External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G63200 108 / 9e-29
PLP9, PLAIIIB ,PLA IIIB
PATATIN-like protein 9 (.1)
AT4G29800 66 / 3e-13
PLP8, PLAIVD ,PLA IVD
PATATIN-like protein 8 (.1.2)
AT2G39220 62 / 5e-12
PLP6, PLAIIB ,PLA IIB
PATATIN-like protein 6 (.1)
AT5G43590 62 / 7e-12
Acyl transferase/acyl hydrolase/lysophospholipase superfamily protein (.1)
AT4G37060 61 / 1e-11
AtPLAIVB, PLP5, PLAIVB ,PLA IVB
phospholipase A IVB, PATATIN-like protein 5 (.1.2)
AT2G26560 59 / 7e-11
PLP2, PLAIIA, PLA2A
PATATIN-LIKE PROTEIN 2, phospholipase A 2A (.1)
AT3G54950 59 / 7e-11
pPLAIIIbeta, PLP7, PLAIIIA ,PLA IIIA
patatin-related phospholipase IIIbeta, PATATIN-LIKE PROTEIN 7, patatin-like protein 6 (.1)
AT4G37070 54 / 2e-09
AtPLAIVA, PLP1, PLAIVA ,PLA IVA
phospholipase A IVA, Acyl transferase/acyl hydrolase/lysophospholipase superfamily protein (.1.2.3.4)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10041402
66 / 4e-13
AT3G54950 637 / 0.0
patatin-related phospholipase IIIbeta, PATATIN-LIKE PROTEIN 7, patatin-like protein 6 (.1)
Lus10036526
64 / 2e-12
AT2G39220 640 / 0.0
PATATIN-like protein 6 (.1)
Lus10038349
63 / 4e-12
AT3G54950 354 / 2e-118
patatin-related phospholipase IIIbeta, PATATIN-LIKE PROTEIN 7, patatin-like protein 6 (.1)
Lus10019637
58 / 1e-10
AT4G37070 486 / 1e-170
phospholipase A IVA, Acyl transferase/acyl hydrolase/lysophospholipase superfamily protein (.1.2.3.4)
Lus10000279
58 / 1e-10
AT4G37070 486 / 1e-170
phospholipase A IVA, Acyl transferase/acyl hydrolase/lysophospholipase superfamily protein (.1.2.3.4)
Lus10023512
58 / 2e-10
AT4G29800 523 / 0.0
PATATIN-like protein 8 (.1.2)
Lus10040394
57 / 3e-10
AT2G39220 642 / 0.0
PATATIN-like protein 6 (.1)
Lus10009340
56 / 6e-10
AT4G37070 489 / 7e-173
phospholipase A IVA, Acyl transferase/acyl hydrolase/lysophospholipase superfamily protein (.1.2.3.4)
Lus10022040
53 / 1e-08
AT3G63200 413 / 3e-145
PATATIN-like protein 9 (.1)
PFAM info
Representative CDS sequence
>Lus10042585 pacid=23181730 polypeptide=Lus10042585 locus=Lus10042584.g ID=Lus10042585.BGIv1.0 annot-version=v1.0
ATGCCGCCAGGAAGACCAGAGTTCTTAGCATTGACGGTGGCGGAAGCACCGGCATTGTCGCTGGAGCCGCTCTTGTCCATCTCTCTTGTCCATCTCGAGG
ACCAGATCCGACTCAAGCTTGGAGATCCACATGCTCGAATCGCTGATTTCTTCGACCTCATTGCCGGCACTGGCATTGGTGCCGTTCTCGCTTCCATGCT
AGCTGCCGACGATGGTTCCGGCCGTCCTCTGTTCTCCGCTCGTGAAGCGGTTGATTTCCTTGCTGACAGAAACTCCGAACTCTTCAAACCGAAGCGCCTC
TCCGCCGTCCTCTCCGGCAGCCGCCCCGCATTCTCCGGGCGGCGTGTCTGTTCAACCCGTTCGACCTCAAATCCGTCGACGGTAAGACATCCTGCTCCGC
TATCGATGGAGGATTGGTGA
AA sequence
>Lus10042585 pacid=23181730 polypeptide=Lus10042585 locus=Lus10042584.g ID=Lus10042585.BGIv1.0 annot-version=v1.0
MPPGRPEFLALTVAEAPALSLEPLLSISLVHLEDQIRLKLGDPHARIADFFDLIAGTGIGAVLASMLAADDGSGRPLFSAREAVDFLADRNSELFKPKRL
SAVLSGSRPAFSGRRVCSTRSTSNPSTVRHPAPLSMEDW
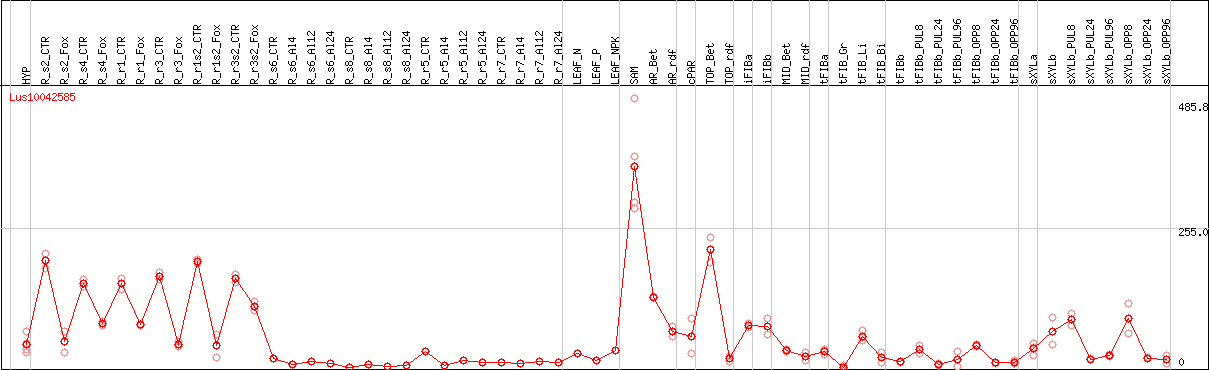
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10042585 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.