Lus10042952 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10042952 pacid=23168400 polypeptide=Lus10042952 locus=Lus10042952.g ID=Lus10042952.BGIv1.0 annot-version=v1.0
ATGACTGCGAAAACCCCTGGGACGCCAGCTTCAAAGATTGAAAACAGAACACCAGCATCAACTCCTGGAGGAGGGAGCAAGGCAAAGGACGAGAAAATCG
TAGTTACTGTACGTCTGCGGCCTTTGAACAGAAGGGAGCAGCTAGCTAAAGATCAAGTGGCGTGGGAGTGTTTGGATGATTACACCATTTCTTACAAGCC
TCCACCCCAGGAGCGCTCTGCTCAACCAGCCTCTTTCACTTTTGATAAAGTGTTTGGCCCAGCTTGTGTAACTGAGTCGGTTTATGAGGATGGAGTAAAG
AACATTGCTCTCTCTGCTCTTATGGGAATAAATGCAACTATCTTTGCTTATGGGCAAACCAGCAGTGGGAAGACATACACGATGAGAGGAATAACTGAAA
AAGCTGTTCAAGACATCTATAACCATATAAGAAACACTCCAGAGAGAGACTTTACAATCAAGATGTCAGGACTTGAGATATATAACGAAAATGTCAGGGA
TTTGTTGAATTCAGAATCTGGCCGTAGCCTCAAACTTCTTGATGATCCGGAGAAAGGTACCGTGGTTGAGAAACTTGTAGAGGAAACAGCAAGCAACGAC
TCGCACTTGCGGAAGTTAATTAGTATATGTGAAGCCCAGAGGCAGGTTGGGGAAACTGCACTCAATGATACTAGTTCACGATCTCATCAGATAATACGAT
TGACAATACAAAGTACTCTAAATGAGAGTTCAGATTGTGCGAGGTCTTTTGTTGCAAGCTTGAATTTCGTTGATCTCGCTGGGAGTGAGAGGGCTTCACA
GACGCATACCGATGGTGCTAGGCTACGAGAAGGTTGCCATATCAATCTTAGCCTGATGACTCTGACTACCGTAATCAGAAAGCTCAGTGTTGGGAAAAGA
AGTTGTCACATACCCTACAGAGACTCCAAGCTTACACGCATACTGCAGCATTCGCTAGGTGGAAATGCTCGTACTGCCATTATCTGTACTTTAAGCCCGG
CATTGACACATGTTGAACAATCTCGAAATACACTTTACTTCGCAACACGAGCTAAGGAAGTTACAAACAGTGCCAAAGTTAACATGGTTATTTCGGACAA
GCAGCTAGTGAAGCATTTACAGAAGGAAGTAGCGAGGCTGGAAGCAGAACGGCGCACTCCTGACCCTTCATCTGAAAAAGACTGGAAAATACAGCAGCTG
GAGATGGAAATTGAGGAACTGAGGCGCCAAAGAGACCTTGCGAAGTGTGAGGTGGACGAGTTACGCAAGAAACTCCAAGATGAAAAACTTGAAGATGGAA
AGATATTCCGCGAAGTTGAATCACCCCACCCACTCGTGAAGAAGTGTCTGTCATACTCAGCAGCAATGACGTCGAAACCTGAGAGCAAGGACTTCATTCA
CTGTGAGAAGATGACAAAAGCAACACTAAGGCAGTCCATGAGACTATCATCAGCTGCACCTTTCACACTGATGCATGAAATTCGCAAGCTCGAACACCTT
CAGGATCAACTTGGGGAAGAAGCTAATCGGGCTCTGGAAGTTTTGCAGAAAGAGGTTGCTTGTCACAGACTAGGTAATCAAGATGCAGCCGAGACAATAG
CTAAACTTCAAGCTGAAATAAGGGAAATGCGCTCAGCTCCGCTACCAAGGGAAGTTGCAATTGGAAGTAACAAGAGTGTCAGTGCTAATCTGAAGGAAGA
AATAACGAGACTTCATTCGCAAGGAAGCAACATCGCAAATCTCGAAGAGCAACTAGAGAACGTCCAGAAGTCGATTGACAAGCTGATCATGTCACTTCCA
AGTAACAATTCAGATGAGTCAGCAGCTTCCAATCCTAAGAACCAGCACAGGAAGAAAAAGACTCACCCTTTATGTTCAAGCAACGGTGCAAACATGCAGA
ACTTCATTAGATCTCCTTGTGCGCCTTTGGCATCATCGAGGCAAGTACTGGAGAACGAGACAGAAAATAGAGCACCGGAGAGCGATGATGTTGCGTCCAA
AGTGTCCGAGAAACAACAACAGACTCCAATGAAGAGTGAGGAAGGTGGAGATGTGTCGTCAAAGGAAGGGACACCTGGTTCTCGGCGTACAAGCTCAGTC
AACATGAAGAAAATGCAGAAGATGTTCCAAAATGCAGCAGAGGAAAATGTGAGAAGCATTAGAAACTATGTCACAGAGCTGAAGGAACGCGTTGCAAAAC
TCCAATATCAAAAGCAGCTGTTAGTCTGCCAGGTACTAGAACTGGAAGCAAATGAAGCAGCTGGGTATAACTTGGAGGACGAGGAGGAAGCAGCGATTGG
ACCGGTAGAACCACAAGTTCCATGGCATGTGACCTTCAGAGAACAACGGCAGCAGATCATAGAATTATGGGACGTTTGCTTTGTTATTTCGGACAAGCAG
CTAGTGAAGCATTTACAGAAGGAAGTAGCGAGGCTGGAAGCAGAACGGCGCACTCCTGACCCTTCATCTGAAAAAGACTGGAAAATACAGCAGCTGGAGA
TGGAAATTGAGGAACTGAGGCGCCAAAGAGACCTTGCGAAGTGTGAGGTGGACGAGTTACGCAAGAAACTCCAAGATGAAAAACTTGAAGATGGAAAGAT
ATTCCGCGAAGTTGAATCACCCCACCCACTCGTGAAGAAGTGTCTGTCATACTCAGCAGCAATGACGTCGAAACCTGAGAGCAAGGACTTCATTCACTGT
GAGAAGATGACAAAAGCAACACTAAGGCAGTCCATGAGACTATCATCAGCTGCACCTTTCACACTGATGCATGAAATTCGCAAGCTCGAACACCTTCAGG
ATCAACTTGGGGAAGAAGCTAATCGGGCTCTGGAAGTTTTGCAGAAAGAGGTTGCTTGTCACAGACTAGGTAATCAAGATGCAGCCGAGACAATAGCTAA
ACTTCAAGCTGAAATAAGGGAAATGCGCTCAGCTCCGCAGCTACCAAGGGAAGTTGCAATTGGAAGTACTGGAGTAGCCACTAACAAGAGTGTCAGTGCT
AATCTGAAGGAAGAAATAACGAGACTTCATTCGCAAGGAAGCAACATCGCAAATCTCGAAGAGCAACTAGAGAACGTCCAGAAGTCGATTGACAAGCTGA
TCATGTCACTTCCAAGTAACAATTCAGATGAGTCAGCAGCTTCCAATCCTAAGAACCAGCACAGGAAGAAAAAGACTCACCCTTTATGTTCAAGCAACGG
TGCAAACATGCAGAACTTCATTAGATCTCCTTGTGCGCCTTTGGCATCATCGAGGCAAGTACTGGAGAACGAGACAGAAAATAGAGCACCGGAGAGCGAT
GATGTTGCGTCCAAAGTGTCCGAGAAACAACAACAGACTCCAATGAAGAGTGAGGAAGGTGGAGATGTGTCGTCAAAGGAAGGGACACCTGGTTCTCGGC
GTACAAGCTCAGTCAACATGAAGAAAATGCAGAAGATGTTCCAAAATGCAGCAGAGGAAAATGTGAGAAGCATTAGAAACTATGTCACAGAGCTGAAGGA
ACGCGTTGCAAAACTCCAATATCAAAAGCAGCTGTTAGTCTGCCAGGTACTAGAACTGGAAGCAAATGAAGCAGCTGGGTATAACTTGGAGGACGAGGAG
GAAGCAGCGATTGGACCGGTAGAACCACAAGTTCCATGGCATGTGACCTTCAGAGAACAACGGCAGCAGATCATAGAATTATGGGACGTTTGCTTTGTAC
CAATCATCCACAGAACACAATTCTATCTTCTGTTCAAAGGAGACCCTGCTGATGAGATATACATGGAGGTGGAACTCAGAAGAATAACATGGCTGCAGCA
GCATCTGGCTGAGGTTGGCGATGCAAGCCCTGGTCGTGGTGGTACCGGAGGAGATGAACCTACAGTTTCCCTTGCGTCAAGTTTGAGAGCACTGAAACGT
GAAAGGGAGTTCCTGGCGAAGAGGCTGGGTCGGCTGAGCATGGAGGAGAGGGAAGCAATGTTCATGAAGTGGAATGTGCCACTGGATGGGAAGCATAGAA
GGCTGCAGTTCATCAGCAAGCTGTGGACGAATCCGCATGATGGGAAGCACGTGGAGGAAAGTGCTGAGATCGTCGGGAGGCTGGTTGGGCTGTACCAAGG
AGGGGTAGTGTCGAAGGAGATGTTCGAGCTTAATTTCGCTCCACCGACGGATAAGAGAGCGTGGTGGGAATGGAACCCGATTTCCAACCTGCTCAACCTC
TGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10042952 pacid=23168400 polypeptide=Lus10042952 locus=Lus10042952.g ID=Lus10042952.BGIv1.0 annot-version=v1.0
MTAKTPGTPASKIENRTPASTPGGGSKAKDEKIVVTVRLRPLNRREQLAKDQVAWECLDDYTISYKPPPQERSAQPASFTFDKVFGPACVTESVYEDGVK
NIALSALMGINATIFAYGQTSSGKTYTMRGITEKAVQDIYNHIRNTPERDFTIKMSGLEIYNENVRDLLNSESGRSLKLLDDPEKGTVVEKLVEETASND
SHLRKLISICEAQRQVGETALNDTSSRSHQIIRLTIQSTLNESSDCARSFVASLNFVDLAGSERASQTHTDGARLREGCHINLSLMTLTTVIRKLSVGKR
SCHIPYRDSKLTRILQHSLGGNARTAIICTLSPALTHVEQSRNTLYFATRAKEVTNSAKVNMVISDKQLVKHLQKEVARLEAERRTPDPSSEKDWKIQQL
EMEIEELRRQRDLAKCEVDELRKKLQDEKLEDGKIFREVESPHPLVKKCLSYSAAMTSKPESKDFIHCEKMTKATLRQSMRLSSAAPFTLMHEIRKLEHL
QDQLGEEANRALEVLQKEVACHRLGNQDAAETIAKLQAEIREMRSAPLPREVAIGSNKSVSANLKEEITRLHSQGSNIANLEEQLENVQKSIDKLIMSLP
SNNSDESAASNPKNQHRKKKTHPLCSSNGANMQNFIRSPCAPLASSRQVLENETENRAPESDDVASKVSEKQQQTPMKSEEGGDVSSKEGTPGSRRTSSV
NMKKMQKMFQNAAEENVRSIRNYVTELKERVAKLQYQKQLLVCQVLELEANEAAGYNLEDEEEAAIGPVEPQVPWHVTFREQRQQIIELWDVCFVISDKQ
LVKHLQKEVARLEAERRTPDPSSEKDWKIQQLEMEIEELRRQRDLAKCEVDELRKKLQDEKLEDGKIFREVESPHPLVKKCLSYSAAMTSKPESKDFIHC
EKMTKATLRQSMRLSSAAPFTLMHEIRKLEHLQDQLGEEANRALEVLQKEVACHRLGNQDAAETIAKLQAEIREMRSAPQLPREVAIGSTGVATNKSVSA
NLKEEITRLHSQGSNIANLEEQLENVQKSIDKLIMSLPSNNSDESAASNPKNQHRKKKTHPLCSSNGANMQNFIRSPCAPLASSRQVLENETENRAPESD
DVASKVSEKQQQTPMKSEEGGDVSSKEGTPGSRRTSSVNMKKMQKMFQNAAEENVRSIRNYVTELKERVAKLQYQKQLLVCQVLELEANEAAGYNLEDEE
EAAIGPVEPQVPWHVTFREQRQQIIELWDVCFVPIIHRTQFYLLFKGDPADEIYMEVELRRITWLQQHLAEVGDASPGRGGTGGDEPTVSLASSLRALKR
EREFLAKRLGRLSMEEREAMFMKWNVPLDGKHRRLQFISKLWTNPHDGKHVEESAEIVGRLVGLYQGGVVSKEMFELNFAPPTDKRAWWEWNPISNLLNL
|
DESeq2's median of ratios [FLAX]
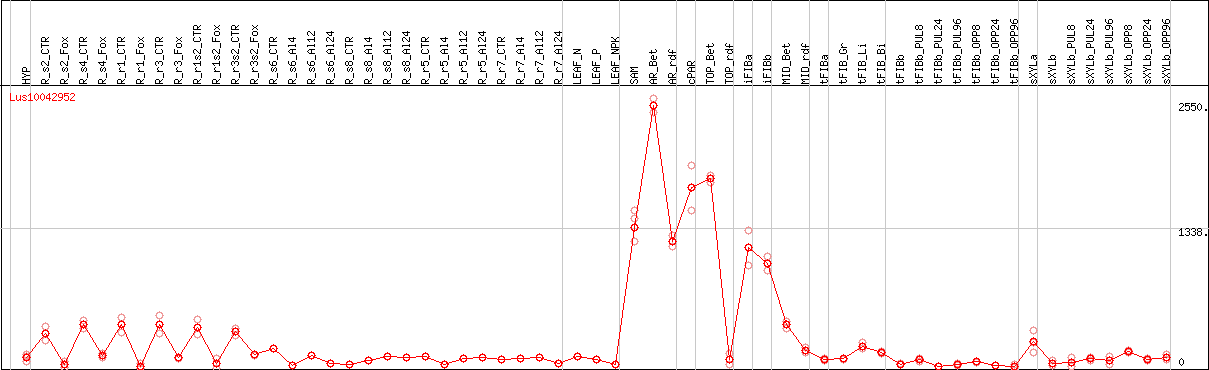
Coexpressed genes
Lus10042952 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
