Lus10042807 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10042807 pacid=23182011 polypeptide=Lus10042807 locus=Lus10042807.g ID=Lus10042807.BGIv1.0 annot-version=v1.0
ATGTTGCACGAGCTTCTACTGGCACTTTTAGGTTACACCGGAGATCTCGTCGTCGACGAAAGAGACCACCAGTCCTCTCTCGGGGTTAACCCTCCCATCT
CCGACCACCGTTCCTTCAAGCTCGCTCCAGACCTCTCTTTTCTCCACTCTTCTGACAGGGATCTTGTGGAAAGGATCATTGCATTAGGGTTCTATTACAG
AGAGCTTGATCGCTTCGCTTCCAAATCTCGCAACTTGAGTTGCGTTAGGTCTTCGTATGCGTTTCCTCCAGTGGCGGAGTTGTCCAACAGCGGGACAGAA
AAACAACAGCAGAGCGTCTACAGGAGGGCTATAGCTAATGGGATCGTGGAGATACTCTCCATTTACAGGTCTGCCGTTCTTCACATTGAACAGAAATTAT
TGTCCGAGACAGTGCCTATCCTTGCTACACTCACTCAAGGCCTGAACAAGTTTTTTGTCCTGTTACCACCTCTGTATGAGCTAGTTTTGGAGATTGAACG
TGAAAATGTTCGAGGGGGACAGCTTCTTACCCTCTTACATAAACGGTGCCATTGTGGCGTTCCTGAACTACAGACATGCATTCAAAGGCTTCTTTGGCAT
GGGCATCAAGTCATGTATAACCAGCTTGCTTCATGGATGGTCTATGGGATCCTTCAAGACCACCATGATGAATTTTTCATCAGAAGGTATGAGAAGAGTA
ATCCTTGTTGGGGTAGAGTACGGATTCTAGTCTCTAGATATCAAGTAGGAAGGTTAGAAAGGAAGGATGAAAAGCCCAGTTCCTCTGATCATGAAACATC
AGAAAGATACATGCGTTTGTCAACGGATGATGCATCTTTGACTGATTGGCACTCAGGATTTGAGACTTGTCTGGATATGTTGCCCTCGTATATCCCTACA
AATATTGCCGATTCAATTCTTTTTGCTGGTAAAGCCATTCGTATTTTGCGGAATCCAAGTCCAACATTCCTATTGAAGGATCCATCAGATAACCAAGAGG
TGATAAAGGGTGGTCAAAAGGTACAGGCATTCATGGGACAGTTTTCCATTCATGAGGAGCCCTTCGTGACAACAACCCATATTGGTGAGGAATTGCTTCC
ACACTCTGAGGCTGATAAGATTGAAGTTATGCTTAAGGACTTAAAGGATTCGTCTGAGTTTCACAAAAGATTGTTTGAAAGTGCTGTTGGATGCATCCGA
GCTGTTGCAGCTAGTCATCTTTGGCAGCTTGTGGTTGTGCGTGCTGACTTGAATGGTCATCTGAAGGCTCTCAAGGATTATTTCCTGCTTGCTAAAGGAG
ACTTTTTCCAGTGTTTTCTTGAGGAGGGCCGGCATTTGATGCGCTTGCCACCTCGGCAATCCACTGCTGAAGCCGATCTCATGGTTCCGTTCCAGTTGGC
TTCAATGAAGACAATTGGGGAAGAAGACATATATTTCTCTCGAGTATCCTTGCGGATGGCTTCGTTTGGAACTTCAACGAAATCCTTCATTGATGCAACC
TCAAGCTCTGGACTATCAAGTACTTCGTCTGAGATGTCACTTAGTGGATGGGACGGTATTGCACTTGAATATACTGTTGAATGGCCGTTGCAGTTGTTCT
TTACACAAGAAGTACTGTCTGATTACCTTAAGATATTCCAGTACTTATTGAGGCTCAAGCGGACGCAAATGGAGTTGGACAAATCATGGGCCTTGGGCAT
GCATCAGGATCATAAACGCTTTCAAAAGCGTTGCAACAAGAACATGGGGTCTTCACCATCTCAGCAGCGACAACAACAATTTAGGCCTCTATGGCGTGTC
AGAGAGCATATGGCGTTTTTAATTAGAAATCTCCAGTTTTATATCCAGGTGGACGTGATAGAATCTCAATGGAATGTTTTGCAAGCCCATCTTCAAGATT
CCCAAGACTTTACTGAACTTGTCGGCTTCCACCAAGCGTACGTAATCTCTCTAGTTGCCTTGTTAACTGATTTCGCTCACCTTTTCTTCCCTTCTGCGTA
CAGATATTTGTCAGCTTTGATATCACAGTCGTTTGTAGACATTGGTTCTTTGTCAAGGATACTGGACAGCATAATGAAGCTTTGCATGCAATTCTGTTGG
AGCCTTGAGAACCAAGAGAGCAACCCGGATACAAAGGAACTTGAACGTATAACTGAGGAGTTCAATAAGAAATCAAATGCGTTGTACACCATACTGCGGA
GTAGCAGGATAGCAGGGAGTCAACGGGCACCATTTCTGAGACGTTTTCTCATGCGTCTCAACTTCAATTTATTCTTCGAGACAACTGCAATCGGCGTGCT
TAATATTGTTAGGCCGAGCCCTGAGGGGACATGGATAGATAGAGCCACGAACATGTTGACGGAAGAAGAAATGATGAACGTTCCGATTTCTGGAGCGAGA
GAAAGAGAAACAGAAGAAGAGACGGAGAGCTGTAGTAGAGCTAGAGAGAGCGAAAGATATATCATTTCTGAGAGAAAGAAAAAGAGGGGCAAAAGGGCCA
GGAAGATCGAGGGATGGGTTCGGGAGCTTATCCAGAAGCAGTTTCGATGTAAGGCTGCCTGCAAAGAAATCGTTAGAAATCCCGAATTCTCTGGGAAGAT
TACTTTCCCAGTTTCCTTGAAGCAGCCAGGGCACAGGGATGGGCCTATCCAGTGCTTCATTAAGAGAGACAAGTCTAAATTAACTTATCATCTTTTCCTT
TGCCTCAGCCCTGCTTTGCTTGTTGAAAATGGGAAGTTCCTTCTCTCTGCTAAACGTACCAGAAGAACTACGTGCACGGAGTATATAATCTCCATGGATG
CTGATAATATATCAAGATCCAGTGGAACATATATTGGAAAGCTAAGGTCAAATTTCCTAGGTACGAGGTTTATAGTCTACGACACTCAGCCGCCCTACAA
CAACGCTCAAATATCGCCTCCCGGTCGAAGCAAGAGATTCTACTCCAGAAAAGTTTCCCCAAAGGTCCCCACGGGAAACTACAACATTTCTCAGATTTCA
TACGAATTGAACGTGCTCGGGACAAGAGGTCCACGGAGGATGCACTGCACCATGCACTCAATTCCCACCTCGGCGCTCGAGTCCGGCGGTGCAGTCCCGG
GCCAACCTGAGATGATCCTTCCGAGGTCCCTCGAGGAGTCATTCAGAAGCATGTCTCTCTCAAAATCAATCATCGACAGCTCGAGCGAGTTTAGCAGCTC
CCGGTTCTTCGATGTTGCTGGACCCCCTCATGAAGGGGGAGAAGAAGAAGAGGAAGGGAAGGAGAGAGCGCTGATTCTGAGGAACAAAGCACCAAGGTGG
CATGAGCAGTTGCAGTGTTGGTGTCTGAACTTTCGTGGCAGAGTCACTGTAGCGTCTGTGAAGAATTTCCAGCTGATAGCTGCGAACCAGCCTGCTGCTG
CTGCTGCTGCTGTTGCACCGATGTCATCAGCCCAACCGGCCCAGTCGGATCACGATAAGGTCATATTGCAGTTTGGGAAGGTGGGGAAGGATATGTTCAC
TATGGATTACCGGTATCCGCTGTCTGCTTTTCAAGCGTTCGCAATCTGCTTGAGCAGCTTTGACGCTAAGCTGGCATGTGAATAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10042807 pacid=23182011 polypeptide=Lus10042807 locus=Lus10042807.g ID=Lus10042807.BGIv1.0 annot-version=v1.0
MLHELLLALLGYTGDLVVDERDHQSSLGVNPPISDHRSFKLAPDLSFLHSSDRDLVERIIALGFYYRELDRFASKSRNLSCVRSSYAFPPVAELSNSGTE
KQQQSVYRRAIANGIVEILSIYRSAVLHIEQKLLSETVPILATLTQGLNKFFVLLPPLYELVLEIERENVRGGQLLTLLHKRCHCGVPELQTCIQRLLWH
GHQVMYNQLASWMVYGILQDHHDEFFIRRYEKSNPCWGRVRILVSRYQVGRLERKDEKPSSSDHETSERYMRLSTDDASLTDWHSGFETCLDMLPSYIPT
NIADSILFAGKAIRILRNPSPTFLLKDPSDNQEVIKGGQKVQAFMGQFSIHEEPFVTTTHIGEELLPHSEADKIEVMLKDLKDSSEFHKRLFESAVGCIR
AVAASHLWQLVVVRADLNGHLKALKDYFLLAKGDFFQCFLEEGRHLMRLPPRQSTAEADLMVPFQLASMKTIGEEDIYFSRVSLRMASFGTSTKSFIDAT
SSSGLSSTSSEMSLSGWDGIALEYTVEWPLQLFFTQEVLSDYLKIFQYLLRLKRTQMELDKSWALGMHQDHKRFQKRCNKNMGSSPSQQRQQQFRPLWRV
REHMAFLIRNLQFYIQVDVIESQWNVLQAHLQDSQDFTELVGFHQAYVISLVALLTDFAHLFFPSAYRYLSALISQSFVDIGSLSRILDSIMKLCMQFCW
SLENQESNPDTKELERITEEFNKKSNALYTILRSSRIAGSQRAPFLRRFLMRLNFNLFFETTAIGVLNIVRPSPEGTWIDRATNMLTEEEMMNVPISGAR
ERETEEETESCSRARESERYIISERKKKRGKRARKIEGWVRELIQKQFRCKAACKEIVRNPEFSGKITFPVSLKQPGHRDGPIQCFIKRDKSKLTYHLFL
CLSPALLVENGKFLLSAKRTRRTTCTEYIISMDADNISRSSGTYIGKLRSNFLGTRFIVYDTQPPYNNAQISPPGRSKRFYSRKVSPKVPTGNYNISQIS
YELNVLGTRGPRRMHCTMHSIPTSALESGGAVPGQPEMILPRSLEESFRSMSLSKSIIDSSSEFSSSRFFDVAGPPHEGGEEEEEGKERALILRNKAPRW
HEQLQCWCLNFRGRVTVASVKNFQLIAANQPAAAAAAVAPMSSAQPAQSDHDKVILQFGKVGKDMFTMDYRYPLSAFQAFAICLSSFDAKLACE
|
DESeq2's median of ratios [FLAX]
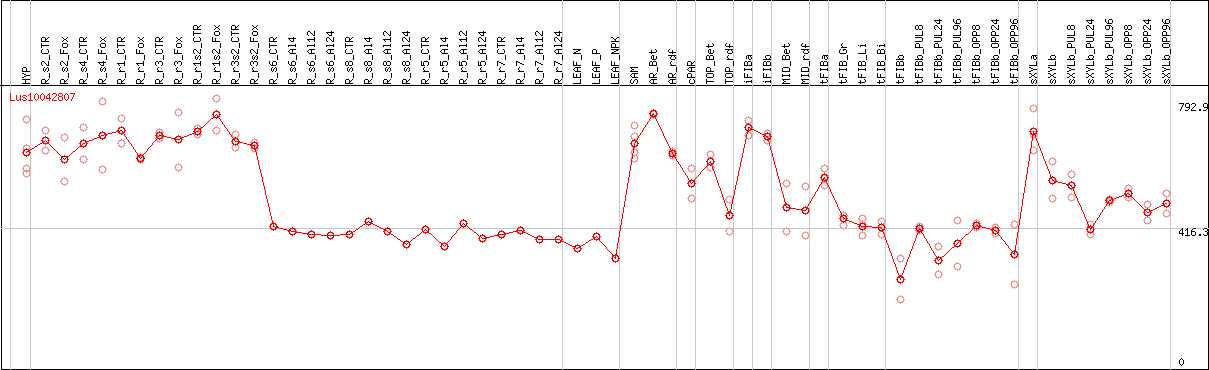
Coexpressed genes
Lus10042807 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
