External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G10960 150 / 2e-43
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
AT2G32070 145 / 1e-41
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
AT1G80780 144 / 6e-41
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1.2.3)
AT1G15920 131 / 6e-36
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1.2)
AT3G44260 130 / 7e-36
AtCAF1a
CCR4- associated factor 1a, Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
AT1G06450 130 / 4e-35
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
AT1G61470 125 / 7e-34
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
AT1G27820 125 / 2e-33
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
AT5G22250 122 / 1e-32
AtCAF1b
CCR4- associated factor 1b, Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
AT3G44240 117 / 2e-31
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10029710
541 / 0
AT5G10960 147 / 2e-42
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Lus10029713
526 / 0
AT5G10960 144 / 4e-41
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Lus10007771
363 / 2e-126
AT1G61470 170 / 5e-51
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Lus10000327
226 / 3e-72
AT1G06450 174 / 1e-51
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Lus10035078
217 / 7e-69
AT1G06450 168 / 2e-49
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Lus10008043
213 / 3e-67
AT1G06450 165 / 5e-48
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Lus10001651
206 / 1e-63
AT1G61470 167 / 4e-49
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Lus10017905
203 / 2e-63
AT1G06450 162 / 5e-48
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Lus10035077
194 / 7e-60
AT1G61470 162 / 6e-48
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G038700
182 / 2e-55
AT1G61470 182 / 8e-56
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Potri.001G046700
161 / 1e-47
AT2G32070 472 / 2e-170
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Potri.003G181100
158 / 2e-46
AT1G80780 468 / 1e-168
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1.2.3)
Potri.006G262500
156 / 1e-45
AT2G32070 443 / 5e-159
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Potri.018G020900
155 / 3e-45
AT2G32070 441 / 2e-158
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Potri.004G200400
139 / 4e-39
AT5G22250 419 / 3e-149
CCR4- associated factor 1b, Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Potri.009G161500
134 / 3e-37
AT5G22250 387 / 1e-136
CCR4- associated factor 1b, Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Potri.006G187200
133 / 5e-37
AT2G32070 403 / 2e-143
Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Potri.006G205600
122 / 2e-32
AT5G22250 302 / 4e-103
CCR4- associated factor 1b, Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
Potri.001G040400
120 / 3e-32
AT5G22250 253 / 4e-84
CCR4- associated factor 1b, Polynucleotidyl transferase, ribonuclease H-like superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0219
RNase_H
PF04857
CAF1
CAF1 family ribonuclease
Representative CDS sequence
>Lus10042746 pacid=23181819 polypeptide=Lus10042746 locus=Lus10042746.g ID=Lus10042746.BGIv1.0 annot-version=v1.0
ATGATTACATACGTTCGACAAGTGTGGGATTCGGATCTCCGGAAAGAGCTTGCAGTTCTGGAACAGGCGTTTCCAAAGTACCCAGTACTCAGTGTGGATT
GCGATGGATTCCTGCTCAAAACACCAAGAAACCCTACTGGCGATCTTTGGTATAGCGATCTCAAACACAAAGTCGACCAAACCCACATCACCCAACTCAG
CGTCGCTCTTTCCGACCACGGCGGAACCGTCTTCTGCACCTGGAAGTTCAATTTTGAGTTTGATGTTGACGAAGATGTTCACACCAAAGACACGGTCGCA
TTTCTGAAGGGTCAGGGAATTGATTTCCAAAAGCATAGAGCGGATGGCATCAAAAGGACTCGATTTGGTGCCCAATTCTGCAACTTGATGGCGAGCAAAA
CCACTCCTCTGACATGGGTCACCTTCCATGGGTTGTATGATTTTGCTCACATCCTCAAGGTGCTTACCAACAACACACCCATGCCTGGCACGCCTTCTGG
CTTTAGAGACTTGTTGGGATATTCTTTTGAGGAGATTGTCGATCTCAAGGTGATGGCCAAGTTTTGCGATGGAATTTCTGATATGGAGGAGGTTGGGTTG
CAGAAACTGGCAGATTTGCTGAGCATCGAGAGGGAGGATGGTGAAGCTTTACATGAAGCTGGTTCTGACAGTTTACTCACTGCAATGGTGTTTTCCAACA
TGAGAACAAGATTCCAAGCTGATCAACATCATGTATATAGTGATTATATCTATGGAATCACTGAAAGGATTGGAAGAAGCCCTACCATCCAGGTTCGCTA
CTCGCACCGATGGTTGTGCCAGCCGCTGCTGCCACTGCCCATTCCATGCTATCGTCCCCGTTATCATCCTTATCCTCGTCCTCCTCGTCGATGTCTTCTT
CTTCCCCTGCATACCCCTTCCAGAATGGGTCAGTACTGA
AA sequence
>Lus10042746 pacid=23181819 polypeptide=Lus10042746 locus=Lus10042746.g ID=Lus10042746.BGIv1.0 annot-version=v1.0
MITYVRQVWDSDLRKELAVLEQAFPKYPVLSVDCDGFLLKTPRNPTGDLWYSDLKHKVDQTHITQLSVALSDHGGTVFCTWKFNFEFDVDEDVHTKDTVA
FLKGQGIDFQKHRADGIKRTRFGAQFCNLMASKTTPLTWVTFHGLYDFAHILKVLTNNTPMPGTPSGFRDLLGYSFEEIVDLKVMAKFCDGISDMEEVGL
QKLADLLSIEREDGEALHEAGSDSLLTAMVFSNMRTRFQADQHHVYSDYIYGITERIGRSPTIQVRYSHRWLCQPLLPLPIPCYRPRYHPYPRPPRRCLL
LPLHTPSRMGQY
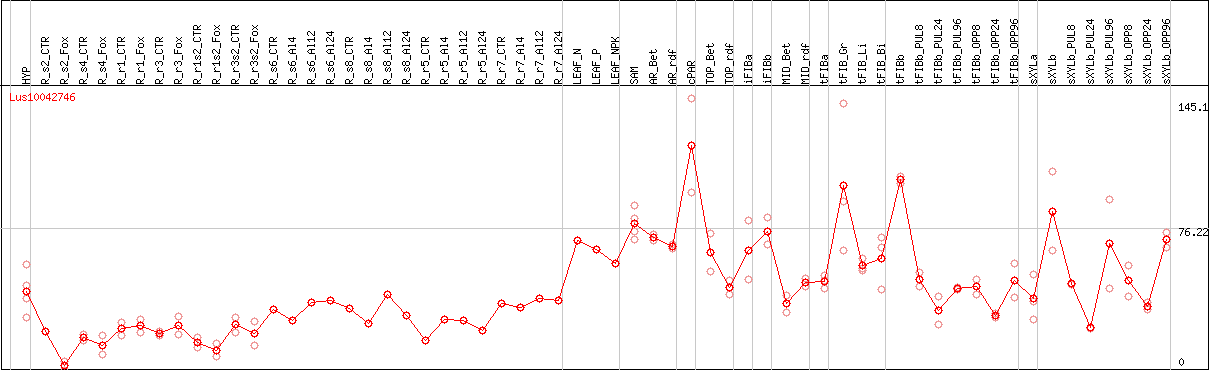
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10042746 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.