External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G17930 48 / 2e-07
Aminotransferase-like, plant mobile domain family protein (.1)
AT2G25010 48 / 2e-07
Aminotransferase-like, plant mobile domain family protein (.1)
AT1G48120 47 / 6e-07
hydrolases;protein serine/threonine phosphatases (.1)
AT2G04865 44 / 4e-06
Aminotransferase-like, plant mobile domain family protein (.1)
AT1G51538 44 / 7e-06
Aminotransferase-like, plant mobile domain family protein (.1)
AT1G50770 43 / 1e-05
Aminotransferase-like, plant mobile domain family protein (.1)
AT1G50820 41 / 4e-05
Aminotransferase-like, plant mobile domain family protein (.1)
AT1G50790 40 / 0.0002
Plant mobile domain protein family (.1)
AT4G16050 39 / 0.0005
Aminotransferase-like, plant mobile domain family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10004818
145 / 1e-43
AT1G17930 56 / 2e-08
Aminotransferase-like, plant mobile domain family protein (.1)
Lus10008761
144 / 1e-42
AT2G25010 79 / 1e-15
Aminotransferase-like, plant mobile domain family protein (.1)
Lus10012081
128 / 4e-38
AT1G17930 53 / 8e-08
Aminotransferase-like, plant mobile domain family protein (.1)
Lus10032693
86 / 5e-23
AT2G25010 52 / 3e-09
Aminotransferase-like, plant mobile domain family protein (.1)
Lus10027047
90 / 4e-22
AT1G17930 119 / 1e-28
Aminotransferase-like, plant mobile domain family protein (.1)
Lus10020970
86 / 8e-21
AT1G17930 123 / 2e-30
Aminotransferase-like, plant mobile domain family protein (.1)
Lus10003430
69 / 1e-14
AT5G51100 60 / 6e-10
Fe superoxide dismutase 2 (.1)
Lus10014633
66 / 1e-14
AT2G04865 61 / 2e-11
Aminotransferase-like, plant mobile domain family protein (.1)
Lus10010613
57 / 4e-11
ND /
No hit found
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF10536
PMD
Plant mobile domain
Representative CDS sequence
>Lus10042711 pacid=23181947 polypeptide=Lus10042711 locus=Lus10042711.g ID=Lus10042711.BGIv1.0 annot-version=v1.0
ATGATGATGTATACCTCGGGACTTCTCCAGTTGGGTCGTTTTGGTATAATCGATATGTTGGACGAGGCTCTGATCCACGCTTTTGTTGAGAGGTGGCAGC
CAGACACTAACACGTTTCCCATGCCATTTGAAGAGATGATCATCACACTTCATGATGTTTGGCATACTCTTCGTATCCCGGTACACGGACGTCCACTTCA
CATTCTACGGTCCAGCGAGGAGTTGTTGGCCGATATAGTTAGGGTCCTCCGTATCCATCCTGGGGATTTGAAGTTTTCAGTTGGAGGCCCAGCAGGTACG
GGATGCCTGATACCCTTGGCTGAAGGGTGCTGCGATGCATGA
AA sequence
>Lus10042711 pacid=23181947 polypeptide=Lus10042711 locus=Lus10042711.g ID=Lus10042711.BGIv1.0 annot-version=v1.0
MMMYTSGLLQLGRFGIIDMLDEALIHAFVERWQPDTNTFPMPFEEMIITLHDVWHTLRIPVHGRPLHILRSSEELLADIVRVLRIHPGDLKFSVGGPAGT
GCLIPLAEGCCDA
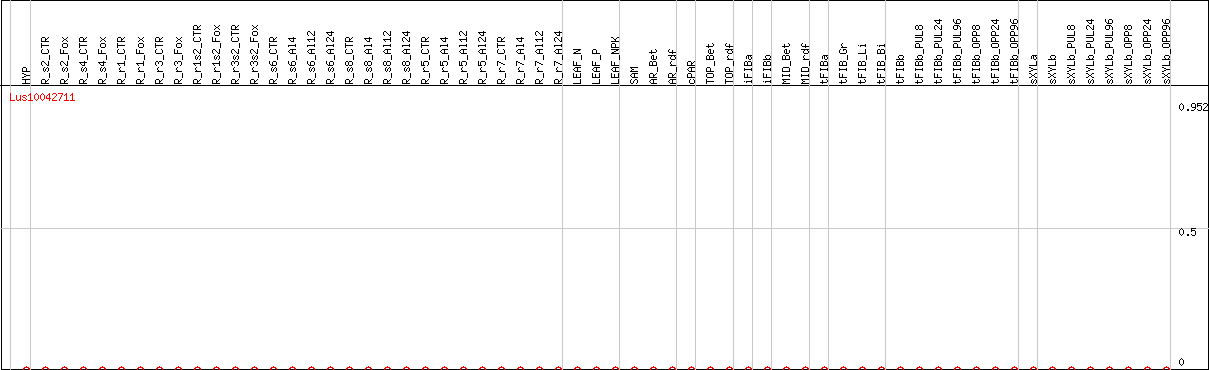
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10042711 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.