External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G54460 123 / 1e-32
TPX2 (targeting protein for Xklp2) protein family (.1)
AT5G28646 112 / 7e-30
WVD2
WAVE-DAMPENED 2, TPX2 (targeting protein for Xklp2) protein family (.1), TPX2 (targeting protein for Xklp2) protein family (.2)
AT3G04630 92 / 1e-21
WDL1
WVD2-like 1 (.1.2.3)
AT3G23090 90 / 2e-20
TPX2 (targeting protein for Xklp2) protein family (.1), TPX2 (targeting protein for Xklp2) protein family (.2)
AT2G35880 70 / 2e-13
TPX2 (targeting protein for Xklp2) protein family (.1)
AT4G32330 65 / 1e-11
TPX2 (targeting protein for Xklp2) protein family (.1), TPX2 (targeting protein for Xklp2) protein family (.2), TPX2 (targeting protein for Xklp2) protein family (.3)
AT2G25480 62 / 1e-10
TPX2 (targeting protein for Xklp2) protein family (.1)
AT1G70950 59 / 2e-09
TPX2 (targeting protein for Xklp2) protein family (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10022003
456 / 5e-163
AT1G54460 172 / 3e-51
TPX2 (targeting protein for Xklp2) protein family (.1)
Lus10000016
272 / 2e-92
AT5G28646 137 / 3e-41
WAVE-DAMPENED 2, TPX2 (targeting protein for Xklp2) protein family (.1), TPX2 (targeting protein for Xklp2) protein family (.2)
Lus10005533
186 / 9e-57
AT1G54460 187 / 3e-57
TPX2 (targeting protein for Xklp2) protein family (.1)
Lus10006563
182 / 2e-55
AT1G54460 187 / 2e-57
TPX2 (targeting protein for Xklp2) protein family (.1)
Lus10021223
87 / 5e-19
AT3G23090 230 / 4e-73
TPX2 (targeting protein for Xklp2) protein family (.1), TPX2 (targeting protein for Xklp2) protein family (.2)
Lus10022213
85 / 3e-18
AT3G23090 220 / 2e-68
TPX2 (targeting protein for Xklp2) protein family (.1), TPX2 (targeting protein for Xklp2) protein family (.2)
Lus10002915
69 / 9e-13
AT4G32330 228 / 2e-70
TPX2 (targeting protein for Xklp2) protein family (.1), TPX2 (targeting protein for Xklp2) protein family (.2), TPX2 (targeting protein for Xklp2) protein family (.3)
Lus10016193
62 / 2e-10
AT2G35880 181 / 7e-52
TPX2 (targeting protein for Xklp2) protein family (.1)
Lus10029355
61 / 3e-10
AT2G35880 185 / 3e-53
TPX2 (targeting protein for Xklp2) protein family (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.013G042800
159 / 4e-46
AT1G54460 176 / 5e-52
TPX2 (targeting protein for Xklp2) protein family (.1)
Potri.005G055900
139 / 1e-38
AT1G54460 207 / 4e-64
TPX2 (targeting protein for Xklp2) protein family (.1)
Potri.008G162800
100 / 1e-23
AT3G23090 214 / 1e-66
TPX2 (targeting protein for Xklp2) protein family (.1), TPX2 (targeting protein for Xklp2) protein family (.2)
Potri.010G076200
99 / 2e-23
AT3G23090 185 / 5e-55
TPX2 (targeting protein for Xklp2) protein family (.1), TPX2 (targeting protein for Xklp2) protein family (.2)
Potri.018G027500
66 / 7e-12
AT4G32330 268 / 2e-85
TPX2 (targeting protein for Xklp2) protein family (.1), TPX2 (targeting protein for Xklp2) protein family (.2), TPX2 (targeting protein for Xklp2) protein family (.3)
Potri.006G254400
66 / 9e-12
AT4G32330 288 / 2e-93
TPX2 (targeting protein for Xklp2) protein family (.1), TPX2 (targeting protein for Xklp2) protein family (.2), TPX2 (targeting protein for Xklp2) protein family (.3)
Potri.016G066600
63 / 6e-11
AT2G35880 149 / 5e-40
TPX2 (targeting protein for Xklp2) protein family (.1)
Potri.006G200400
59 / 1e-09
AT2G35880 168 / 1e-46
TPX2 (targeting protein for Xklp2) protein family (.1)
Potri.008G129300
58 / 3e-09
AT1G70950 241 / 2e-72
TPX2 (targeting protein for Xklp2) protein family (.1)
Potri.010G113000
58 / 4e-09
AT1G70950 147 / 5e-38
TPX2 (targeting protein for Xklp2) protein family (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF06886
TPX2
Targeting protein for Xklp2 (TPX2) domain
Representative CDS sequence
>Lus10042543 pacid=23181675 polypeptide=Lus10042543 locus=Lus10042543.g ID=Lus10042543.BGIv1.0 annot-version=v1.0
ATGGATAAGAAGCATAAAGGTGTTGCAAGGAAGTTGAAAAACGGTAGTATGCATCAAGAGAAAGAAACCAAGGAGGACTTGGATGTGTTAGGAGTGAAGA
ACATGAATAATGGTAGTAGTACCCCTGTTTCTGATCAGAAAAGCCAGAAGCCAAAAGCTGAGAAGTCAATTGATGGTAAAGTGACGAGCTTTCCCACGCC
AAAGTCGTCTAATCTCGGTCCAGGAAGCGCTGAAGCAGTTTGTTTAGAGACTGAAGAGCATGGAGATTATGGGAAGATTTCAGAAAATGATGAGAAACAC
AATGATGAAGATGATAAGAACTCGGTTGCTTCCTCGTCAGCTGCATATAAGAATACTAAATCTGCAGTGACTGTTGGCACAGCTCCTCAATTTAAATCTG
CTGAGCGTGCCCAAAAGCGGAAGGAGTACTATATGAAATTGGAGGAAAAACGGAAAACTCTCGAGGCTGAGAAAGCTCAAGCTGAGGCCAGGAACAAGGA
AGAGCAACAAGCAGCGATCCGACAGATGAGAAAGAACATGATCTTCAAAGCGAACCCAGTCCCCAATTTCTATTATGAGCCACCTCCACCAAAGGTTGAA
CTCAAGAAGCTGCCATTGACGCGGCCCGTGTCCCCAAAGCTAAACCGGAGAAAGAGCTGTGGCGACCTGGTCTACACATCTCAGGCGCAACAAGTCAGCA
GCAAGCACTGTGCCAAGCATCGCGAAAGCCTCGGTCCGGACCATAATGAATCTCCATCTGCAACAAGAGCCATTCTTCGTCGCACACAGACCGCAAAGGC
GAAAAGCTATTCAGGGTTTGAACAAGAAGAACATCAGGACGTCAGTGAAACCAATGATGGTCCTGAAGAGGTCAACAATGAGCAACCGAGCGCCAATGTT
TCTGTTGAGTCATGA
AA sequence
>Lus10042543 pacid=23181675 polypeptide=Lus10042543 locus=Lus10042543.g ID=Lus10042543.BGIv1.0 annot-version=v1.0
MDKKHKGVARKLKNGSMHQEKETKEDLDVLGVKNMNNGSSTPVSDQKSQKPKAEKSIDGKVTSFPTPKSSNLGPGSAEAVCLETEEHGDYGKISENDEKH
NDEDDKNSVASSSAAYKNTKSAVTVGTAPQFKSAERAQKRKEYYMKLEEKRKTLEAEKAQAEARNKEEQQAAIRQMRKNMIFKANPVPNFYYEPPPPKVE
LKKLPLTRPVSPKLNRRKSCGDLVYTSQAQQVSSKHCAKHRESLGPDHNESPSATRAILRRTQTAKAKSYSGFEQEEHQDVSETNDGPEEVNNEQPSANV
SVES
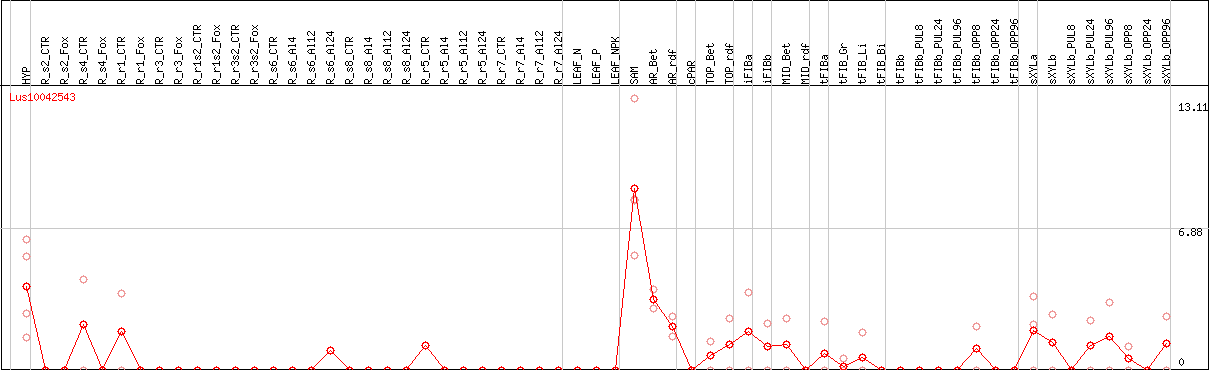
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10042543 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.