External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G44410 134 / 6e-38
FAD-binding Berberine family protein (.1)
AT4G20860 130 / 2e-36
FAD-binding Berberine family protein (.1)
AT5G44360 127 / 3e-35
FAD-binding Berberine family protein (.1)
AT5G44400 125 / 1e-34
FAD-binding Berberine family protein (.1)
AT2G34790 124 / 3e-34
MEE23, EDA28
MATERNAL EFFECT EMBRYO ARREST 23, EMBRYO SAC DEVELOPMENT ARREST 28, FAD-binding Berberine family protein (.1)
AT5G44380 123 / 5e-34
FAD-binding Berberine family protein (.1)
AT5G44440 122 / 1e-33
FAD-binding Berberine family protein (.1)
AT5G44390 122 / 2e-33
FAD-binding Berberine family protein (.1)
AT1G30760 121 / 4e-33
FAD-binding Berberine family protein (.1)
AT4G20820 117 / 8e-32
FAD-binding Berberine family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10026290
242 / 4e-79
AT5G44390 473 / 1e-162
FAD-binding Berberine family protein (.1)
Lus10038446
159 / 5e-47
AT4G20860 520 / 0.0
FAD-binding Berberine family protein (.1)
Lus10038445
139 / 8e-40
AT1G30760 415 / 6e-141
FAD-binding Berberine family protein (.1)
Lus10023361
138 / 4e-39
AT2G34790 674 / 0.0
MATERNAL EFFECT EMBRYO ARREST 23, EMBRYO SAC DEVELOPMENT ARREST 28, FAD-binding Berberine family protein (.1)
Lus10018119
134 / 6e-38
AT2G34790 641 / 0.0
MATERNAL EFFECT EMBRYO ARREST 23, EMBRYO SAC DEVELOPMENT ARREST 28, FAD-binding Berberine family protein (.1)
Lus10036147
133 / 2e-37
AT2G34790 626 / 0.0
MATERNAL EFFECT EMBRYO ARREST 23, EMBRYO SAC DEVELOPMENT ARREST 28, FAD-binding Berberine family protein (.1)
Lus10038436
132 / 5e-37
AT4G20820 508 / 1e-176
FAD-binding Berberine family protein (.1)
Lus10033516
131 / 1e-36
AT2G34790 451 / 1e-153
MATERNAL EFFECT EMBRYO ARREST 23, EMBRYO SAC DEVELOPMENT ARREST 28, FAD-binding Berberine family protein (.1)
Lus10023369
121 / 2e-36
AT1G30710 156 / 4e-46
FAD-binding Berberine family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0277
FAD-oxidase_C
PF08031
BBE
Berberine and berberine like
Representative CDS sequence
>Lus10042383 pacid=23153466 polypeptide=Lus10042383 locus=Lus10042383.g ID=Lus10042383.BGIv1.0 annot-version=v1.0
ATGAACGAGATTTCAGAATCGTACACGCCGTTCCCGAACAGGAAAGGGCATCTTTACATCATTGAGTACATTGCGAGGTGGGGGCATAAGGAGGAAGAGG
AGAGGCACCTTGAATGGATGAAGAAGCTTTACGACTACATGGAACCATATGTTTCGAAGTCCTTGAGGAGGGCGTATTTGAATTACAGGGATCTGGACAT
TGGGACCAACAATCTTGACACGACCATGACGACAAGCTTCTCGGAGGCGAGTGTGTGGGGAAGGAAGTACTTCAGGGACAATTTTGAGAAGTTGTGTAAG
GTAAAGAGTATGATCGATCCTGAAGATTTCTTTAGCGATGAACAGAGCATTCCTCAGTTTAGTGTTTATCTAAGGGACAATTAG
AA sequence
>Lus10042383 pacid=23153466 polypeptide=Lus10042383 locus=Lus10042383.g ID=Lus10042383.BGIv1.0 annot-version=v1.0
MNEISESYTPFPNRKGHLYIIEYIARWGHKEEEERHLEWMKKLYDYMEPYVSKSLRRAYLNYRDLDIGTNNLDTTMTTSFSEASVWGRKYFRDNFEKLCK
VKSMIDPEDFFSDEQSIPQFSVYLRDN
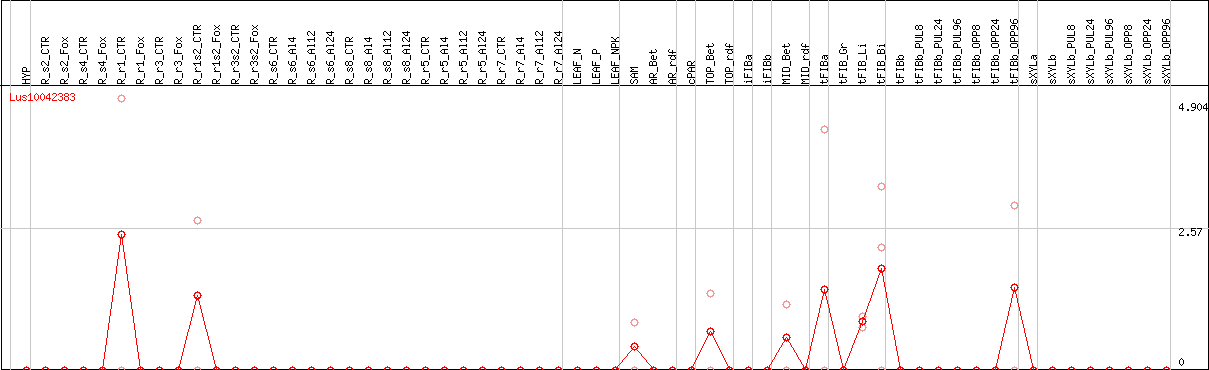
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10042383 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.