Lus10042297 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10042297 pacid=23153870 polypeptide=Lus10042297 locus=Lus10042297.g ID=Lus10042297.BGIv1.0 annot-version=v1.0
ATGGCGGACACAAATTCGGTTGACGTGATCCTTGACTTCTTGCGGAGAAACCGGTTCACTAGAGCTGAGGCAGCGTTAAGGAGCGAGCTTGGCGTTCAAC
CTGACCTAAATGGGTTCTTGCAGAAACTTAGTCTTGATGACAAGGAAGCGAGGGAAGTCATAGAGGAAAGGAATGTGAGTAACAAACCAGCTGCTAAATT
TCAGGGCTCTGGTTCCCGAGATAGTTGTGAGGATTCTAAAGAACTTATAGTGAAAGAGATAGACTGCAGCACTCACGGAAATGGGTCTGAGGGTAGATGG
AAAAGTTCTGCTTCTGCTGACGACCGTAATGTCAAACCTAATGAAACAATCGATGGCACAGACTCTGGGGATACCGTCTTAGATTTGTATTCGTGGAAGT
CCATAGCTAACAACGGTCCTAATGCTTATCAGAATGATAGAGCTAATACAAGCAACTACTCTGGTAGAACAGAAGGGAAATCTAGAGAGGAGGCTACCTT
CTCAGGTGTAAAGAAAACTGCATGGCAGGAAAGCAGCAGTGGTTCATCCATCCCTGATTCTAGAAATAGCAAAATTCAGACAAGTGATCCGATAGAGTTT
GGTCAGCAAGTTAAAACAACTGTCTCTTATCCTGCTGACAACCCATGGTCAGTAACTGAGCAGCCAACAAGTTCTTCTGCGGACATATGGAAAAATTGTT
CTGTGAAGACTGTTCTTCCGTTCTCCATTCTGGATGGATCTACTAGTTATGACACGTCATCAGATAAGAAGGAAGGGAATAAGAAAGCAGATGTGCCTGA
TGTAAGGGCTGCCATTAAAGAAAAGGTAGATGAAGTTGGCAGAGCTTTGTACTTTGGGAACTCACAAGGAAGTATTGAACCGAAGAATTTAAGCTCTTCA
GGGTTTACTATTGCGGCTGAGAATCATAAAGAAGACTATCCGAGACTTCCGCCCGTTAGACTAAAGTCAGAAGACAAATTATTGAATGCCAACTGGCAGG
AGAAGTTTGAGCGTGATAAGCAAGGGGCAAAGCTTAGCAGTGGTGATAGTTCCTTTCTGATAGGGTCATATCTTGACGTTCCTATTGGACAGGAAATCAA
CTCTTCAGTTGACGATTCAGCTGGAAAAAGAATAGCCGGAGGCAGCTGGCTGTCCGTGAGTCAGGGAATTGCAGAAGACGACCTAGTTTCTGGTTTTGCT
ACAGTTGGTGATGGATTGAGTGAGTCTATTGACTATCCTAATCAATATTGGGATTCTGATGAATATGATGATGATGACGATGTCGGGTACACGAGGCAAC
CTATAGAGGATGAGGCCTGGTTTCTGGCTCATGAAATTGATTATCCTAGTGACAACGAGAAGGGGACAGTAACTGGGAGCGTTCCAGACCCTCAGGAAAG
GGGTCCAACAAAAGACGAGGATGATGATCAATCTTTTGCTGAGGAAGACTCGTACTTTTCTGGGGAGCGGTCTTTCCAAACAAAAAATGTTGAACCTGCT
ACAGCTTCAGATGATCCTATTGGACTGTCAGTGACTGAAATTTACGGGAGAACACATGATAATGACTTGATGGCACAGTATGATGGCCAATTGTTAGATG
AAGAAGAACTAAATTTGATGTGTGGTGAACCTGTGTGGCAGGGGTTTGTCACGCAAACAAATGAACTAGTCATGCTAGGAGACGGGAAAGTAAACAATGA
CTCCGGGAGGACACGAATGGATGATGATCAACATGGCTCGGTTCGATCAATTGGTGTGGGTATTAACAGCGATGCTGCTGATTTTGGGAGTGAAATACAT
GATAGTTTGATTGGTGGTAGCAGTGAAGGAGATCTGGAATACTTTCGTGATCATGATGTCGAGGTTAGAGGGTATAGAGCTAGTCACCAAGATTTAGACA
AGAAATATGGTGATAAACAAAGCAAGGATAAAAAGAGGGCCGGCAAACATGATGCAGGGAATTTTGGGAATGAAAAAGTTGCTTCTACTCAACTGAAAGC
TCTTAAAGATGGAGGGTTTTCATTTCCTCCACCATTGAGGGATAGTCAAGAGGTCCAGGCAGGGTCTAGCAAGTCTTTGTGGTCTAACAAGTCTAGCACA
GTTTTGGCTAATGAAACTGGTGACATGGATCATTTGGATGCTTTGACGTTGCCTGATGACATGCTTGGTGGATTGAAGCTTAAAAGTAGTAGTTCTTCAA
CTGCGAAAAGTTACATGGAAGAAAATATTGCCCATGCTATTGTTTCTGGAGACTCAACTCCCTCTACGCTTTCAAACTATGCATACAACGAAAAAGAACG
TGCTAAGAAAGAGGAGGATGATAAGATTGACGACGCTGGGGAAGAACCTGGGATGTCACTTGATGATGAAGAAGCAATTGCTGTGCAAGAACAAGTGAGG
CAGATCAAAGCTCAGGAGGAAGAATTTGAGACTTTCAACCTAAAGATTGTGCATCGGAAAAACAGGCATAAATGTTTCGCCTTCAGTATGGGAATTGGAT
ACAAAATCTCTATCGATGCTGCTTCTTCATCAGCTGGGACTGGCTTTGAGGAGGATAAGAACTTCCATGTTGTGTTAAATTCTGTGTTAGCTGGGCGATA
TCATGTTACAGAGTATCTTGGCTCAGCTGCTTTCAGTAAAGCCATTCAGGCGCATGATCTGCACACAGGCATGGATGTCTGTGTGAAAATCATAAAAAAC
AACAAGGATTTCTTTGACCAGAGCCTTGATGAGATAAAACTTCTCAAATACGTGAACAAGCATGATCCAGCTGACAAGTATCACCTTCTCAGATTATACG
ACTATTTCTACTACCGGGAACATTTGTTGATTGTGTGTGAACTTCTCAAAGCAAACTTATATGAATTCCATAAATTCAACAGAGAATCTGGAGGAGAAGT
GTATTTCACAATGCCCAGATTGCAGTCAATTACAATTCAGTGTTTGGAAGCACTTCAATTCTTGCATGGTCTCGGCCTTATTCACTGCGATTTGAAGCCT
GAAAATATACTAGTGAAAAGCTACAGTCGGTGCGAGGTGAAAGTCATCGATCTCGGAAGTAGCTGCTTCGAGACGGATCACCTCTGCTCTTACGTCCAAT
CTAGATCATATCGGGCACCCGAGGTCATCCTCGGCCTTCCATATGATAAGAAGATCGACGTTTGGTCACTCGGCTGTATCTTGGCAGAGCTTTGTACTGG
CAATGTCCTCTTCCAAAATGATTCACCAGCAACCCTTCTGGCCAGAGTCATCGGAATCATTGGGGCCGTTGAACAGGAAATGTTGGCTAAAGGACGAGAC
ACATACAAGTATTTCACCAAAAATCACATGCTTTACGAACGGAATCAGGAGACAAACCGGCTAGAGTATTTGGTACCGAAGAAGACATCATTGAGGCACC
GATTGCCGATGGGGGACCACGGATTCATTGATTTCGTCAGGCATTTGCTCGAAGTGAACCCCAACAAGCGGCCTTCTGCGGCAGAGGCTCTGAAGCACCC
TTGGCTATCCTTCCCTTACGAACCAATATCTGCGTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10042297 pacid=23153870 polypeptide=Lus10042297 locus=Lus10042297.g ID=Lus10042297.BGIv1.0 annot-version=v1.0
MADTNSVDVILDFLRRNRFTRAEAALRSELGVQPDLNGFLQKLSLDDKEAREVIEERNVSNKPAAKFQGSGSRDSCEDSKELIVKEIDCSTHGNGSEGRW
KSSASADDRNVKPNETIDGTDSGDTVLDLYSWKSIANNGPNAYQNDRANTSNYSGRTEGKSREEATFSGVKKTAWQESSSGSSIPDSRNSKIQTSDPIEF
GQQVKTTVSYPADNPWSVTEQPTSSSADIWKNCSVKTVLPFSILDGSTSYDTSSDKKEGNKKADVPDVRAAIKEKVDEVGRALYFGNSQGSIEPKNLSSS
GFTIAAENHKEDYPRLPPVRLKSEDKLLNANWQEKFERDKQGAKLSSGDSSFLIGSYLDVPIGQEINSSVDDSAGKRIAGGSWLSVSQGIAEDDLVSGFA
TVGDGLSESIDYPNQYWDSDEYDDDDDVGYTRQPIEDEAWFLAHEIDYPSDNEKGTVTGSVPDPQERGPTKDEDDDQSFAEEDSYFSGERSFQTKNVEPA
TASDDPIGLSVTEIYGRTHDNDLMAQYDGQLLDEEELNLMCGEPVWQGFVTQTNELVMLGDGKVNNDSGRTRMDDDQHGSVRSIGVGINSDAADFGSEIH
DSLIGGSSEGDLEYFRDHDVEVRGYRASHQDLDKKYGDKQSKDKKRAGKHDAGNFGNEKVASTQLKALKDGGFSFPPPLRDSQEVQAGSSKSLWSNKSST
VLANETGDMDHLDALTLPDDMLGGLKLKSSSSSTAKSYMEENIAHAIVSGDSTPSTLSNYAYNEKERAKKEEDDKIDDAGEEPGMSLDDEEAIAVQEQVR
QIKAQEEEFETFNLKIVHRKNRHKCFAFSMGIGYKISIDAASSSAGTGFEEDKNFHVVLNSVLAGRYHVTEYLGSAAFSKAIQAHDLHTGMDVCVKIIKN
NKDFFDQSLDEIKLLKYVNKHDPADKYHLLRLYDYFYYREHLLIVCELLKANLYEFHKFNRESGGEVYFTMPRLQSITIQCLEALQFLHGLGLIHCDLKP
ENILVKSYSRCEVKVIDLGSSCFETDHLCSYVQSRSYRAPEVILGLPYDKKIDVWSLGCILAELCTGNVLFQNDSPATLLARVIGIIGAVEQEMLAKGRD
TYKYFTKNHMLYERNQETNRLEYLVPKKTSLRHRLPMGDHGFIDFVRHLLEVNPNKRPSAAEALKHPWLSFPYEPISA
|
DESeq2's median of ratios [FLAX]
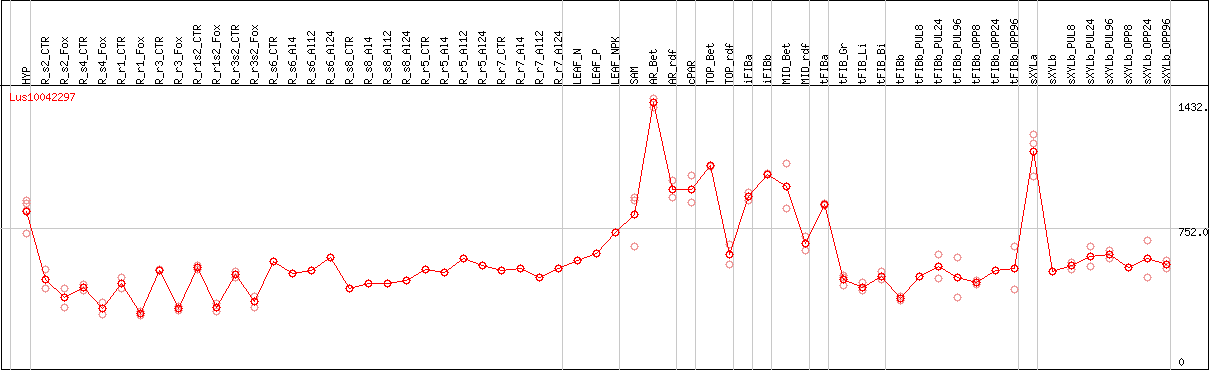
Coexpressed genes
Lus10042297 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
