External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G50920 229 / 9e-70
CLPC1, CLPC, ATHSP93-V, HSP93-V, DCA1
HEAT SHOCK PROTEIN 93-V, DE-REGULATED CAO ACCUMULATION 1, CLPC homologue 1 (.1)
AT3G48870 226 / 1e-68
ClpC2, ATCLPC, ATHSP93-III, HSP93-III
ClpC2, Clp ATPase (.1.2)
AT3G45450 137 / 7e-39
Double Clp-N motif-containing P-loop nucleoside triphosphate hydrolases superfamily protein (.1)
AT1G21470 57 / 2e-10
unknown protein
AT5G51070 56 / 4e-09
SAG15, CLPD, ERD1
SENESCENCE ASSOCIATED GENE 15, EARLY RESPONSIVE TO DEHYDRATION 1, Clp ATPase (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10032543
376 / 5e-134
AT5G50920 265 / 9e-83
HEAT SHOCK PROTEIN 93-V, DE-REGULATED CAO ACCUMULATION 1, CLPC homologue 1 (.1)
Lus10036017
267 / 1e-83
AT5G50920 1542 / 0.0
HEAT SHOCK PROTEIN 93-V, DE-REGULATED CAO ACCUMULATION 1, CLPC homologue 1 (.1)
Lus10016723
256 / 1e-79
AT5G50920 1526 / 0.0
HEAT SHOCK PROTEIN 93-V, DE-REGULATED CAO ACCUMULATION 1, CLPC homologue 1 (.1)
Lus10043014
53 / 7e-08
AT5G51070 664 / 0.0
SENESCENCE ASSOCIATED GENE 15, EARLY RESPONSIVE TO DEHYDRATION 1, Clp ATPase (.1)
Lus10043013
50 / 6e-07
AT5G51070 1288 / 0.0
SENESCENCE ASSOCIATED GENE 15, EARLY RESPONSIVE TO DEHYDRATION 1, Clp ATPase (.1)
Lus10032514
49 / 2e-06
AT5G51070 1280 / 0.0
SENESCENCE ASSOCIATED GENE 15, EARLY RESPONSIVE TO DEHYDRATION 1, Clp ATPase (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.012G105900
267 / 9e-84
AT5G50920 1505 / 0.0
HEAT SHOCK PROTEIN 93-V, DE-REGULATED CAO ACCUMULATION 1, CLPC homologue 1 (.1)
Potri.015G105100
263 / 4e-82
AT5G50920 1508 / 0.0
HEAT SHOCK PROTEIN 93-V, DE-REGULATED CAO ACCUMULATION 1, CLPC homologue 1 (.1)
Potri.012G112000
64 / 7e-12
AT5G51070 1328 / 0.0
SENESCENCE ASSOCIATED GENE 15, EARLY RESPONSIVE TO DEHYDRATION 1, Clp ATPase (.1)
Potri.015G110000
60 / 2e-10
AT5G51070 1282 / 0.0
SENESCENCE ASSOCIATED GENE 15, EARLY RESPONSIVE TO DEHYDRATION 1, Clp ATPase (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF02861
Clp_N
Clp amino terminal domain, pathogenicity island component
Representative CDS sequence
>Lus10043198 pacid=23167908 polypeptide=Lus10043198 locus=Lus10043198.g ID=Lus10043198.BGIv1.0 annot-version=v1.0
ATGGCGAGATTTCTAGTTCATTCCACCATTGCCCCTTCCCTGGCTCCTCCTGCTAAGGGAATCCGGCCTCAAGGATCCGAAAACTCGAAAAGGACTGTTG
TCAAGGCGATGGCAAGTTTACAAGCACCCTTGAGGGGTTACTCTGGGTTGCGGAGTTCAACCTCTTTCGGTACATGGCAAGCTTTTGGTTCAAGGCTTGC
TATCTCTACCTTTCCTAGGCAGAGGAGAGCTAAAAGATTTGTAGCTAAAGCTATGTTTGAGCGGTTCACAGAAAAAGCTATCAAAGTTATCATGCTTGCT
CAAGAGGAGGCTAGACGTCTTGGCCACAACTTTGTTGGCACTGAGCAAATCCTTTTGGGTCTTATCGGCGAGGGAACTGGCATTGCTGCTAAGGTCCTCA
AATCTATGGGTATCAATCTTAAGGACGCTCGTGTAGAAGTCGAAAAGATCATTGGACGAGGCAGTGGATTTGTAGCTGTGGAAATCCCATTCACTCCTCG
AGCTAAGCGCGTTCTAGAATTGTCGCTTGAAGAAGCTCGCCAACTCGGTCACAACTACATCGGATCTGAGCACTTGCTCTTGGGGCTGCTCCGTGAAGGT
GAGGGTGTTGCGGCCCGTGTTCTTGAGAATTTGGGTGCTGATCCTAGTAATATAAGAACACAGGCAAGTTCTAAACTCTTTTAA
AA sequence
>Lus10043198 pacid=23167908 polypeptide=Lus10043198 locus=Lus10043198.g ID=Lus10043198.BGIv1.0 annot-version=v1.0
MARFLVHSTIAPSLAPPAKGIRPQGSENSKRTVVKAMASLQAPLRGYSGLRSSTSFGTWQAFGSRLAISTFPRQRRAKRFVAKAMFERFTEKAIKVIMLA
QEEARRLGHNFVGTEQILLGLIGEGTGIAAKVLKSMGINLKDARVEVEKIIGRGSGFVAVEIPFTPRAKRVLELSLEEARQLGHNYIGSEHLLLGLLREG
EGVAARVLENLGADPSNIRTQASSKLF
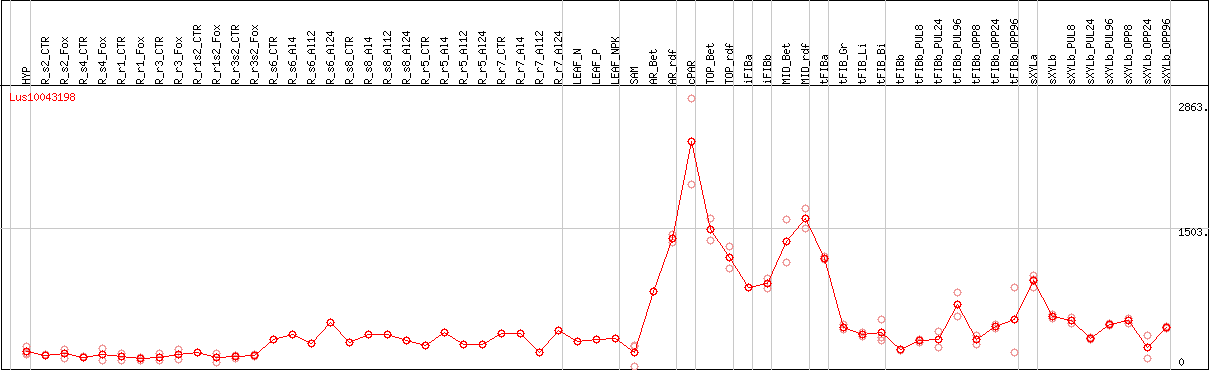
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10043198 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.