External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G19270 612 / 0
CYP707A4
"cytochrome P450, family 707, subfamily A, polypeptide 4", cytochrome P450, family 707, subfamily A, polypeptide 4 (.1)
AT4G19230 532 / 0
CYP707A1
"cytochrome P450, family 707, subfamily A, polypeptide 1", cytochrome P450, family 707, subfamily A, polypeptide 1 (.1.2)
AT5G45340 525 / 0
CYP707A3
"cytochrome P450, family 707, subfamily A, polypeptide 3", cytochrome P450, family 707, subfamily A, polypeptide 3 (.1.2)
AT2G29090 499 / 1e-174
CYP707A2
"cytochrome P450, family 707, subfamily A, polypeptide 2", cytochrome P450, family 707, subfamily A, polypeptide 2 (.1.2)
AT5G05690 259 / 9e-81
CBB3, DWF3, CYP90A1, CYP90A, CPD
DWARF 3, CYTOCHROME P450 90A1, CONSTITUTIVE PHOTOMORPHOGENIC DWARF, CABBAGE 3, Cytochrome P450 superfamily protein (.1.2.3)
AT1G05160 248 / 3e-76
ATKAO1, CYP88A3
ENT-KAURENOIC ACID OXYDASE 1, "cytochrome P450, family 88, subfamily A, polypeptide 3", cytochrome P450, family 88, subfamily A, polypeptide 3 (.1)
AT2G32440 241 / 7e-74
ATKAO2, CYP88A4, KAO2
ARABIDOPSIS ENT-KAURENOIC ACID HYDROXYLASE 2, ent-kaurenoic acid hydroxylase 2 (.1.2)
AT1G19630 235 / 1e-71
CYP722A1
"cytochrome P450, family 722, subfamily A, polypeptide 1", cytochrome P450, family 722, subfamily A, polypeptide 1 (.1)
AT3G50660 236 / 2e-71
PSC1, CYP90B1, CLM, SNP2, DWF4, SAV1
SUPPRESSOR OF NPH4 2, SHADE AVOIDANCE 1, PARTIALLY SUPPRESSING COI1 INSENSITIVITY TO JA 1, DWARF 4, CYTOCHROME P450 90B1, CLOMAZONE-RESISTANT, Cytochrome P450 superfamily protein (.1)
AT4G36380 235 / 5e-71
ROT3
ROTUNDIFOLIA 3, Cytochrome P450 superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10021725
832 / 0
AT3G19270 633 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 4", cytochrome P450, family 707, subfamily A, polypeptide 4 (.1)
Lus10019858
562 / 0
AT3G19270 643 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 4", cytochrome P450, family 707, subfamily A, polypeptide 4 (.1)
Lus10012675
527 / 0
AT3G19270 586 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 4", cytochrome P450, family 707, subfamily A, polypeptide 4 (.1)
Lus10033308
508 / 7e-178
AT5G45340 732 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 3", cytochrome P450, family 707, subfamily A, polypeptide 3 (.1.2)
Lus10034768
501 / 2e-175
AT5G45340 731 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 3", cytochrome P450, family 707, subfamily A, polypeptide 3 (.1.2)
Lus10035685
496 / 3e-173
AT4G19230 714 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 1", cytochrome P450, family 707, subfamily A, polypeptide 1 (.1.2)
Lus10016515
440 / 4e-151
AT2G29090 554 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 2", cytochrome P450, family 707, subfamily A, polypeptide 2 (.1.2)
Lus10040785
352 / 1e-109
AT1G07510 1078 / 0.0
FTSH protease 10 (.1)
Lus10037273
322 / 3e-106
AT4G19230 461 / 2e-161
"cytochrome P450, family 707, subfamily A, polypeptide 1", cytochrome P450, family 707, subfamily A, polypeptide 1 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.014G029100
679 / 0
AT3G19270 665 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 4", cytochrome P450, family 707, subfamily A, polypeptide 4 (.1)
Potri.002G126100
677 / 0
AT3G19270 666 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 4", cytochrome P450, family 707, subfamily A, polypeptide 4 (.1)
Potri.004G140900
630 / 0
AT3G19270 731 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 4", cytochrome P450, family 707, subfamily A, polypeptide 4 (.1)
Potri.009G101700
615 / 0
AT3G19270 711 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 4", cytochrome P450, family 707, subfamily A, polypeptide 4 (.1)
Potri.004G235400
527 / 0
AT4G19230 778 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 1", cytochrome P450, family 707, subfamily A, polypeptide 1 (.1.2)
Potri.009G033900
506 / 2e-177
AT2G29090 677 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 2", cytochrome P450, family 707, subfamily A, polypeptide 2 (.1.2)
Potri.001G242600
501 / 5e-175
AT2G29090 652 / 0.0
"cytochrome P450, family 707, subfamily A, polypeptide 2", cytochrome P450, family 707, subfamily A, polypeptide 2 (.1.2)
Potri.002G069600
271 / 2e-85
AT5G45340 296 / 1e-95
"cytochrome P450, family 707, subfamily A, polypeptide 3", cytochrome P450, family 707, subfamily A, polypeptide 3 (.1.2)
Potri.008G067500
254 / 3e-79
AT5G05690 710 / 0.0
DWARF 3, CYTOCHROME P450 90A1, CONSTITUTIVE PHOTOMORPHOGENIC DWARF, CABBAGE 3, Cytochrome P450 superfamily protein (.1.2.3)
Potri.010G189800
251 / 1e-77
AT5G05690 717 / 0.0
DWARF 3, CYTOCHROME P450 90A1, CONSTITUTIVE PHOTOMORPHOGENIC DWARF, CABBAGE 3, Cytochrome P450 superfamily protein (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00067
p450
Cytochrome P450
Representative CDS sequence
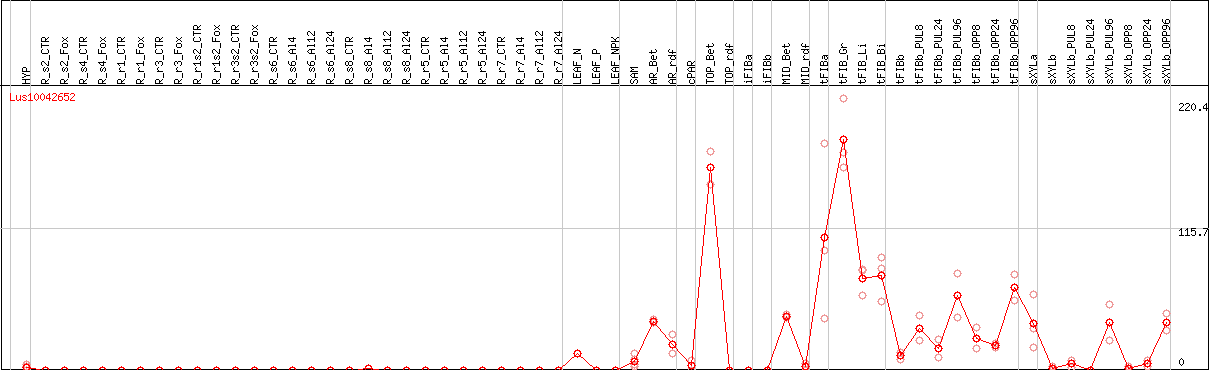
>Lus10042652 pacid=23182042 polypeptide=Lus10042652 locus=Lus10042652.g ID=Lus10042652.BGIv1.0 annot-version=v1.0
ATGGGCGATGGATTCTTTCTCAATATCCTACTACTAGTAGTAGTATTCATCATCATCATCATCATCATCATCACTCTCTCTTGCTACTTATTACTGATCA
CCAAAATCAGAAAACCTCACCTCCTCCCTCCTGGCTCCATGGGGTGGCCTGTGGTGGGCGAGACTCTGCAGCTATACTCACAGGATCCATCCATTTTCTT
CTCATCCAAACAGAAAAGATATGGAGAAATATTCAAGACCCACATCCTAGGTTGCCCTTGCATCATGCTTGCTAGCCCTGAAGCTGCACGATTCGTTCTT
GTCACTCAAGCTCATCTCTTTAAGCCCACCTACCCTAAATCTAAAGAGCGACTCATTGGCCCCAAAGCCCTCTTCTTCCACATGGGAGATTACCATCTCC
GTCTCCGCAAGCTCGTCCAGTCAGCTCTTTCTCTCCACCGTCTCCGCAGCTTGGTCGGAGATGTCTCCTCCCTCACCCTCTCCGCCTTGGATTCTTGGCA
CGGCGGCCAGATCGTTACCACATATGAACAACTCAAAATGTTGTCGTTCGAAGTTGGAATCCTGGCGATTTTCGGACACCTGGAAGGGCATTACAGGACA
GAGTTGAAAATGAACTATGGCATCGTTGAAAAAGGCTACAATTCTTTTCCCACCTCTGTTCCTGGATCTCCTTACAAAAGGGCTCTTACGGCGAGGAGGA
AGCTAAATGAGATATTAGGGGCCATAATTAGCGAGAGGAAGGAGAAGAAGTTACTAGAGAAGGACCTGTTGGGCTGTTTACTAAACTCCAAGGACCAAAG
GGGAGAGGAGTTGACCGACGACCAAGTTGCCGACAACATCATCGGAGTTCTGTTCGCCGCTCAGGACACGACGGCCAGTGCCATGACTTGGATTATCAAG
TACCTCCACGACAACCCCAAACTTCTCGAGGCCGTCAAGGCTGAACAGAAGGCAATTCAAGAATTGAATGAAACAGACAAGCGTAGTAGTAGTAGTAGTA
ATGAGACGACGGTGACTTGGAGTCAAACTCGTAAAAATATGCCTTTCACTCACAAGGTTGTGTTAGAGAGCCTGAGGATGGCAAGCATTATAGCGTTCAC
GTTTAGAGAAGCAGTCACTGACGTCGAATACAAAGGGTACCTTATTCCAAAAGGTTGGAAGGTGATGCCCTTATTCAGGAACATTCATCACAATCCAGAA
TATTTCAGTGATCCTCATAAATTTGAACCTTCAAGATTTGAGGTTGCAGCAAAGGCGAACACATTTATGCCATTTGGAAATGGGGTGCATGCTTGTCCAG
GAAATGAGCTTGCCAAGCTGGAGATGCTCATCTTCATCCACCATCTGGTCACCAAGTACAGGTGGGAAGTGGTTGGAGATGAAAGTGGGACACAGAACTG
CCCATTTCCAGTGCCAATGAATGGACTCCCAGCTAGATTTTGGAATACAACAACACCAAGTTCCCAGCACCTTAACTAA
AA sequence
>Lus10042652 pacid=23182042 polypeptide=Lus10042652 locus=Lus10042652.g ID=Lus10042652.BGIv1.0 annot-version=v1.0
MGDGFFLNILLLVVVFIIIIIIIITLSCYLLLITKIRKPHLLPPGSMGWPVVGETLQLYSQDPSIFFSSKQKRYGEIFKTHILGCPCIMLASPEAARFVL
VTQAHLFKPTYPKSKERLIGPKALFFHMGDYHLRLRKLVQSALSLHRLRSLVGDVSSLTLSALDSWHGGQIVTTYEQLKMLSFEVGILAIFGHLEGHYRT
ELKMNYGIVEKGYNSFPTSVPGSPYKRALTARRKLNEILGAIISERKEKKLLEKDLLGCLLNSKDQRGEELTDDQVADNIIGVLFAAQDTTASAMTWIIK
YLHDNPKLLEAVKAEQKAIQELNETDKRSSSSSNETTVTWSQTRKNMPFTHKVVLESLRMASIIAFTFREAVTDVEYKGYLIPKGWKVMPLFRNIHHNPE
YFSDPHKFEPSRFEVAAKANTFMPFGNGVHACPGNELAKLEMLIFIHHLVTKYRWEVVGDESGTQNCPFPVPMNGLPARFWNTTTPSSQHLN