Lus10042915 [FLAX]

| External link |
|
|||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||
|
Paralogs
|
|
|||||||||||||||
|
Poplar homologues |
|
|||||||||||||||
| PFAM info |
| |||||||||||||||
|
Representative CDS sequence |
>Lus10042915 pacid=23182037 polypeptide=Lus10042915 locus=Lus10042915.g ID=Lus10042915.BGIv1.0 annot-version=v1.0
ATGCAATGCAGCAACACGGAGGTTGGAGCTGGGACTATGAATCCTCTCACTTTCCTGAGGGTCCTCGGTCCAGAGCCATGGAATGTGGCGTATGTAGAGC
CTAGTATTCGGCCTGATGATAGTCGCTATGGAGAAAATCCTAATAGGCTTCAGCGCCATACTCAATTTCAGGTTATTTTGAAGCCTGATCCTGGAAATTC
ACAAGACCTTTTCATTCGCAGTCTGTCAGCTTTAGGCATCAATGTTAATGACCATGATATACGGTTTGTGGAGGACAACTGGGAAAGCCCTGTACTTGGT
GCTTGGGGTTTAGGCTGGGAAATATGGATGGATGGGATGGAAATAACTCAATTCACGTATTTTCAGCAGGCTGGAAGTCTTCAGTTGTCACCAATATCTG
TTGAAATCACTTACGGCCTTGAGCGCATACTCATGTTGCTTCAGGGAGTTGATCATTTCAAAAAGATACAGTATGCTGATGGTATTACCTATGGCGAGTT
GTTCTCCGAAAATGAGAAGGAAATGAGTGCATATTATCTGGAACATGCTAGTGTTCAGCAGCTTCAGAATCACTTTGATTTATTTGAGGCAGAAGCACGT
TCTTTGCTTGCTTTGGGTTTGGCCATTCCTGCGTATGATCAGCTTCTCAAGACATCTCACGCTTTCAATATTTTGGACTCTAGAGGATTTGTCGGTGTGA
CTGAGCGTGCTCGTTATTTTGGTCGGATGCGTAGTTTAGCTCGTCAGTGTGCTGAACTTTGGTTGAAGGCCAGGGAGTCGCTGGGATACCCATTGGGTAT
CGTATCTGAACCTGCTGATTCTACATGTTCTCAACAGTTTGTGGAGGTGGAAGACAACAACATGATTCTCAATGACAAAAGGGCATTTGTTCTTGAAATT
GGCACTGAAGAGTTGCCTCCACTTGATGTAGTTCATGGCACCGAGCAGCTTGAAAGTTTAATGTTTAAATTACTAGAGAAACAAAGATTAGGCCATGGCA
AGATTCAAGTGTTTGGCACCCCTCGCAGATTGGTGGTCTTGGTGGAGAACCTTTGCTCTAAGCAGTCAGAAAGTGAGGTCGAGGTTAGAGGTCCTCCAGT
TTCAAAGGCATTTGATGAGCAGGGAAATCCTACAAAGGCTGCCGAGGGCTTTTGTCGTAGATACGATGTATCGCTGGACTCTTTGTTTAGAAAGATTGAT
GGGAAAACTGAGTACATTCATGCCCTTGTAAAGAAGACTACTCGGAGGGCATCAGAGGTTATCTTGAGTGAAGATTTACCTGGTATAATCAGTAAATTAT
CATTCCCAAAGTCCATGCGTTGGAACTCTCAGGTCATGTTCAGCAGGCCAATTCGCTGGGTTATGGCCTTGCATGGACCCACTGTTGTTCCATTCACATT
TGTGGGAGTTTCAAGTGGAAACCTATCGTATGGTCTTCGCAACTACCCTTCATCTACAATTCAGGTCAAGAGTGCCGAATCCTATGCAAGTGCATTGATA
ACGGCTGGAATTAATATTGTTATCGAGGATCGAAAGAAAAGCATCTTGGAGCAGTCAAATTTGTTGGCAGAAAGTGTCAATGGTCGTCTCTTGGTGCAGG
AAAACTTGCTTCATGAGGTAGTTAATCTTGTCGAGGCACCAGTACCCGTGCTTGGAAAGTTTAAAGAGTCATTTTTGGAGCTTCCAGATGATCTCTTAAC
AATGGTCATGCAGAAGCACCAGAAGTATTTCGCAATCACTGATGATGGTGGAAAGCTACTTCCTTACTTCATTGCCGTAGCAAATGGGCCCATAAACGAG
ATGGTGGTTAGAGAAGGGAATGAAGCTGTGCTTAGAGCACGCTATGAAGATGCTAAATTTTTCTATGAGATGGATACTCGAAGGAGTTTCTCGGAGTTTC
AAAGTCAGTTGAAAGGGATACTTTTCCATGAGAAGCTCGGGACAATGTTTGACAAGGTAGTTCGCATGCAAAAAACGGTTGGAAATTTGAGTTTGGAGTT
GGGTAATGGTGAAGTCATGTTTCAAACTGTCAAGGATGCCACGTCACTAGCAATGTCGGATCTTGCTACTGCAGTAGTAACAGAATTCACTGCGCTTTCT
GGAATAATGGCACGCCATTATGCCCTAAGAGATGGATATTCGGAGCAGATTGCGGAGGCATTGTTCGAAATCACCCTTCCAAGATTTTCTGGCGACGTTC
TTCCAAAAAGTGACGTGGGGATTGTTTTGGCAATTGCTGACAGATTAGACAGCCTTGTTGGTTTGTTTGCTGCCGGATGCCGGCCTAGTTCTACTAATGA
CCCATTTGGACTGAGAAGAATCTCATATGGCCTGGTTCAGATATTGGTGGAAAGGGAGAAAGATTTGGATTTGCATCGCGCTATAACACTTGCTGCAGCT
GTACAACCTATACCTGTTGATGCTCATATAATAGATGACGCACATGCATTTGTTACTCGAAGATTAGAGCAATTCTTGGTCGACAAAGGAATAAGTTCAG
AATTGGTCCGTTCTGCCCTGGCTGAGCGTGGGAATTCCCCATATCTGGCAGCTAAAACAGCATCCAAAGTGCTGATAGATACTCTACAGATGCAAATGTT
ATCAGAAAGTCATCTATTCCCTAAGGTGGTTGAAGCATACTCTCGACCCACTAGAATAGTTCGAGGAAAGGACTTTGATTCCAGCTACCAGAAATTCAAC
TTGGGTATTGACAAATTCCCTTCGTTCTGCAAGGTGAATGACACAAAATTCGAGACAAATGAAGAGAGAGCCTTGTGGAAGACTTTCTTGTCCACTAGAA
GCAAAATTCACCCTGGTATCGAGGTTGACGAATTCGTTGAAGTTTCTTCGGAGCTGGTGCAGCCTTTGGAGGATTTCTTCAATAATGTCTTCGTTATGGT
GGAAGACGAAAGCGTTCGAAACAACAGGCTTGCCCTACTTGCGCAAATTGCGGATCTTCCCAGAGGCATTGCGGACCTGTCACTCTTGCCAGGGTTTTAG
|
|||||||||||||||
|
AA sequence
|
>Lus10042915 pacid=23182037 polypeptide=Lus10042915 locus=Lus10042915.g ID=Lus10042915.BGIv1.0 annot-version=v1.0
MQCSNTEVGAGTMNPLTFLRVLGPEPWNVAYVEPSIRPDDSRYGENPNRLQRHTQFQVILKPDPGNSQDLFIRSLSALGINVNDHDIRFVEDNWESPVLG
AWGLGWEIWMDGMEITQFTYFQQAGSLQLSPISVEITYGLERILMLLQGVDHFKKIQYADGITYGELFSENEKEMSAYYLEHASVQQLQNHFDLFEAEAR
SLLALGLAIPAYDQLLKTSHAFNILDSRGFVGVTERARYFGRMRSLARQCAELWLKARESLGYPLGIVSEPADSTCSQQFVEVEDNNMILNDKRAFVLEI
GTEELPPLDVVHGTEQLESLMFKLLEKQRLGHGKIQVFGTPRRLVVLVENLCSKQSESEVEVRGPPVSKAFDEQGNPTKAAEGFCRRYDVSLDSLFRKID
GKTEYIHALVKKTTRRASEVILSEDLPGIISKLSFPKSMRWNSQVMFSRPIRWVMALHGPTVVPFTFVGVSSGNLSYGLRNYPSSTIQVKSAESYASALI
TAGINIVIEDRKKSILEQSNLLAESVNGRLLVQENLLHEVVNLVEAPVPVLGKFKESFLELPDDLLTMVMQKHQKYFAITDDGGKLLPYFIAVANGPINE
MVVREGNEAVLRARYEDAKFFYEMDTRRSFSEFQSQLKGILFHEKLGTMFDKVVRMQKTVGNLSLELGNGEVMFQTVKDATSLAMSDLATAVVTEFTALS
GIMARHYALRDGYSEQIAEALFEITLPRFSGDVLPKSDVGIVLAIADRLDSLVGLFAAGCRPSSTNDPFGLRRISYGLVQILVEREKDLDLHRAITLAAA
VQPIPVDAHIIDDAHAFVTRRLEQFLVDKGISSELVRSALAERGNSPYLAAKTASKVLIDTLQMQMLSESHLFPKVVEAYSRPTRIVRGKDFDSSYQKFN
LGIDKFPSFCKVNDTKFETNEERALWKTFLSTRSKIHPGIEVDEFVEVSSELVQPLEDFFNNVFVMVEDESVRNNRLALLAQIADLPRGIADLSLLPGF
|
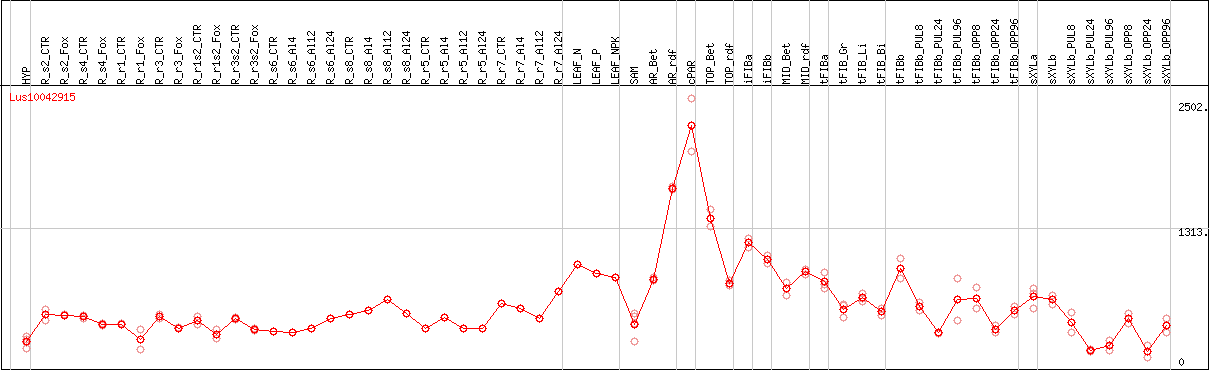
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10042915 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
