External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G21620 177 / 3e-56
RD2
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
AT3G53990 61 / 9e-12
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
AT3G62550 61 / 1e-11
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
AT3G58450 54 / 7e-09
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
AT3G11930 54 / 8e-09
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2.3.4)
AT3G17020 52 / 2e-08
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
AT5G14680 52 / 3e-08
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
AT1G68300 49 / 4e-07
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
AT3G03270 47 / 1e-06
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
AT2G47710 46 / 2e-06
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10026346
390 / 2e-137
AT2G21620 269 / 5e-90
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
Lus10042316
184 / 6e-59
AT2G21620 284 / 3e-99
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
Lus10022602
56 / 1e-09
AT3G53990 225 / 2e-76
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
Lus10021501
54 / 2e-09
AT3G53990 144 / 3e-45
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
Lus10006701
53 / 1e-08
AT2G47710 218 / 1e-73
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
Lus10021104
53 / 1e-08
AT3G53990 215 / 2e-72
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
Lus10014545
53 / 1e-08
AT5G14680 295 / 6e-104
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
Lus10017207
52 / 2e-08
AT3G53990 217 / 2e-73
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
Lus10032142
52 / 2e-08
AT5G14680 296 / 3e-104
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.004G156100
202 / 5e-66
AT2G21620 241 / 6e-82
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
Potri.004G156200
179 / 3e-57
AT2G21620 256 / 3e-88
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
Potri.009G117500
175 / 1e-55
AT2G21620 258 / 5e-89
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
Potri.006G092700
66 / 1e-13
AT3G53990 207 / 1e-69
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
Potri.014G122000
65 / 5e-13
AT3G62550 195 / 1e-64
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
Potri.002G196700
62 / 6e-12
AT3G62550 196 / 6e-65
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
Potri.016G104600
61 / 9e-12
AT3G53990 229 / 6e-78
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
Potri.006G198200
61 / 2e-11
AT3G11930 209 / 9e-69
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2.3.4)
Potri.005G015200
60 / 3e-11
AT3G62550 156 / 4e-49
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
Potri.016G064000
59 / 2e-10
AT3G11930 214 / 4e-71
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2.3.4)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0039
HUP
PF00582
Usp
Universal stress protein family
Representative CDS sequence
>Lus10042317 pacid=23153479 polypeptide=Lus10042317 locus=Lus10042317.g ID=Lus10042317.BGIv1.0 annot-version=v1.0
ATGGAGGATCCACACTGTAAGAAAAGAGAGAAGCAAACAGCGAAAACACTGGAAACAGTGGAGGAAGATGAGGTGTATGATCATGGTTGGAATGATGAAG
AAGAAGAACAAAGGAAGAGAATATCAGAGCTGGACATGGAAACAGCACAAGGACGAAGAAGAAGAAAAATAGTCAGTGGCGGAGAAGGAGGAAGACACAT
AATTGTAGGGATCGATCATGGACCAAACAGCAAGCATGCTTTCAACTGGGCTCTGCTTCACCTCTGCCGCCAGGCTGACACCCTTCACCTTGTTCATGCC
CTTTCTGATGCAAAGAACAAGCTGCTTTCTGACATGGCTGAAGGGTTGGTGGAGAAGTTTACACTTGAGGCTTTGAGGATGGCTAAAGTGAAGGCAGTGG
GGAGAGTAGTGGAAGGAGAGGCAAGTAAGGTTATTTGCAAGGAAGCAGAGAGGCTAAAACCTGTGGCAGTTGTACTAGGAACCAGAGGGAGATCTCTTTT
CCAAAGTGTGGTGCAAGGAAGTGTCGGGGAGTATTGTTTCCACCACATCAAAGTTCCAGTCGTAATTGTTCCTCTGCAATGTCAGTGTCTAACTACACAA
CCTTTCCTTTATGTTTTCCTCTTTCGGATCTCTGACACTTTCCATGGAAATGACTGTATATGA
AA sequence
>Lus10042317 pacid=23153479 polypeptide=Lus10042317 locus=Lus10042317.g ID=Lus10042317.BGIv1.0 annot-version=v1.0
MEDPHCKKREKQTAKTLETVEEDEVYDHGWNDEEEEQRKRISELDMETAQGRRRRKIVSGGEGGRHIIVGIDHGPNSKHAFNWALLHLCRQADTLHLVHA
LSDAKNKLLSDMAEGLVEKFTLEALRMAKVKAVGRVVEGEASKVICKEAERLKPVAVVLGTRGRSLFQSVVQGSVGEYCFHHIKVPVVIVPLQCQCLTTQ
PFLYVFLFRISDTFHGNDCI
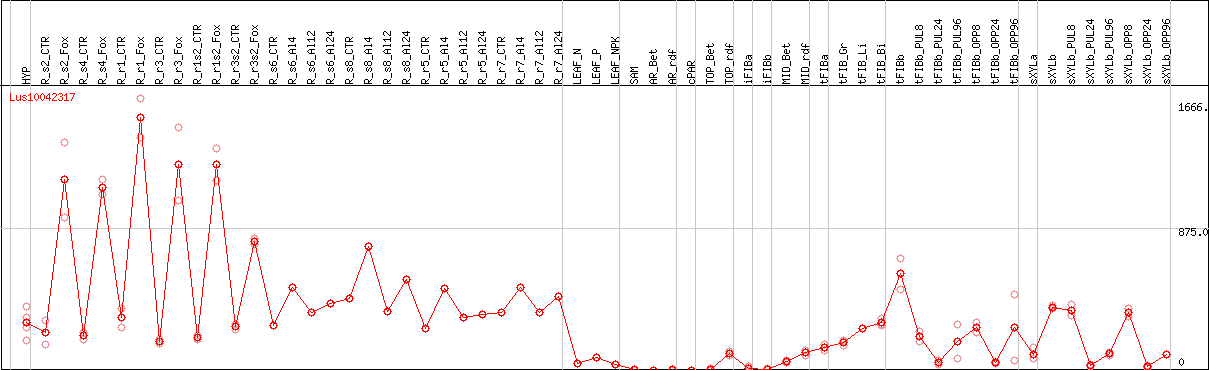
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10042317 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.