External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G77290 348 / 5e-122
Glutathione S-transferase family protein (.1.2)
AT3G62760 57 / 9e-10
ATGSTF13
Glutathione S-transferase family protein (.1)
AT1G49860 55 / 1e-08
ATGSTF14
glutathione S-transferase (class phi) 14 (.1)
AT2G30860 52 / 1e-07
GLUTTR, ATGSTF7, ATGSTF9
glutathione S-transferase PHI 9 (.1.2)
AT2G30870 51 / 1e-07
ERD13, ATGSTF4, ATGSTF10
EARLY DEHYDRATION-INDUCED 13, ARABIDOPSIS THALIANA GLUTATHIONE S-TRANSFERASE PHI 10, glutathione S-transferase PHI 10 (.1)
AT1G02940 50 / 3e-07
ATGSTF5
glutathione S-transferase (class phi) 5 (.1)
AT5G17220 49 / 1e-06
GST26, TT19, ATGSTF12
TRANSPARENT TESTA 19, GLUTATHIONE S-TRANSFERASE 26, ARABIDOPSIS THALIANA GLUTATHIONE S-TRANSFERASE PHI 12, glutathione S-transferase phi 12 (.1)
AT1G02920 46 / 5e-06
ATGST11, GST11, ATGSTF8, ATGSTF7
ARABIDOPSIS GLUTATHIONE S-TRANSFERASE 11, glutathione S-transferase 7 (.1)
AT1G02930 46 / 9e-06
ATGST1, GST1, ERD11, ATGSTF3, ATGSTF6
EARLY RESPONSIVE TO DEHYDRATION 11, ARABIDOPSIS THALIANA GLUATIONE S-TRANSFERASE F3, ARABIDOPSIS GLUTATHIONE S-TRANSFERASE 1, glutathione S-transferase 6 (.1.2)
AT2G47730 45 / 2e-05
GST6, ATGSTF5, ATGSTF8
Arabidopsis thaliana glutathione S-transferase phi 8, GLUTATHIONE S-TRANSFERASE \(CLASS PHI\) 5, glutathione S-transferase phi 8 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10029723
483 / 4e-175
AT1G77290 392 / 3e-139
Glutathione S-transferase family protein (.1.2)
Lus10020735
71 / 2e-14
AT2G30860 334 / 4e-118
glutathione S-transferase PHI 9 (.1.2)
Lus10029815
70 / 3e-14
AT2G30860 332 / 2e-117
glutathione S-transferase PHI 9 (.1.2)
Lus10020736
64 / 7e-12
AT2G30860 308 / 6e-108
glutathione S-transferase PHI 9 (.1.2)
Lus10029816
63 / 1e-11
AT2G30860 308 / 1e-107
glutathione S-transferase PHI 9 (.1.2)
Lus10001419
58 / 9e-10
AT2G30860 297 / 3e-103
glutathione S-transferase PHI 9 (.1.2)
Lus10023511
52 / 9e-08
AT5G17220 240 / 2e-80
TRANSPARENT TESTA 19, GLUTATHIONE S-TRANSFERASE 26, ARABIDOPSIS THALIANA GLUTATHIONE S-TRANSFERASE PHI 12, glutathione S-transferase phi 12 (.1)
Lus10004151
52 / 1e-07
AT2G30860 194 / 1e-62
glutathione S-transferase PHI 9 (.1.2)
Lus10040393
48 / 2e-06
AT3G03190 263 / 1e-89
ARABIDOPSIS GLUTATHIONE-S-TRANSFERASE 6, glutathione S-transferase F11 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G182000
397 / 3e-141
AT1G77290 414 / 4e-148
Glutathione S-transferase family protein (.1.2)
Potri.002G015200
54 / 2e-08
AT3G03190 208 / 4e-68
ARABIDOPSIS GLUTATHIONE-S-TRANSFERASE 6, glutathione S-transferase F11 (.1)
Potri.017G138800
54 / 2e-08
AT5G17220 265 / 1e-90
TRANSPARENT TESTA 19, GLUTATHIONE S-TRANSFERASE 26, ARABIDOPSIS THALIANA GLUTATHIONE S-TRANSFERASE PHI 12, glutathione S-transferase phi 12 (.1)
Potri.002G015100
51 / 1e-07
AT3G03190 196 / 2e-63
ARABIDOPSIS GLUTATHIONE-S-TRANSFERASE 6, glutathione S-transferase F11 (.1)
Potri.014G132200
49 / 6e-07
AT3G62760 295 / 2e-102
Glutathione S-transferase family protein (.1)
Potri.002G207672
49 / 6e-07
AT3G62760 302 / 2e-105
Glutathione S-transferase family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0172
Thioredoxin
PF02798
GST_N
Glutathione S-transferase, N-terminal domain
CL0497
GST_C
PF00043
GST_C
Glutathione S-transferase, C-terminal domain
Representative CDS sequence
>Lus10042756 pacid=23181743 polypeptide=Lus10042756 locus=Lus10042756.g ID=Lus10042756.BGIv1.0 annot-version=v1.0
ATGCAGCTATACCACCATGCTTTCTCCATAGATAGCCAGAAAGTGAGGCTAGCTTTGGAAGAGAAGGGGATTGACTATACTTCAAACCATGTGAATCCTA
TAACAGCCAAGAATCTGGATGCTTCGTTTTTCAGGATGAACCCTAGTGCAAGGCTCCCCGTCTTCCAAAATGGTTCCCATATCATCTTCGAAACAATCGA
GATAATCCAGTACATAGAAAGGATCGCATTCGTCTCCACAGGCGCCAACGACATGCTATTCAGCAACAGAGAAGTTGTTGAATGGATGAAGAAGATACAG
AAATGGAACCCCAAGTTCTTTACTCTTTCACATGTCCCGGAAAAATACCGCCTCCACGTCTCCAAGTTCACAAGGCGAGTGCTGATAGCTCGGATGGCCG
AATCTTCTGATCTAGCCAGTGCTTACCACCGGAAGCTGAAAGAGACGTACGAAACAGAAGAGAAGCTGAAAACTCCGGAAGTTGTAAAACGAAGCACTGA
GCACCTAGTTAGACTTCTTGATGAAGTAGAGACAAAGCTAGGGGAAACAGCGTACTTAGCTGGCGAAGACTTTTCATTGGCAGATGTAATGCTGATCCCG
GTGCTTGCGAGGTTGGTACTGTTGAATTTGGAAGACGAGTATATACGAACGAGGCCGAACGTAGCTGAGTACTGGGAAGTGGTGAAGCAAAGACCGAGTT
ATAGAAACGTGATTGGGAAGTACTTCAATGGATGGAGGAAATACAAGACGTTGATGAAAACTTGGTGTTTGGTCCGTTTCAGAAGTCAGTTCAAGATATA
CTGA
AA sequence
>Lus10042756 pacid=23181743 polypeptide=Lus10042756 locus=Lus10042756.g ID=Lus10042756.BGIv1.0 annot-version=v1.0
MQLYHHAFSIDSQKVRLALEEKGIDYTSNHVNPITAKNLDASFFRMNPSARLPVFQNGSHIIFETIEIIQYIERIAFVSTGANDMLFSNREVVEWMKKIQ
KWNPKFFTLSHVPEKYRLHVSKFTRRVLIARMAESSDLASAYHRKLKETYETEEKLKTPEVVKRSTEHLVRLLDEVETKLGETAYLAGEDFSLADVMLIP
VLARLVLLNLEDEYIRTRPNVAEYWEVVKQRPSYRNVIGKYFNGWRKYKTLMKTWCLVRFRSQFKIY
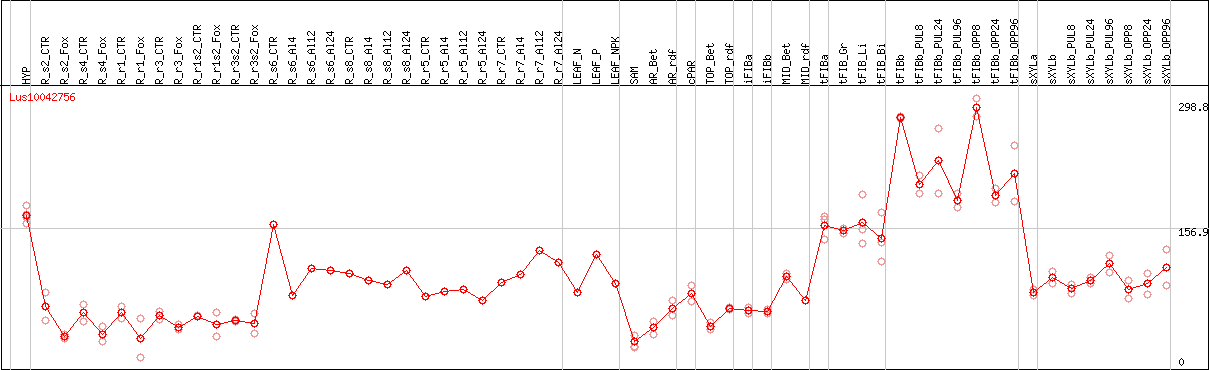
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10042756 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.