External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G18870 276 / 2e-92
Mitochondrial transcription termination factor family protein (.1)
AT2G34620 144 / 1e-40
Mitochondrial transcription termination factor family protein (.1)
AT2G03050 136 / 4e-38
SOLDAT10, EMB93
SINGLET OXYGEN-LINKED DEATH ACTIVATOR 10, EMBRYO DEFECTIVE 93, Mitochondrial transcription termination factor family protein (.1)
AT2G36000 114 / 3e-29
EMB3114
EMBRYO DEFECTIVE 3114, Mitochondrial transcription termination factor family protein (.1.2)
AT5G55580 87 / 6e-19
Mitochondrial transcription termination factor family protein (.1)
AT1G78930 86 / 2e-18
Mitochondrial transcription termination factor family protein (.1)
AT2G21710 81 / 9e-17
EMB2219
embryo defective 2219, Mitochondrial transcription termination factor family protein (.1)
AT4G02990 78 / 7e-16
RUG2, BSM
RUGOSA 2, BELAYA SMERT, Mitochondrial transcription termination factor family protein (.1)
AT4G14605 76 / 3e-15
Mitochondrial transcription termination factor family protein (.1)
AT2G44020 74 / 1e-14
Mitochondrial transcription termination factor family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10026325
595 / 0
AT3G18870 278 / 3e-93
Mitochondrial transcription termination factor family protein (.1)
Lus10014567
146 / 4e-41
AT2G34620 300 / 6e-101
Mitochondrial transcription termination factor family protein (.1)
Lus10032120
137 / 4e-38
AT2G34620 300 / 2e-101
Mitochondrial transcription termination factor family protein (.1)
Lus10029422
129 / 2e-34
AT2G36000 294 / 3e-98
EMBRYO DEFECTIVE 3114, Mitochondrial transcription termination factor family protein (.1.2)
Lus10004219
123 / 2e-32
AT2G36000 293 / 8e-98
EMBRYO DEFECTIVE 3114, Mitochondrial transcription termination factor family protein (.1.2)
Lus10016971
119 / 4e-31
AT2G36000 271 / 2e-89
EMBRYO DEFECTIVE 3114, Mitochondrial transcription termination factor family protein (.1.2)
Lus10021297
117 / 2e-30
AT2G36000 264 / 8e-87
EMBRYO DEFECTIVE 3114, Mitochondrial transcription termination factor family protein (.1.2)
Lus10030451
113 / 5e-29
AT2G03050 283 / 3e-95
SINGLET OXYGEN-LINKED DEATH ACTIVATOR 10, EMBRYO DEFECTIVE 93, Mitochondrial transcription termination factor family protein (.1)
Lus10026612
109 / 2e-27
AT2G03050 285 / 4e-96
SINGLET OXYGEN-LINKED DEATH ACTIVATOR 10, EMBRYO DEFECTIVE 93, Mitochondrial transcription termination factor family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.004G150600
309 / 2e-105
AT3G18870 311 / 1e-106
Mitochondrial transcription termination factor family protein (.1)
Potri.011G081400
159 / 1e-46
AT2G34620 392 / 5e-138
Mitochondrial transcription termination factor family protein (.1)
Potri.006G205000
136 / 2e-37
AT2G36000 284 / 2e-94
EMBRYO DEFECTIVE 3114, Mitochondrial transcription termination factor family protein (.1.2)
Potri.016G072200
136 / 2e-37
AT2G36000 280 / 1e-92
EMBRYO DEFECTIVE 3114, Mitochondrial transcription termination factor family protein (.1.2)
Potri.010G167400
114 / 1e-29
AT2G03050 303 / 3e-103
SINGLET OXYGEN-LINKED DEATH ACTIVATOR 10, EMBRYO DEFECTIVE 93, Mitochondrial transcription termination factor family protein (.1)
Potri.001G361800
92 / 2e-20
AT5G55580 584 / 0.0
Mitochondrial transcription termination factor family protein (.1)
Potri.007G001800
90 / 7e-20
AT1G78930 633 / 0.0
Mitochondrial transcription termination factor family protein (.1)
Potri.004G209400
84 / 4e-18
AT4G38160 481 / 3e-172
pigment defective 191, Mitochondrial transcription termination factor family protein (.1.2.3)
Potri.009G170300
80 / 1e-16
AT4G38160 479 / 8e-170
pigment defective 191, Mitochondrial transcription termination factor family protein (.1.2.3)
Potri.017G067600
77 / 1e-15
AT4G14605 571 / 0.0
Mitochondrial transcription termination factor family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF02536
mTERF
mTERF
Representative CDS sequence
>Lus10042342 pacid=23153782 polypeptide=Lus10042342 locus=Lus10042342.g ID=Lus10042342.BGIv1.0 annot-version=v1.0
ATGCTACACTCACTAACTATCCAAGCTTCCTCCATAACCAGAACCTTCACCAAACACCCATCAATACCTACTCCATCTCTACACCCTCCCCTTCACATCA
ACTTCCAAACTACCCGCCTAGAACATCTCCGATACCTCAAATCCGTCGGCGTGGTTCGCCCCGACACCAAACTCCACAAGCTCCCCTCCCCGGACGCCAT
CTCCCACATCCTCTCCACCGTAAACTTCTTCAAATCCAAAGGCTTCTCCAACCCCGACTTCCCCAGGCTCGTCTCCGTTTGCCCCGAAATCTTCTCCCCC
AACTTCGATGTCTCCGACATTCAACCCGTGTTCAAGTTCCTCGCCATGGATTTGGGTGCCTTGCCCCACCAATCTTGCGGCCTGATCGCCAATTGCCCCG
AGATCCTCCTCACTGATGTGGAGTACTGCCTGAGGCCGACGCTGGACTATCTCAGGCATATTGGGGTTGAGAATTTGAACGCTCCTACAATCCTCAACGC
TAACCTTTTGAACGTTCGAATCAGGAAGTTGCAGTCGAAGGTGATGTTTCTGAGGAGCATTGGGTTTTCGGAGGAGGAAGCTGAGAAATGCTGTGCGAGG
TTGCCTGCCATATTGAAGTACAGCGTTGAGAATAATATGAAGAGGAAGTACGTGTACCTGGTGAAGGAGATGGGGCGGAGCGTGGAGGAGTTGAAGGAGT
TTCCTCAGTATTTCGGGTTTAGCCTCGAAACGAGGATACACCCGAGGCATCTGCATCTGAAAGCCATGAATGTTAAGGCGAAGCTGGATAGAATGTTGAA
ATGTAGTGACGAGAATTTCTACTACCGTTTTTGGGAGAAACCTGGCCTTGCTGGGCGCAACAAATCCAGACATTCTAAGAGGATGAATCCGAGAGGAGAA
AAACGCATTTCGACCAATGCTCCGAGTCTAACACTCTGA
AA sequence
>Lus10042342 pacid=23153782 polypeptide=Lus10042342 locus=Lus10042342.g ID=Lus10042342.BGIv1.0 annot-version=v1.0
MLHSLTIQASSITRTFTKHPSIPTPSLHPPLHINFQTTRLEHLRYLKSVGVVRPDTKLHKLPSPDAISHILSTVNFFKSKGFSNPDFPRLVSVCPEIFSP
NFDVSDIQPVFKFLAMDLGALPHQSCGLIANCPEILLTDVEYCLRPTLDYLRHIGVENLNAPTILNANLLNVRIRKLQSKVMFLRSIGFSEEEAEKCCAR
LPAILKYSVENNMKRKYVYLVKEMGRSVEELKEFPQYFGFSLETRIHPRHLHLKAMNVKAKLDRMLKCSDENFYYRFWEKPGLAGRNKSRHSKRMNPRGE
KRISTNAPSLTL
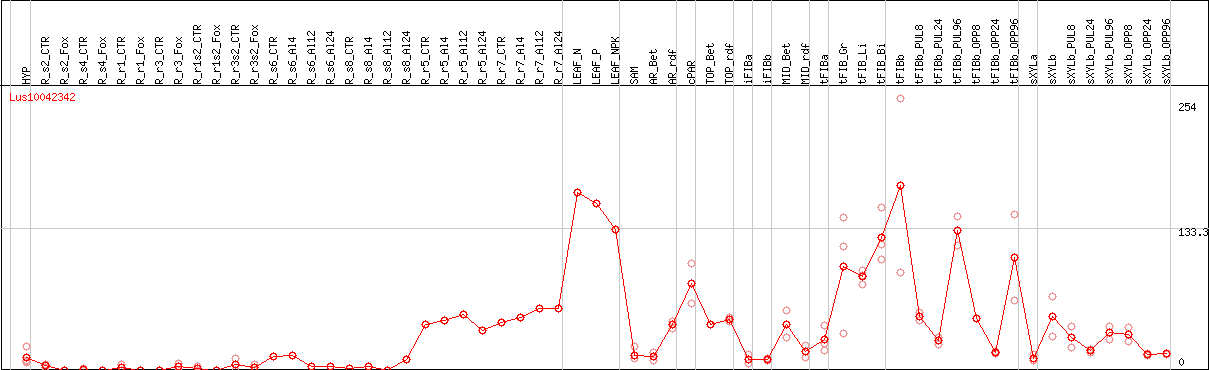
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10042342 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.