Lus10043229 [FLAX]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Poplar homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Lus10043229 pacid=23168062 polypeptide=Lus10043229 locus=Lus10043229.g ID=Lus10043229.BGIv1.0 annot-version=v1.0
ATGATGCCAGGAATCCTATGTTCGACAAGCAGAATTGCACTCATTCCAGGAACTCCTTTCACATTAACAAAGAACAACGGTTCTACTAAATGTAGTTTAT
CCAGGAAACCAACCAAGCATGCAACATTCTTACCCTTCCCTGCATCGGTTAAGTCCTTGCCCAATTACAGAAGGTGTCATCATGCTTTCGTCAGGACGAG
GACGATGCACGAGCTATCAGCCACAGGCACTGACGTAGCGGTTGGTGAGGCGGAAACACCTGCTGTCACTGAGGAAACTTCTGATGGAGCCGATGAAAAT
GGAACTTCCTCTTCTCCCCCATCGGATGCCAGTACTCCAACAGCTCAACCTAAGCGTGCAAGACCAGGAAGAAGAAGCGAGATGCCACCGGTAAAGAATG
AGGATTTGGTTCCTGGTGCAACATTCACCGGAAAAGTCAGGTCTATTCAACCATTTGGCGCGTTCATCGATTTTGGAGCTTTCACGGATGGCCTTGTACA
TGTTTCAAAGCTGAGTGATGGCTTTGTTAAGGATGTTGGAGATGTTGTTTCTGTTGGACAAGAGGTTCAGGTAAGGCTAGTTGAAGCAAGCACTGAGACA
GGGCGTATTTCTCTTAGTATGCGAGACGATGATTCAAGTAGTAAGCCGCAGCAAAGAAGAGATACATCTTCTGCAGCCAGTGGAGGAGATAAGCCAAGAC
CTGCAAGAAGAAATACACAGAGGACAGGCCAGAAGAGGGATGAAAAGAGTTCAAAGTTTACTATTGGGCAGCAACTAGAAGGAACAGTTAAGAACTTGGC
CAGGTCTGGTGCTTTCATTTCTCTCCCTGACGGTGAGGAAGGATTCTTGCCTTCATCAGAAGAAGCTACTGAGGGATTCGTAAACATGATGGGTGGCTCT
AAACTGGAGGTAGGACAACAAGTCAGTGTGAGGGTTTTGCGTGTGAATAGAGGCCAGGCTACATTGACAATGAAAAAGGAGGTAGACAATAAAAATGTAG
GTGGGTTACTAGGCCAGGGAGTGGTCCATGCGGCAACAAATCCTTTCCTGCTGGCTTTCCGTAAGAACGAGGATATAGCAAGCTTTCTAGACGAAAGAGA
GAAGGCAGAGAAAGAGGCAGAGACTACTTTAGTGGAGGAAGAGCTAAAACAGGAAGGTAATGTGTCTGATGTTCAATTAGAAACCGGTTCCGACAGCTTA
GTTGGTGTCTCTTCCTTGGTAGACCAAACGGTAGAAGATGCTGTTGCTTCTTCGGAGGTGATTCTAGACTCTAGTTTATCAGAAGTGGAGAGTACTGTTC
AAATTATAGCTGATAAGACGATTGAAGAAGAGAATGTAGCTGTGGATATCGCAATTCCAGATCAAAAGGTTGAGGAAGAACCTTTGGCAGAAGCAGAGGA
AAGCGAGGTTAAGTCCGGTTTATCTGAAGAAATAGCTGGTCAAGCCTTATCGTCAGAAACTCCCGTCAATGAAACTATTGCCACACCAGTAGTAGATAAT
GAAATCGTGGCACCAGATTCTGAAGGAAATGGGAGTACTGCCATTGCAGATGAAGTGGAAAATCTTACAGCGGCAGAAGTCCCAGAGGAAGTCGAAGCCA
ACAAGGATGCAGTGGAGATTCAAACCTCTGCCATGGACAATGAGACTACTTCTGCCTCTCTGGTTCCAGATGAGATACCTGAGAAAAATGCAACAACTAT
CTCACCAGCTCTGGTAAAGCAACTTCGAGAGGAAACAGGAGCAGGAATGATGGATTGCAAGAACGCTCTTTCCGAGACAGGAGGCGACATCATTAAAGCA
CAGGAATTCCTCCGAACCAAAGGGCTGGCAAGTGCAGAGAAAAAGGCGAGCAGAGCCACTGCAGAAGGAAGAATCGGCTCTTACATTCATGACAGCAGAA
TCGGTATCCTCATTGAGGTCAACTGCGAAACAGACTTTGTTTCCCGTGGCGAAATTTTCAAGGAACTGGTTGATGACCTATCGATGCAAGTGGCTGCGTG
CCCACAAGTGAAGTATGTAGCCAAGGAAGATGTTGCTGAAGAACTCGTTAGCAAGGAAAGAGAGATCGAGATGCAAAAGGAAGATCTCCTGTCCAAACCG
GAGCAGATCAGAGCTAAAATCGTGGAAGGGCGGATCCAGAAGAGGCTCGACGAACTGGCACTGCTGGAGCAACCATACATCAAGAATGATAAGGTTGTGG
TTAAGGACTGGATTAAGCAGACAATAGCCACCATTGGCGAAAATATGAAAGTTAAGAGGTTTGTCCGCTTCAATCTCGGTGAGGGTCTCGAGAAGAAAAG
CCAGGATTTTGCTGCTGAGGTTGCTGCACAAACTGCAGCTAGATCAGCCCCAGCGCCACCGAAAGAAGAACCTGCTGCAACCGAGGACAAAGAAGATGTT
CAGAAGGCACCAGCGGCACCGGTTTCTGCAGCACTGGTGAAGCAACTGAGGGAAGAAACAGGAGCAGGGATGATGGACTGCAAGAAAGCACTCTCTGAAA
CTGGAGGTGATCTGGAGAAGGCACAGGAGTATCTCAGAAAGAAAGGTCTTTCATCCGCGGATAAGAAATCCAGCCGAATTGCTGCTGAGGGCCGAATTGG
TTCCTACATCCACGATTCTCGCATCGGAGTACTGATTGAGGTCAACTGCGAGACTGATTTCGTCGGCAGAAGCGAGAAATTCAAGGAATTAGTTGATGAT
CTGGCAATGCAGGTTGTGGCCTGCCCACAGGTGCGGTTTGTATCTGTCGAAGATGTGCCGGAGAGCGTAATAATGAAAGAAAAGGAACTCGAAATGCAGA
GGGAGGACCTCCAGTCGAAACCGGAGAACATTAGGGAAAAGATTGTGGAAGGAAGGATGTCGAAAATGCTGGGAGAGCTTGCACTTCTGGAGCAGCCGTT
CATTAAAGACGACAAACTCCTGGTGAAGGATTTGGTGAAGCAAACCATTGCTGCTCTGGGTGAGAACATAAAGGTCCGCAGGTTTGTCAAGTTTACTCTT
GGAGAAGCCATTGAAGATGCAAACCCGGAAGCTTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Lus10043229 pacid=23168062 polypeptide=Lus10043229 locus=Lus10043229.g ID=Lus10043229.BGIv1.0 annot-version=v1.0
MMPGILCSTSRIALIPGTPFTLTKNNGSTKCSLSRKPTKHATFLPFPASVKSLPNYRRCHHAFVRTRTMHELSATGTDVAVGEAETPAVTEETSDGADEN
GTSSSPPSDASTPTAQPKRARPGRRSEMPPVKNEDLVPGATFTGKVRSIQPFGAFIDFGAFTDGLVHVSKLSDGFVKDVGDVVSVGQEVQVRLVEASTET
GRISLSMRDDDSSSKPQQRRDTSSAASGGDKPRPARRNTQRTGQKRDEKSSKFTIGQQLEGTVKNLARSGAFISLPDGEEGFLPSSEEATEGFVNMMGGS
KLEVGQQVSVRVLRVNRGQATLTMKKEVDNKNVGGLLGQGVVHAATNPFLLAFRKNEDIASFLDEREKAEKEAETTLVEEELKQEGNVSDVQLETGSDSL
VGVSSLVDQTVEDAVASSEVILDSSLSEVESTVQIIADKTIEEENVAVDIAIPDQKVEEEPLAEAEESEVKSGLSEEIAGQALSSETPVNETIATPVVDN
EIVAPDSEGNGSTAIADEVENLTAAEVPEEVEANKDAVEIQTSAMDNETTSASLVPDEIPEKNATTISPALVKQLREETGAGMMDCKNALSETGGDIIKA
QEFLRTKGLASAEKKASRATAEGRIGSYIHDSRIGILIEVNCETDFVSRGEIFKELVDDLSMQVAACPQVKYVAKEDVAEELVSKEREIEMQKEDLLSKP
EQIRAKIVEGRIQKRLDELALLEQPYIKNDKVVVKDWIKQTIATIGENMKVKRFVRFNLGEGLEKKSQDFAAEVAAQTAARSAPAPPKEEPAATEDKEDV
QKAPAAPVSAALVKQLREETGAGMMDCKKALSETGGDLEKAQEYLRKKGLSSADKKSSRIAAEGRIGSYIHDSRIGVLIEVNCETDFVGRSEKFKELVDD
LAMQVVACPQVRFVSVEDVPESVIMKEKELEMQREDLQSKPENIREKIVEGRMSKMLGELALLEQPFIKDDKLLVKDLVKQTIAALGENIKVRRFVKFTL
GEAIEDANPEA
|
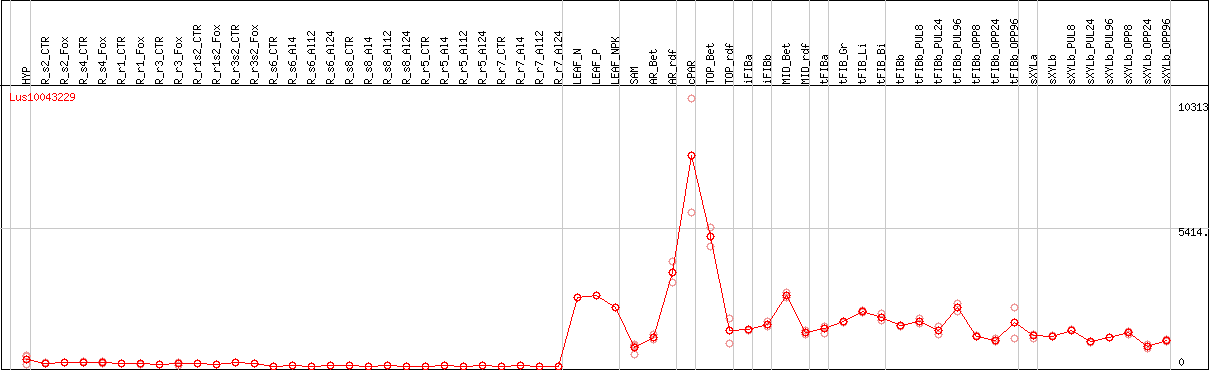
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10043229 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
