External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G17680 84 / 1e-18
disease resistance protein (TIR-NBS-LRR class), putative (.1)
AT4G12020 69 / 2e-13
WRKY
MEKK4, MAPKKK11, ATWRKY19, WRKY19
MAPK/ERK KINASE KINASE 4, MITOGEN-ACTIVATED PROTEIN KINASE KINASE KINASE 11, protein kinase family protein (.1.2.3)
AT4G19500 67 / 4e-13
nucleoside-triphosphatases;transmembrane receptors;nucleotide binding;ATP binding (.1.2)
AT3G25510 67 / 8e-13
disease resistance protein (TIR-NBS-LRR class), putative (.1)
AT5G36930 66 / 9e-13
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
AT5G41750 66 / 1e-12
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
AT5G46270 66 / 2e-12
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT5G22690 65 / 3e-12
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT5G40060 64 / 6e-12
Disease resistance protein (NBS-LRR class) family (.1)
AT1G31540 63 / 1e-11
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10011104
221 / 9e-67
AT5G17680 574 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10003749
116 / 5e-30
AT5G17680 517 / 5e-161
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10041060
92 / 2e-21
AT5G17680 641 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10029722
85 / 5e-19
AT5G17680 608 / 0.0
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10041078
74 / 3e-15
AT5G17680 399 / 6e-126
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10030345
72 / 1e-14
AT5G17680 528 / 5e-168
disease resistance protein (TIR-NBS-LRR class), putative (.1)
Lus10039850
69 / 9e-14
AT5G36930 573 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10008222
68 / 2e-13
AT4G12010 327 / 4e-96
Disease resistance protein (TIR-NBS-LRR class) family (.1)
Lus10008221
67 / 5e-13
AT4G12010 400 / 8e-120
Disease resistance protein (TIR-NBS-LRR class) family (.1)
PFAM info
Representative CDS sequence
>Lus10043227 pacid=23168204 polypeptide=Lus10043227 locus=Lus10043227.g ID=Lus10043227.BGIv1.0 annot-version=v1.0
ATGGGTAGAGATATTGTTTTTCACGAATCTCCCAGGAATCCTGGAAGGCGCAGTCCATTGTGGGATCCCGAAGATGTCCTTTCTGTGTTGATTACACCAA
TGGGAAGGGAAGAGGTGGAAGCCATTGTATTGGATCTATCAGAGATGGAGGAGATGCAATTAAATAGGGAAGCTTTCTCAAATATGAGTCTGAGACTGCT
TATCATACGCAAGGCCAAATTTGATGCAGGACCCGTCAATCTTCCCAGTGAATTGAGATTTCTTGAATGGGATGGGTACCCTGCAGACTCGTTTCCGGCT
CCTTTTAGACATAGTAAACTCGTGGAACTTATCATGCACCGCCCTCTATTTTCTATCTCAAGAAACTCAAGAGTAATATCATTTCAGGGCTGCAAGGGAC
CATTATTATACGAGTCAACCCAGCGCACATTTACATCTTCAAGTCAACATCACTTGTTAATGAACTGTTACAACATGAAGTTGCCTGCCTTGTTTGGTTA
CCATTCAATAACAACTCTGGACCTGGGGGACTGTGGGCTGAGGGAAGGAGATCTTGTTTCCGTCCTTGGAGTTGTTGAATCTAAGCAATAG
AA sequence
>Lus10043227 pacid=23168204 polypeptide=Lus10043227 locus=Lus10043227.g ID=Lus10043227.BGIv1.0 annot-version=v1.0
MGRDIVFHESPRNPGRRSPLWDPEDVLSVLITPMGREEVEAIVLDLSEMEEMQLNREAFSNMSLRLLIIRKAKFDAGPVNLPSELRFLEWDGYPADSFPA
PFRHSKLVELIMHRPLFSISRNSRVISFQGCKGPLLYESTQRTFTSSSQHHLLMNCYNMKLPALFGYHSITTLDLGDCGLREGDLVSVLGVVESKQ
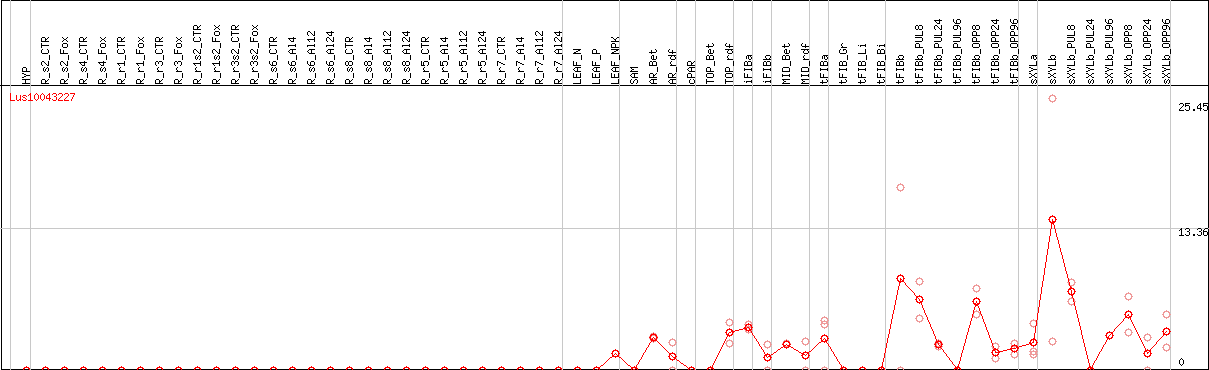
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10043227 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.