External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G27040 151 / 2e-42
OCP11, AGO4
OVEREXPRESSOR OF CATIONIC PEROXIDASE 11, ARGONAUTE 4, Argonaute family protein (.1.2)
AT5G21150 142 / 2e-39
AGO9
ARGONAUTE 9, Argonaute family protein (.1)
AT2G32940 130 / 5e-35
AGO6
ARGONAUTE 6, Argonaute family protein (.1)
AT5G21030 117 / 2e-30
PAZ domain-containing protein / piwi domain-containing protein (.1)
AT2G27880 59 / 3e-10
AGO5
ARGONAUTE 5, Argonaute family protein (.1)
AT5G43810 54 / 1e-08
PNH, ZLL, AGO10
ZWILLE, PINHEAD, ARGONAUTE 10, Stabilizer of iron transporter SufD / Polynucleotidyl transferase (.1.2)
AT1G48410 51 / 1e-07
AGO1
ARGONAUTE 1, Stabilizer of iron transporter SufD / Polynucleotidyl transferase (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10015155
273 / 1e-86
AT2G32940 1069 / 0.0
ARGONAUTE 6, Argonaute family protein (.1)
Lus10035331
221 / 1e-67
AT2G32940 1080 / 0.0
ARGONAUTE 6, Argonaute family protein (.1)
Lus10025537
147 / 4e-41
AT5G21030 1234 / 0.0
PAZ domain-containing protein / piwi domain-containing protein (.1)
Lus10000295
135 / 3e-38
AT2G27040 514 / 2e-175
OVEREXPRESSOR OF CATIONIC PEROXIDASE 11, ARGONAUTE 4, Argonaute family protein (.1.2)
Lus10026750
70 / 5e-14
AT5G21150 1253 / 0.0
ARGONAUTE 9, Argonaute family protein (.1)
Lus10014386
69 / 9e-14
AT2G27880 773 / 0.0
ARGONAUTE 5, Argonaute family protein (.1)
Lus10031904
59 / 2e-10
AT1G48410 1481 / 0.0
ARGONAUTE 1, Stabilizer of iron transporter SufD / Polynucleotidyl transferase (.1.2.3)
Lus10031331
59 / 2e-10
AT1G48410 1621 / 0.0
ARGONAUTE 1, Stabilizer of iron transporter SufD / Polynucleotidyl transferase (.1.2.3)
Lus10017983
59 / 3e-10
AT1G48410 1602 / 0.0
ARGONAUTE 1, Stabilizer of iron transporter SufD / Polynucleotidyl transferase (.1.2.3)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.014G159400
202 / 1e-60
AT2G32940 1169 / 0.0
ARGONAUTE 6, Argonaute family protein (.1)
Potri.001G219700
160 / 1e-45
AT2G27040 1403 / 0.0
OVEREXPRESSOR OF CATIONIC PEROXIDASE 11, ARGONAUTE 4, Argonaute family protein (.1.2)
Potri.006G025900
159 / 3e-45
AT2G27040 1236 / 0.0
OVEREXPRESSOR OF CATIONIC PEROXIDASE 11, ARGONAUTE 4, Argonaute family protein (.1.2)
Potri.016G024200
151 / 2e-42
AT2G27040 1246 / 0.0
OVEREXPRESSOR OF CATIONIC PEROXIDASE 11, ARGONAUTE 4, Argonaute family protein (.1.2)
Potri.008G010500
140 / 1e-38
AT2G27040 1281 / 0.0
OVEREXPRESSOR OF CATIONIC PEROXIDASE 11, ARGONAUTE 4, Argonaute family protein (.1.2)
Potri.009G001500
65 / 2e-12
AT2G27880 1192 / 0.0
ARGONAUTE 5, Argonaute family protein (.1)
Potri.001G213700
60 / 1e-10
AT2G27880 1214 / 0.0
ARGONAUTE 5, Argonaute family protein (.1)
Potri.015G029000
59 / 2e-10
AT1G48410 1549 / 0.0
ARGONAUTE 1, Stabilizer of iron transporter SufD / Polynucleotidyl transferase (.1.2.3)
Potri.012G037100
59 / 4e-10
AT1G48410 1560 / 0.0
ARGONAUTE 1, Stabilizer of iron transporter SufD / Polynucleotidyl transferase (.1.2.3)
Potri.006G118600
52 / 6e-08
AT2G27880 1068 / 0.0
ARGONAUTE 5, Argonaute family protein (.1)
PFAM info
Representative CDS sequence
>Lus10043291 pacid=23168150 polypeptide=Lus10043291 locus=Lus10043291.g ID=Lus10043291.BGIv1.0 annot-version=v1.0
ATGTCGAAAGTATCAACTGAATCCTCTCAGCTGCCTCCTCCAAAGCACTCGATTGTGAGTAGAAAGGGGTTTGGAACATTTGGGCGTCGGATCCCTTTGC
TTACGAACCACTTCAGAGTTTCTGTTTCGATTAAGTATGAAGACGACAGACCTGTCGAAGGGAAGGGAATTGGGAGAAAGCTCATTGACAAGCTTTACCA
GACCTATTCATCTGAGTTTGCTGGTAAAAGATTTGCGTACGACGGAGAAAAGGCCCTCTACACTGTAGGTCCTTTACCTCAGAAGAAGCTGGCGTTCACA
GTTGTTCTTGAGGATGCTATCGCCAAAAGCGAACATGGAAGCCCCCAGTCAACAAGTAAGCGGTCCAAGCATTCTATAAGATCAAAGACTTTTAAAGTAG
AGATCAATTATGCTGCCATGATCCCCATGAAGCCAATAGCTCTTGCCCTCAAAGGAGTTGATGCCGATAATGGTGGCATTCAGGATGCAGGGGTGCCTTT
TGGTGAGACAGACAATTCTTTCACGATGATTCAAGGAACTACACTGATGTTGGAGAGGGTATAA
AA sequence
>Lus10043291 pacid=23168150 polypeptide=Lus10043291 locus=Lus10043291.g ID=Lus10043291.BGIv1.0 annot-version=v1.0
MSKVSTESSQLPPPKHSIVSRKGFGTFGRRIPLLTNHFRVSVSIKYEDDRPVEGKGIGRKLIDKLYQTYSSEFAGKRFAYDGEKALYTVGPLPQKKLAFT
VVLEDAIAKSEHGSPQSTSKRSKHSIRSKTFKVEINYAAMIPMKPIALALKGVDADNGGIQDAGVPFGETDNSFTMIQGTTLMLERV
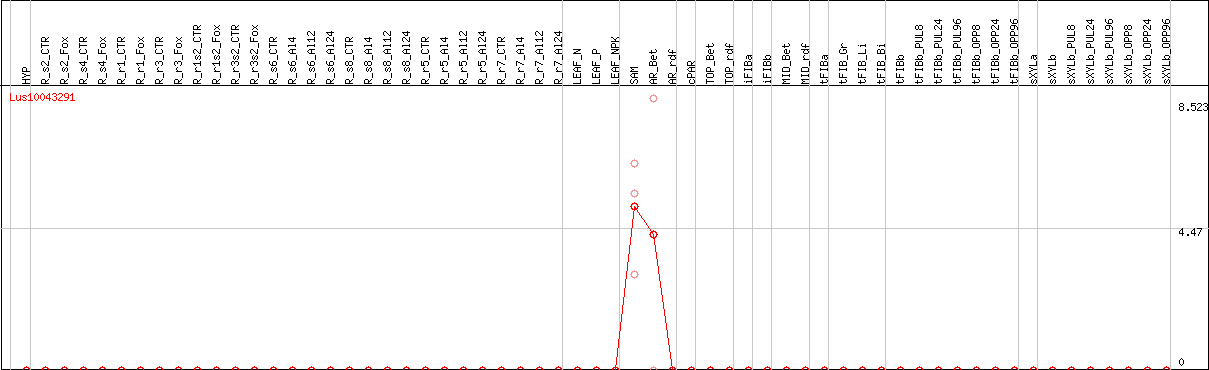
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10043291 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.