potri.013G103600
173 / 7e-51
AT1G08070 477 / 7e-160
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.016G019900
169 / 9e-49
AT4G02750 542 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.003G006800
169 / 1e-48
AT3G16610 687 / 0.0
pentatricopeptide (PPR) repeat-containing protein (.1)
Potri.013G044700
169 / 1e-48
AT3G22690 582 / 0.0
unknown protein
Potri.005G019400
165 / 1e-48
AT1G09190 574 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.005G005700
167 / 3e-48
AT3G49142 912 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.004G208000
166 / 4e-48
AT2G03880 939 / 0.0
required for efficiency of mitochondrial editing 1, Pentatricopeptide repeat (PPR) superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF01535
PPR
PPR repeat
Representative CDS sequence
>Lus10042198 pacid=23153537 polypeptide=Lus10042198 locus=Lus10042198.g ID=Lus10042198.BGIv1.0 annot-version=v1.0
ATGTACGCAAAATGTGGGGATATGGATCGAGCTCGATCGGTCTTCGATGAGATACTGTGTAAGGATGTAATATCTTGGACAACTATGATTCTAGGTCATG
CCATCAATGGTGAAGGAGAGGAAGCCCTCCTTGCTTTCCACGAAATGCGTGAGGAAGGAATCAGGCCGAATTTCGTGACATTCATCGGGGTTCTGTCGGC
ATTTAATCATTCCGGTCTGGTAGAAGCAGGCCAGAAGTTATACGAGATGATGATCCAGGCTGACGATATTGAGCCAAAGATGGAGCATTACGGATGTATG
ATCGATATGCTTGCCCGAGCAGGGATGCTCGAGGAGGCAAAGAAGTTCGTGGAGAGAATGCCGGTGGAGCCAAATGCAGCTGTTTGGAGGATGCTGGTTA
ATGCTTGCAGAGGTCACAGTGCTACTAAACTCGGATTGAGCTCAGTGGCTAACCTCTTTCCAGGGAGGAATTCAACTGTGGTTGCTGAAAATCGGATTAT
TTGTGCTAACATGTTTGCTGAGGCAGAGAGATGGAATGAAGTTTTGCAAGATCGACCTATTATGGTTGCCCAGAAGGATCGTAAAGTAGCTGGCAAAAGC
TCCACTGTAACGTAA
AA sequence
>Lus10042198 pacid=23153537 polypeptide=Lus10042198 locus=Lus10042198.g ID=Lus10042198.BGIv1.0 annot-version=v1.0
MYAKCGDMDRARSVFDEILCKDVISWTTMILGHAINGEGEEALLAFHEMREEGIRPNFVTFIGVLSAFNHSGLVEAGQKLYEMMIQADDIEPKMEHYGCM
IDMLARAGMLEEAKKFVERMPVEPNAAVWRMLVNACRGHSATKLGLSSVANLFPGRNSTVVAENRIICANMFAEAERWNEVLQDRPIMVAQKDRKVAGKS
STVT
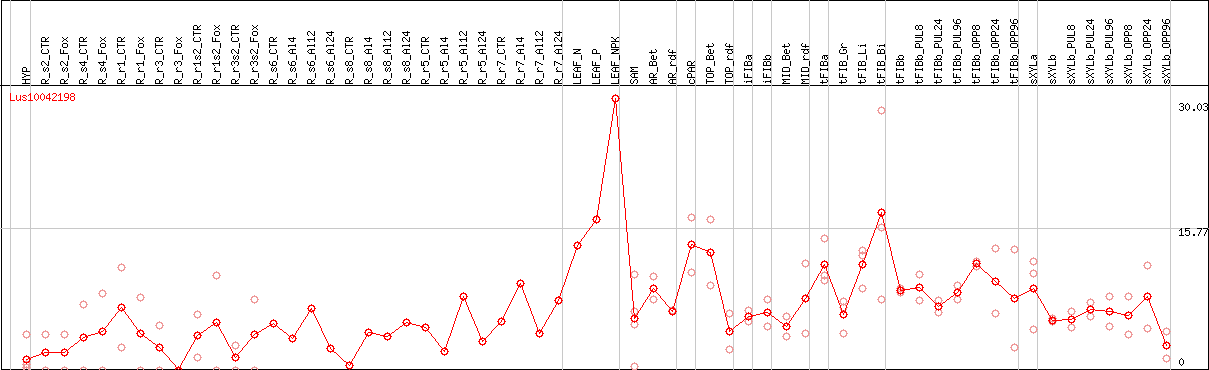
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10042198 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.