External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G24470 130 / 1e-35
GATA
GATA25, TIFY1, ZIM
Zinc-finger protein expressed in Inflorescence Meristem, GATA TRANSCRIPTION FACTOR 25, GATA-type zinc finger protein with TIFY domain (.1.2.3)
AT3G21175 119 / 1e-31
GATA
GATA24, TIFY2B, ZML1
ZIM LIKE 1, GATA TRANSCRIPTION FACTOR 24, ZIM-like 1 (.1.2.3)
AT1G51600 119 / 2e-31
GATA
GATA28, TIFY2A, ZML2
GATA TRANSCRIPTION FACTOR 28, ZIM-LIKE 2 (.1.2)
AT2G46670 50 / 3e-07
CCT motif family protein (.1)
AT2G46790 50 / 1e-06
TL1, APRR9
TOC1-LIKE PROTEIN 1, Arabidopsis pseudo-response regulator 9, pseudo-response regulator 9 (.1.2)
AT5G02810 50 / 1e-06
APRR7, PRR7
pseudo-response regulator 7 (.1)
AT5G60100 47 / 9e-06
APRR3
pseudo-response regulator 3 (.1.2.3)
AT5G24470 46 / 2e-05
APRR5
pseudo-response regulator 5 (.1)
AT5G61380 45 / 5e-05
AtTOC1, TOC1, APRR1, ARR1
PSEUDO-RESPONSE REGULATOR 1, TIMING OF CAB EXPRESSION 1, CCT motif -containing response regulator protein (.1)
AT4G14713 42 / 0.0003
ZIM
TIFY4A, PPD1
PEAPOD 1, TIFY domain/Divergent CCT motif family protein (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10028166
275 / 3e-88
AT4G24470 233 / 4e-72
Zinc-finger protein expressed in Inflorescence Meristem, GATA TRANSCRIPTION FACTOR 25, GATA-type zinc finger protein with TIFY domain (.1.2.3)
Lus10032839
194 / 2e-60
AT4G24470 231 / 8e-75
Zinc-finger protein expressed in Inflorescence Meristem, GATA TRANSCRIPTION FACTOR 25, GATA-type zinc finger protein with TIFY domain (.1.2.3)
Lus10002412
193 / 4e-60
AT4G24470 231 / 6e-75
Zinc-finger protein expressed in Inflorescence Meristem, GATA TRANSCRIPTION FACTOR 25, GATA-type zinc finger protein with TIFY domain (.1.2.3)
Lus10023230
53 / 2e-07
AT5G24470 300 / 2e-92
pseudo-response regulator 5 (.1)
Lus10019077
52 / 3e-07
AT5G61380 491 / 2e-168
PSEUDO-RESPONSE REGULATOR 1, TIMING OF CAB EXPRESSION 1, CCT motif -containing response regulator protein (.1)
Lus10008878
52 / 4e-07
AT5G24470 296 / 8e-91
pseudo-response regulator 5 (.1)
Lus10015720
51 / 8e-07
AT5G61380 470 / 3e-160
PSEUDO-RESPONSE REGULATOR 1, TIMING OF CAB EXPRESSION 1, CCT motif -containing response regulator protein (.1)
Lus10000566
50 / 8e-07
AT5G61380 433 / 5e-146
PSEUDO-RESPONSE REGULATOR 1, TIMING OF CAB EXPRESSION 1, CCT motif -containing response regulator protein (.1)
Lus10032934
50 / 1e-06
AT5G24470 356 / 2e-114
pseudo-response regulator 5 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G110900
179 / 2e-54
AT4G24470 253 / 2e-83
Zinc-finger protein expressed in Inflorescence Meristem, GATA TRANSCRIPTION FACTOR 25, GATA-type zinc finger protein with TIFY domain (.1.2.3)
Potri.005G152800
175 / 4e-53
AT4G24470 264 / 1e-87
Zinc-finger protein expressed in Inflorescence Meristem, GATA TRANSCRIPTION FACTOR 25, GATA-type zinc finger protein with TIFY domain (.1.2.3)
Potri.002G110800
164 / 3e-48
AT3G21175 230 / 1e-73
ZIM LIKE 1, GATA TRANSCRIPTION FACTOR 24, ZIM-like 1 (.1.2.3)
Potri.005G152500
156 / 5e-45
AT3G21175 208 / 4e-65
ZIM LIKE 1, GATA TRANSCRIPTION FACTOR 24, ZIM-like 1 (.1.2.3)
Potri.007G116700
131 / 2e-35
AT3G21175 155 / 3e-44
ZIM LIKE 1, GATA TRANSCRIPTION FACTOR 24, ZIM-like 1 (.1.2.3)
Potri.017G042200
130 / 7e-35
AT3G21175 171 / 1e-50
ZIM LIKE 1, GATA TRANSCRIPTION FACTOR 24, ZIM-like 1 (.1.2.3)
Potri.010G251600
122 / 1e-32
AT1G51600 298 / 9e-101
GATA TRANSCRIPTION FACTOR 28, ZIM-LIKE 2 (.1.2)
Potri.007G116550
122 / 2e-32
AT3G21175 228 / 3e-73
ZIM LIKE 1, GATA TRANSCRIPTION FACTOR 24, ZIM-like 1 (.1.2.3)
Potri.015G061900
54 / 4e-08
AT5G61380 474 / 2e-161
PSEUDO-RESPONSE REGULATOR 1, TIMING OF CAB EXPRESSION 1, CCT motif -containing response regulator protein (.1)
Potri.010G215200
54 / 6e-08
AT5G02810 441 / 2e-144
pseudo-response regulator 7 (.1)
PFAM info
Representative CDS sequence
>Lus10042865 pacid=23181940 polypeptide=Lus10042865 locus=Lus10042865.g ID=Lus10042865.BGIv1.0 annot-version=v1.0
ATGGATATTCCTCACCAGATTGACGTCGTCCCCGGTCCCGGCGAAGACGACGATCCTGCTGGGGCACCGGATTGTATGGGCCAGCAAAACCACATTGGAT
ATGATGACGCTTCCGCCGCCGCCGCTGGAATCGTCGTCGACAACATGTCTCCTGACTCCGTATATGTTTCTACCGGTGGTGGCCCTGGCTCGGAGTTTGG
TTTCCAGGGCGGGGATATCACCAACCAGCTTACTTTGACGTTTCGTGGCCAGGTTTATGTGTTTGACGACATCACTCCAGACAGGGTTCAGGCAGTGCTA
TTACTTCTAGGAGGGTGTGAAGTAAATGCAGGTCTGTATGGCATGGATGTCGCACCTCAAAGCCAAAAGGGCCAGGTGATGGAATATCCAGGGCGTTCTA
CACAACCTCATAGAGAAGCATCGTTAAATAGATTTAGACAGAAGAGGAAAGAACGATGTTTTGATAAAAAAGTCAGATATGGCGTTCGGCAACAGGTTGC
ACAAAGGCCATGGGGGATTTCTCCAAAACGCTCTAATCTTTCGCGTCGAATAGCATCCCTTGTCAGTTTCCGGGAGAAGAGGAAGGAGCAATGTCTCGAC
AAGAAAATACGATACACTGTCCGAAAAGAAGTTGCACAGAGGCTTAGGAGGTGTCAACATTGTGGTGTAAGTGAGAAGAATACTCCTGCAATGCGACGTG
GACCAGCTGGAAGAATACTCCTGCAATGCGACGTGGACCAGCTGGACCAAGAACTTTATGTAATGCATGCGGCCTCATGTGGGCAAATACGGTCTGGTAC
CCTCAGAGATCTCAGTAAAGGTGGAAGGAGTCTTCCTGCAGATCAAGTTGAACCTGTATGTAAAGAGTGGCTACTCATGTACTTTTATATTTGCTTCTGG
GATTTCTCAGCATGA
AA sequence
>Lus10042865 pacid=23181940 polypeptide=Lus10042865 locus=Lus10042865.g ID=Lus10042865.BGIv1.0 annot-version=v1.0
MDIPHQIDVVPGPGEDDDPAGAPDCMGQQNHIGYDDASAAAAGIVVDNMSPDSVYVSTGGGPGSEFGFQGGDITNQLTLTFRGQVYVFDDITPDRVQAVL
LLLGGCEVNAGLYGMDVAPQSQKGQVMEYPGRSTQPHREASLNRFRQKRKERCFDKKVRYGVRQQVAQRPWGISPKRSNLSRRIASLVSFREKRKEQCLD
KKIRYTVRKEVAQRLRRCQHCGVSEKNTPAMRRGPAGRILLQCDVDQLDQELYVMHAASCGQIRSGTLRDLSKGGRSLPADQVEPVCKEWLLMYFYICFW
DFSA
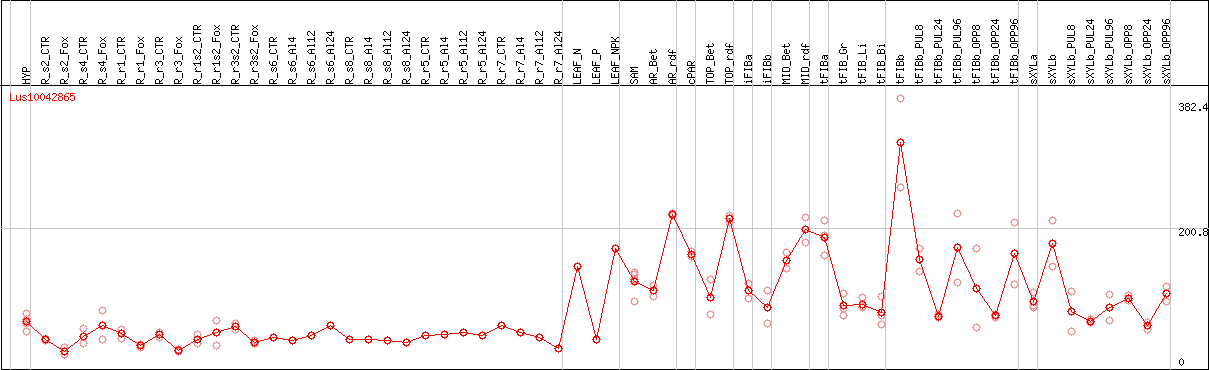
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10042865 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.