Lus10043045 [FLAX]

| External link |
|
||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
||||||||||||||||||||||||||||||||||||||||
| PFAM info | |||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10043045 pacid=23168337 polypeptide=Lus10043045 locus=Lus10043045.g ID=Lus10043045.BGIv1.0 annot-version=v1.0
ATGGCTCCTGCAGCAGCTCCTTTGTCACACCTTCTGTTCTCCTTCGTCTTCTGCTTCTGCTTCCTGTCTACTCCTTGCATGAGTGTCGCCGATCTGGATG
ATGGCGGATTCTCCATCTCCGGCTTCGATTCTGTTTTCTATCCTGGCGACTACAGTCCTCCGTCGCCCCCTGCGCCGCCTCCGCCTCCGCTCCCGCCCTC
TGTGACCTGCGAGGGGGATCTCAAAGGTGTAGGCTCCTTAGACTCTGTGTGCAAGCTGAGTTCGTCTTTGAATTTCACTCATGACGTGTACATAGGAGGG
GGCGGGTCATTGGTCTTGCTTCCTGATGTCGTTTTGAGCTGCGCAGTCCGCGGCTGTGCAATTTTGGTCAGTGTCGGAGATGAGATCAGGATGGGGGAGA
AGGCGGCGATAATTGCTGGGACTGTCTGGCTCACTGCTCAGAATGCAACTTTTAGTTACGGCTCCTTGATTAACTCTACTGCCTTGGCTGGTGCGCCTCC
TGAAGAGACCAGTGGAACTCCTGGTGGCGTTCAAGGAGCTGGTGGTGGATATGGAGGAAGGGGAGCTAGCTGTGTCTCGGATAATACCAAGTTGCCGGAG
GATGTGTGGGGAGGGGACGCTTATTCTTGGGCGGACTTGAACGCTCCTTGGAGTTATGGTAGTAAAGGTGGAACTACTAGCAAGGAGGAGAATTATGGGG
GAGAAGGAGGCGGAAGGATTGGAATGAATATTACCAACTCAATTGAGCTTTATGGGACTTTGTTGGCGGAAGGTGGAGATGGTGGTATAAAGGGTGGAGG
GGGTTCAGGTGGCAGCATCCGTATCATCGGTCATAGAATGGTAGGGGATGGTACATTAAGCGCTTCTGGTGGTAATGGATTTGCTGGAGGTGGCGGTGGA
AGAGTTTCTGTTGATGTTTTCAGCAGACATGATGATATATCGATCTTTGTCCATGGTGGAAGGAGTCTTGGCTGCACCGAAAATTCAGGTGCTGCAGGCA
CATATTATGATGCTGTTCCTCGGAGTCTTACAGTTAATAACAATAACATGTCCACCAACACTGATACACTTCTGTTGGAATTCCCCAAGCAGCCACTTTG
GACCAATGTTAACATTCAGAATCGTGCGAGGGCTTCTGTTCCTTTATTTTGGAGTCGGGTTCAGGTCCATGGACAAATCAGTTTATCCAGTGAGGCAGTG
ATGAGCTTTGGCCTTCCGCATTATGCGACATCTGAATTTGAGTTGATGGCTGAAGAACTTCTAATGAGCAACTCAGTTGTTAAGGCAGGTGACAGACTAT
TGTTCTTTCTTCCAAATCTTTTTCCTTCCTTTTCTGCACTTTGTGATGATGTTTCATGCTGCCAAATGCCTTATTTTCTAATGCTCATTTATAACCAGAT
ATATGGGGCCCTTCGTATGTCCGTTAAGATATTGTTGATGTGGAATTCCAAGATGCTTATAGATGGAGGTGGTGATGAGCTTGTGGCTACTTCTGCTCTC
GAGGCCAGCAACCTAGTTGTTCTCAAGGAATCTTCAGCAGTACACTCCAATGCAAATCTGGAAGTTCATGGGCAAGGCTTTTTTAATTTGTCTGGACCAG
GGAACATGATTGAAGCTCAAAGGCTAATTCTATCCTTGTTTTTTGGTATCTATGTTGAACCTGGATCTGTTCTTAGAGGCCCCTTGAAGGATGCAGATGA
TCGTAGCATGACGCCACGGCTCTACTGTGAAGATCGAGAATGCCCAAGGGAACTGATTCATCCGCCTGAAGATTGTAATATGAACTCTTCATTGGCCTTT
ACTCTCCAGATTTGTCGAGTTGAAGAGGTTATTGTTGGAGGCTCAGTATCGGGATCTGTTGTTCATTTTCGTCTAGTCAGAAATATTATTGTCAATTCTT
CTGGAGTAATCAGTGCTTCTGGATTGGGATGTACTGGTGGTATTGGTAGAGGAAACATTTTTGACAATGGTCTTGGTGGTGGAGGTGGACATGGCGGAAA
AGGTGGTAAAGGATATTATAATGGAAGTTTCGTGGATGGTGGTGTTGCTTATGGAAATGCTGAATTGCCTTGTGAACTTGGAAGCGGCAGGGGAAATTAT
AGTTTAGCTGGATCGACTGCAGGCGGTGGTATAATTGTGATCGGTTCAATGGAGCATGCGCTGTCCAGTTTGTCTGTTTTTGGCTCACTTAGAGCAGATG
GTGAAAGCTTCGGAGAGGACCCAAAGAAAGATGGTGGAATGACTTCAAGCATAGGTCCTGGAGGTGGTGCTGGTGGAACTATTCTTTTATTTGTGCATTC
AGTCACACTTGCGAATTCTTCTGCTATTTCCACCATGGGAGGGCATGGGAGTCCTAACGGCGGAGGTGGTGGAGGTGGTGGAAGGGTTCACTTCCATTGG
GCGAATATTCCTTCAGGAGATGAGTATATGCCAATAGCAAGCTCAAATGGGAGCATTCAGACTGGAGGGGGGCTTGGTAGAGGCCAGGGTCATACAGGAG
ATGATGGAACTGTTACTGGGAAGGCCTGTCCGGAAGGGCTCTATGGTATCTTCTGCGAGGAGTGCCCCTTGGGTACATTCAAGAATATCAGTGGATCTGA
TAAAGCCCTGTGTTATAGTTGCCCATTCTACGAGCTACCAAGTCGTGCCATTTATGTCACTGTCCGTGGTGGTGTTACTGAAAGACCTTGTCCATACAAG
TGTGTGACAGACAGATATCACATGCCAAACTGTTACACAACCCTAGAGGAGTTGGTATACACTTTTGGCGGGCCGTGGTTGTTTAGTCTTATTCTATTGG
GTCTACTCATCCTGTTAACTCTAGTGCTTAGTGTTGCACGAATGAAATATGTTGCGGGGGAGGAATTACCAACTCTTGTGCCTGCTCAGCGGGGTTCTCG
AATTGATCACTCGTTCCCTTTCCTCGAGTCACTGAATGAGGTATTGGAAACGAACAGAACCGAAGAATCTCGGAGTCATGTGCATAGGATGTTTTTTATG
GGGGCAAATACTTTCAGTGAACCTTGGCACTTAACTCATTCTCCTCCTAAACAAGTTATTGAGATTGTGTATGAGGATGCGTTCAACAGGTTTGTAGATG
AGATCAATGGTTTGGCTGCATATCAGTGGTGGCAAGGATCTGTATATAGCATCCTTTCTGTTCTCGCATATCCACTTGCCTACTCCTGGCTACAGCAGTG
TAGGAAAAGGAAACTTCAGCAGCTACGTGATTTTGTACGTTCAGAATATGATCATGCTTGCTTACGGTCTTGTCGTTCCCGTGCTCTTTACGAAGGACTC
AAGGTGGGAGCCACTTTGGACCTAATGCTTGCATACTTGGACTTTTTCCTTGGTGGAGATGAGAAGAGACCTGATCTTCCTCCTAGTCTTCATCAGAGAT
TTCCCATGACTTTAGTTTTGGGGGGAGATGGAAGTTACATGGCACCTTTCTCACTTCACAATGATAATATTCTCACCAGTCTCATGAGCCAGGCTGTTCC
ACCAACCATCTGGTATAGGCTAGTGGCTGGTCTTAATGCTCAATTACGTTTGGTTCAACGCGGACGCTTGAAATTTACATTTCTTCGAGTTATTTCCTGG
ATTGAAACACATGCGAATCCTACGTTGGCTACCTATGGAATACGTGTGGGTCTTGCTTGGTTTCAGCCCACTGCCCCTGGTTATTGTCAGTTTGGACTTG
TAGTATATGCCACTGGAGATAATAATACACTGCAGTCAGGCGACGGGAGTGGGATTGTATGTTCCCCTCTGGCGAACAGGGGTGATCAAAGGCAATATCC
GAAGGGTACGGAAGCTCTGCTGACACGTAAAGTATGTGGAGAAGGAATTTTGAATTCGAGTACTCTACAGACACTTAAAATAGGAAGGGCTTCATTTTCT
TTCCCTTTCTTGATATTCAACATCAGACCTGTTGGGCACCAGGACCTTGTGGGTTTGCTGATATCTATGGTCCTTCTAGGAGATATTAGCTTAGTGATGC
TTAGTCTGCTACAGATGTACTCGATATCGCTGCTCAACTTCCTATTAGTTTTGTTTATTCTTCCTCTGGGGATATTGTTTCCATTCCCTGCTGGGATCAG
TGCTTTGTTCAGTCATGGACCGAGACGTTCGGCAGGTCTTTCTCGTCTATATGCTTTGTGGAATTTTACATCCTTGATCAACGTTGTGGTAGCATTAGTA
TGCGGATTTGTTCATTACATGAGTCGCTCAAGGGAAAGCCACCTCAACTTGCCGTCCTGGAATTTTAGCATGGATGATAGCGAGTGGTGGATACTTCCAT
GTGGGCTGTTGGTTTGTAAGATTGTGCAAGCACGGCTGATTGATTGTCACATTGCTAACCAAGAAATCCAAGACTACTCCTTGTACAGTAGCGATCCAGG
AGTGTTCTGGCAAGCTTGA
|
||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10043045 pacid=23168337 polypeptide=Lus10043045 locus=Lus10043045.g ID=Lus10043045.BGIv1.0 annot-version=v1.0
MAPAAAPLSHLLFSFVFCFCFLSTPCMSVADLDDGGFSISGFDSVFYPGDYSPPSPPAPPPPPLPPSVTCEGDLKGVGSLDSVCKLSSSLNFTHDVYIGG
GGSLVLLPDVVLSCAVRGCAILVSVGDEIRMGEKAAIIAGTVWLTAQNATFSYGSLINSTALAGAPPEETSGTPGGVQGAGGGYGGRGASCVSDNTKLPE
DVWGGDAYSWADLNAPWSYGSKGGTTSKEENYGGEGGGRIGMNITNSIELYGTLLAEGGDGGIKGGGGSGGSIRIIGHRMVGDGTLSASGGNGFAGGGGG
RVSVDVFSRHDDISIFVHGGRSLGCTENSGAAGTYYDAVPRSLTVNNNNMSTNTDTLLLEFPKQPLWTNVNIQNRARASVPLFWSRVQVHGQISLSSEAV
MSFGLPHYATSEFELMAEELLMSNSVVKAGDRLLFFLPNLFPSFSALCDDVSCCQMPYFLMLIYNQIYGALRMSVKILLMWNSKMLIDGGGDELVATSAL
EASNLVVLKESSAVHSNANLEVHGQGFFNLSGPGNMIEAQRLILSLFFGIYVEPGSVLRGPLKDADDRSMTPRLYCEDRECPRELIHPPEDCNMNSSLAF
TLQICRVEEVIVGGSVSGSVVHFRLVRNIIVNSSGVISASGLGCTGGIGRGNIFDNGLGGGGGHGGKGGKGYYNGSFVDGGVAYGNAELPCELGSGRGNY
SLAGSTAGGGIIVIGSMEHALSSLSVFGSLRADGESFGEDPKKDGGMTSSIGPGGGAGGTILLFVHSVTLANSSAISTMGGHGSPNGGGGGGGGRVHFHW
ANIPSGDEYMPIASSNGSIQTGGGLGRGQGHTGDDGTVTGKACPEGLYGIFCEECPLGTFKNISGSDKALCYSCPFYELPSRAIYVTVRGGVTERPCPYK
CVTDRYHMPNCYTTLEELVYTFGGPWLFSLILLGLLILLTLVLSVARMKYVAGEELPTLVPAQRGSRIDHSFPFLESLNEVLETNRTEESRSHVHRMFFM
GANTFSEPWHLTHSPPKQVIEIVYEDAFNRFVDEINGLAAYQWWQGSVYSILSVLAYPLAYSWLQQCRKRKLQQLRDFVRSEYDHACLRSCRSRALYEGL
KVGATLDLMLAYLDFFLGGDEKRPDLPPSLHQRFPMTLVLGGDGSYMAPFSLHNDNILTSLMSQAVPPTIWYRLVAGLNAQLRLVQRGRLKFTFLRVISW
IETHANPTLATYGIRVGLAWFQPTAPGYCQFGLVVYATGDNNTLQSGDGSGIVCSPLANRGDQRQYPKGTEALLTRKVCGEGILNSSTLQTLKIGRASFS
FPFLIFNIRPVGHQDLVGLLISMVLLGDISLVMLSLLQMYSISLLNFLLVLFILPLGILFPFPAGISALFSHGPRRSAGLSRLYALWNFTSLINVVVALV
CGFVHYMSRSRESHLNLPSWNFSMDDSEWWILPCGLLVCKIVQARLIDCHIANQEIQDYSLYSSDPGVFWQA
|
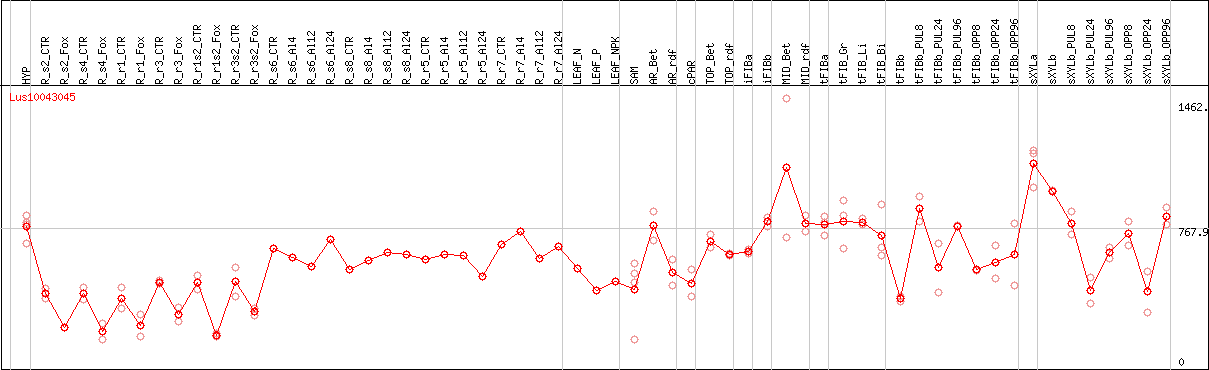
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10043045 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
