External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G19050 152 / 8e-47
ARR7
response regulator 7 (.1)
AT1G74890 150 / 5e-46
ARR15
response regulator 15 (.1)
AT3G48100 139 / 8e-42
ATRR2, IBC6, ARR5
INDUCED BY CYTOKININ 6, ARABIDOPSIS THALIANA RESPONSE REGULATOR 2, response regulator 5 (.1)
AT5G62920 135 / 3e-40
ARR6
response regulator 6 (.1)
AT1G59940 135 / 6e-40
ARR3
response regulator 3 (.1)
AT1G10470 135 / 3e-39
IBC7, ATRR1, ARR4, MEE7
maternal effect embryo arrest 7, INDUCED BY CYTOKININ 7, RESPONCE REGULATOR 1, response regulator 4 (.1)
AT3G57040 117 / 2e-32
ATRR4, ARR9
RESPONSE REGULATOR 4, response regulator 9 (.1)
AT2G41310 116 / 2e-32
ARR8, ATRR3
RESPONSE REGULATOR 8, response regulator 3 (.1)
AT3G56380 114 / 2e-32
ARR17
response regulator 17 (.1)
AT2G40670 107 / 9e-30
ARR16
response regulator 16 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10036642
292 / 6e-102
AT1G19050 160 / 6e-50
response regulator 7 (.1)
Lus10042153
224 / 7e-75
AT3G48100 213 / 1e-70
INDUCED BY CYTOKININ 6, ARABIDOPSIS THALIANA RESPONSE REGULATOR 2, response regulator 5 (.1)
Lus10004243
218 / 3e-72
AT3G48100 220 / 2e-73
INDUCED BY CYTOKININ 6, ARABIDOPSIS THALIANA RESPONSE REGULATOR 2, response regulator 5 (.1)
Lus10010723
155 / 2e-47
AT1G10470 155 / 1e-46
maternal effect embryo arrest 7, INDUCED BY CYTOKININ 7, RESPONCE REGULATOR 1, response regulator 4 (.1)
Lus10030664
144 / 2e-42
AT1G10470 206 / 2e-65
maternal effect embryo arrest 7, INDUCED BY CYTOKININ 7, RESPONCE REGULATOR 1, response regulator 4 (.1)
Lus10029211
143 / 3e-42
AT1G10470 211 / 4e-68
maternal effect embryo arrest 7, INDUCED BY CYTOKININ 7, RESPONCE REGULATOR 1, response regulator 4 (.1)
Lus10030822
142 / 7e-42
AT1G59940 201 / 2e-64
response regulator 3 (.1)
Lus10039524
122 / 1e-34
AT3G57040 222 / 2e-73
RESPONSE REGULATOR 4, response regulator 9 (.1)
Lus10016334
118 / 4e-33
AT3G57040 211 / 5e-69
RESPONSE REGULATOR 4, response regulator 9 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0304
CheY
PF00072
Response_reg
Response regulator receiver domain
Representative CDS sequence
>Lus10042938 pacid=23168369 polypeptide=Lus10042938 locus=Lus10042938.g ID=Lus10042938.BGIv1.0 annot-version=v1.0
ATGGCGGTCGCTGGTTCTGAATCACTGCGTTGGAGCTTGTCCGAGGAGGTTGGAATTTCGGGGTCTGGTGGAACTGTTTCAGAGGAGTTTCATGTTCTTG
CAGTGGATGATAGTATTGTTGACCGCAAGTTAATCGAGCGCTTACTCAAGAGCTCTTCCTTTAAAGTGACGGCTGTGGAGAGTGGTAGTAGAGCTCTGCA
GTACCTTGGATTGGACGGAGGGAAGAGTGCTGCTGGATTCAATGATTTGAAGGAATCATCGGCTTGTAAAGAGATACCGGTTGTAGTAATGTCGTCAGAG
AACGTTTTGACCCGTATCCATAGTTGTCTGGAGGAAGGAGCAGAGGAATTCTTGATGAAGCCTGTGAAGAAGTCGGATGTTGCAAGGCTTAAAAACTTCA
TAGAGAAGGGCGACCGTGCTGAGAAAACCCAGAGAAAGAGCCTGAAGAGGATGATTGAGGATGGCTTATTTACAACTACATTTTCTTGGTCAGAGGAGGA
GGAGGAGCGATCCCCCTCGTCATCCCCATCACCATCACAATCATCATCTACATGTTCATCACCATCTTCAAAAAGGCCCAAGCAGCAATAG
AA sequence
>Lus10042938 pacid=23168369 polypeptide=Lus10042938 locus=Lus10042938.g ID=Lus10042938.BGIv1.0 annot-version=v1.0
MAVAGSESLRWSLSEEVGISGSGGTVSEEFHVLAVDDSIVDRKLIERLLKSSSFKVTAVESGSRALQYLGLDGGKSAAGFNDLKESSACKEIPVVVMSSE
NVLTRIHSCLEEGAEEFLMKPVKKSDVARLKNFIEKGDRAEKTQRKSLKRMIEDGLFTTTFSWSEEEEERSPSSSPSPSQSSSTCSSPSSKRPKQQ
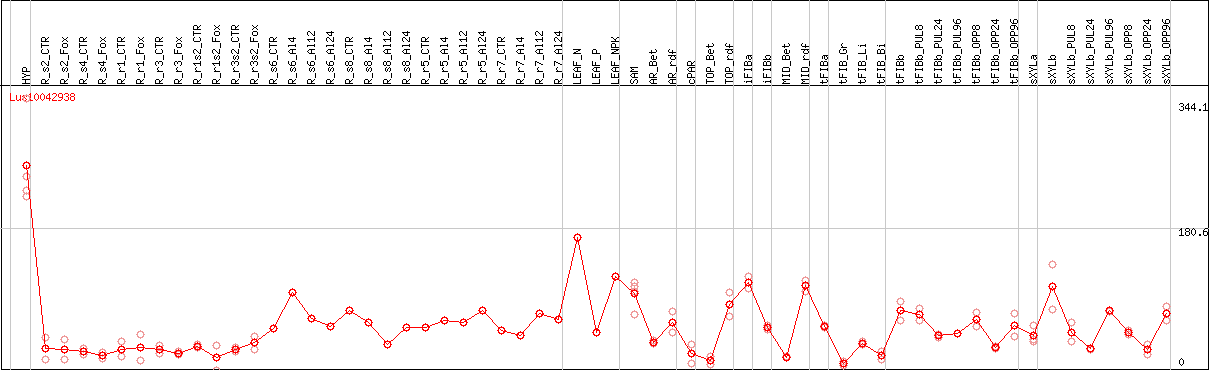
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10042938 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.