External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G17810 67 / 6e-13
BETA-TIP
beta-tonoplast intrinsic protein (.1)
AT1G73190 61 / 6e-11
ALPHA-TIP, TIP3;1
ALPHA-TONOPLAST INTRINSIC PROTEIN, Aquaporin-like superfamily protein (.1)
AT3G61430 61 / 7e-11
ATPIP1, PIP1;1, PIP1A
PLASMA MEMBRANE INTRINSIC PROTEIN 1;1, ARABIDOPSIS THALIANA PLASMA MEMBRANE INTRINSIC PROTEIN 1, plasma membrane intrinsic protein 1A (.1.2)
AT4G00430 61 / 1e-10
PIP1;4, TMP-C, PIP1E
TRANSMEMBRANE PROTEIN C, PLASMA MEMBRANE INTRINSIC PROTEIN 1E, plasma membrane intrinsic protein 1;4 (.1.2)
AT4G23400 60 / 3e-10
PIP1D, PIP1;5
plasma membrane intrinsic protein 1;5 (.1)
AT1G01620 59 / 4e-10
TMP-B, PIP1C, PIP1;3
PLASMA MEMBRANE INTRINSIC PROTEIN 1;3, plasma membrane intrinsic protein 1C (.1.2)
AT2G45960 59 / 5e-10
PIP1;2, ATHH2, TMP-A, PIP1B
TRANSMEMBRANE PROTEIN A, NAMED PLASMA MEMBRANE INTRINSIC PROTEIN 1;2, plasma membrane intrinsic protein 1B (.1.2.3)
AT3G26520 58 / 9e-10
TIP1;2, SITIP, GAMMA-TIP2, TIP2
SALT-STRESS INDUCIBLE TONOPLAST INTRINSIC PROTEIN, tonoplast intrinsic protein 2 (.1)
AT2G16850 58 / 1e-09
PIP3B, PIP2;8
PLASMA MEMBRANE INTRINSIC PROTEIN 3B, plasma membrane intrinsic protein 2;8 (.1)
AT3G16240 57 / 2e-09
DELTA-TIP1, ATTIP2;1, AQP1, DELTA-TIP
delta tonoplast integral protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10007568
258 / 1e-85
AT2G45960 108 / 3e-27
TRANSMEMBRANE PROTEIN A, NAMED PLASMA MEMBRANE INTRINSIC PROTEIN 1;2, plasma membrane intrinsic protein 1B (.1.2.3)
Lus10042375
139 / 3e-39
AT1G73190 104 / 3e-26
ALPHA-TONOPLAST INTRINSIC PROTEIN, Aquaporin-like superfamily protein (.1)
Lus10040652
66 / 2e-12
AT1G17810 367 / 2e-129
beta-tonoplast intrinsic protein (.1)
Lus10018256
66 / 2e-12
AT1G17810 370 / 7e-131
beta-tonoplast intrinsic protein (.1)
Lus10036187
62 / 4e-11
AT1G73190 382 / 4e-135
ALPHA-TONOPLAST INTRINSIC PROTEIN, Aquaporin-like superfamily protein (.1)
Lus10038324
62 / 4e-11
AT1G73190 380 / 1e-134
ALPHA-TONOPLAST INTRINSIC PROTEIN, Aquaporin-like superfamily protein (.1)
Lus10039222
60 / 3e-10
AT2G16850 518 / 0.0
PLASMA MEMBRANE INTRINSIC PROTEIN 3B, plasma membrane intrinsic protein 2;8 (.1)
Lus10027467
59 / 5e-10
AT2G16850 518 / 0.0
PLASMA MEMBRANE INTRINSIC PROTEIN 3B, plasma membrane intrinsic protein 2;8 (.1)
Lus10014978
58 / 1e-09
AT2G16850 514 / 0.0
PLASMA MEMBRANE INTRINSIC PROTEIN 3B, plasma membrane intrinsic protein 2;8 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.009G128500
303 / 2e-103
AT4G23400 85 / 6e-19
plasma membrane intrinsic protein 1;5 (.1)
Potri.009G128300
284 / 2e-96
AT3G61430 93 / 6e-22
PLASMA MEMBRANE INTRINSIC PROTEIN 1;1, ARABIDOPSIS THALIANA PLASMA MEMBRANE INTRINSIC PROTEIN 1, plasma membrane intrinsic protein 1A (.1.2)
Potri.004G167000
285 / 3e-96
AT3G61430 96 / 6e-23
PLASMA MEMBRANE INTRINSIC PROTEIN 1;1, ARABIDOPSIS THALIANA PLASMA MEMBRANE INTRINSIC PROTEIN 1, plasma membrane intrinsic protein 1A (.1.2)
Potri.009G128000
283 / 1e-95
AT4G23400 100 / 3e-24
plasma membrane intrinsic protein 1;5 (.1)
Potri.009G127900
275 / 2e-92
AT2G45960 107 / 6e-27
TRANSMEMBRANE PROTEIN A, NAMED PLASMA MEMBRANE INTRINSIC PROTEIN 1;2, plasma membrane intrinsic protein 1B (.1.2.3)
Potri.009G128100
163 / 3e-49
AT1G73190 91 / 1e-21
ALPHA-TONOPLAST INTRINSIC PROTEIN, Aquaporin-like superfamily protein (.1)
Potri.018G152100
64 / 7e-12
AT1G17810 350 / 8e-123
beta-tonoplast intrinsic protein (.1)
Potri.003G128600
62 / 5e-11
AT4G00430 514 / 0.0
TRANSMEMBRANE PROTEIN C, PLASMA MEMBRANE INTRINSIC PROTEIN 1E, plasma membrane intrinsic protein 1;4 (.1.2)
Potri.006G098100
61 / 1e-10
AT4G00430 482 / 4e-174
TRANSMEMBRANE PROTEIN C, PLASMA MEMBRANE INTRINSIC PROTEIN 1E, plasma membrane intrinsic protein 1;4 (.1.2)
Potri.016G113300
59 / 4e-10
AT4G00430 524 / 0.0
TRANSMEMBRANE PROTEIN C, PLASMA MEMBRANE INTRINSIC PROTEIN 1E, plasma membrane intrinsic protein 1;4 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00230
MIP
Major intrinsic protein
Representative CDS sequence
>Lus10042385 pacid=23153929 polypeptide=Lus10042385 locus=Lus10042385.g ID=Lus10042385.BGIv1.0 annot-version=v1.0
ATGGCTGGAAATGGGATTGAAGATGAAGAAAGTGGAGGCGGCTATGGCGGAAACAGAGTATGGCGAGCATCAGTTGCTGAGCTGGTGGGAACCGCGGTAC
TAGTCTTCGCAATGGACACAATCGCCATTTCCTCGTACGAGACCAAGACGTCGACCCCTAACTTGGTGATGTCGATCCTAATCGCTGTCATTGTGACCCT
TATTCTCCTGGCAACTTTCCCAGTCTCCGGTGGCCACATCAACCCAGTCATCACCATCGCCGCTATGCTTACTGGCCTCGTCACCGTTTCTCGGGCAGCC
ATCTACATTGCGGCTCAATGTGCGGGCGCCATACTGGGCGCCCTGGCATTGAAAGCCGTGGTTAGTAGCACCATCGAACAGACATTTTCGCTCGGAGGCT
GTACTCTAAACGTCGTCGTGCCCGGCCCACAGGGACCCGTTGTCATTGGGCTCGAAACCAAGCAGGCCCTTTGGATGGAGATACTATGCACCTTCTTGTT
TCTGTTCGTCGCCATTTGGGTAGCCTTCGACGGCAGGCAGTTTAGAAGATTGGGCCGGGTAGTGGTGTTCTGCATAATTGGGCTTATGGTGGGCCTCATT
GTCTTTATTTCGACGACGCTTACTGGAACCAAGGGTTATGCTGGGGTTGGGATGAACCCAGCAAGATGTTTGGGCCCAGCGATTGTGAGAGGAGGCCATC
TGTGGACTGGACATTGGATATTTTGGGCCGGACCGGGTATTGCTTGCGTGGCTTTTGCTATCTACGCAAAAATCATCCCCAAGGAAAATCTATTCGGGTA
G
AA sequence
>Lus10042385 pacid=23153929 polypeptide=Lus10042385 locus=Lus10042385.g ID=Lus10042385.BGIv1.0 annot-version=v1.0
MAGNGIEDEESGGGYGGNRVWRASVAELVGTAVLVFAMDTIAISSYETKTSTPNLVMSILIAVIVTLILLATFPVSGGHINPVITIAAMLTGLVTVSRAA
IYIAAQCAGAILGALALKAVVSSTIEQTFSLGGCTLNVVVPGPQGPVVIGLETKQALWMEILCTFLFLFVAIWVAFDGRQFRRLGRVVVFCIIGLMVGLI
VFISTTLTGTKGYAGVGMNPARCLGPAIVRGGHLWTGHWIFWAGPGIACVAFAIYAKIIPKENLFG
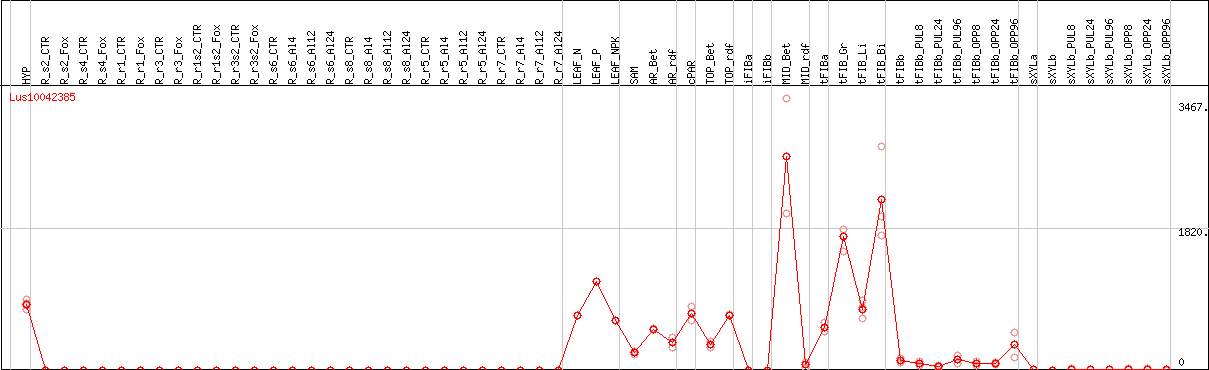
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10042385 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.