External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G07040 114 / 5e-29
RPS3, RPM1
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
AT1G63350 51 / 3e-07
Disease resistance protein (CC-NBS-LRR class) family (.1)
AT1G58400 49 / 2e-06
Disease resistance protein (CC-NBS-LRR class) family (.1)
AT5G44700 46 / 1e-05
GSO2, EDA23
GASSHO 2, EMBRYO SAC DEVELOPMENT ARREST 23, Leucine-rich repeat transmembrane protein kinase (.1)
AT1G53350 43 / 0.0001
Disease resistance protein (CC-NBS-LRR class) family (.1)
AT1G27180 42 / 0.0002
disease resistance protein (TIR-NBS-LRR class), putative (.1)
AT3G44480 42 / 0.0003
COG1, RPP10, RPP1
recognition of peronospora parasitica 1, Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT2G19330 41 / 0.0004
PIRL6
plant intracellular ras group-related LRR 6 (.1)
AT3G46730 41 / 0.0004
NB-ARC domain-containing disease resistance protein (.1)
AT1G59780 41 / 0.0006
NB-ARC domain-containing disease resistance protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10041243
294 / 1e-93
AT3G07040 526 / 1e-172
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Lus10002945
227 / 1e-68
AT3G07040 580 / 0.0
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Lus10028639
221 / 1e-66
AT3G07040 579 / 0.0
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Lus10020016
217 / 4e-65
AT3G07040 563 / 0.0
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Lus10018937
215 / 3e-64
AT3G07040 587 / 0.0
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Lus10014712
206 / 7e-61
AT3G07040 492 / 3e-159
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Lus10003094
126 / 5e-34
AT3G07040 169 / 4e-46
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Lus10038983
124 / 3e-32
AT3G07040 468 / 5e-150
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Lus10027281
120 / 5e-31
AT3G07040 459 / 5e-147
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.001G134500
136 / 1e-36
AT3G07040 422 / 1e-132
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Potri.001G134450
132 / 6e-35
AT3G07040 488 / 1e-157
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Potri.001G134300
131 / 1e-34
AT3G07040 487 / 2e-157
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Potri.001G134701
130 / 1e-34
AT3G07040 464 / 6e-149
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Potri.001G134601
128 / 1e-33
AT3G07040 443 / 2e-142
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Potri.001G134400
123 / 6e-32
AT3G07040 427 / 3e-134
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Potri.001G134475
123 / 6e-32
AT3G07040 413 / 1e-129
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Potri.002G216800
121 / 2e-31
AT3G07040 573 / 0.0
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Potri.003G099100
117 / 5e-30
AT3G07040 482 / 8e-156
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Potri.005G009101
103 / 1e-25
AT3G07040 153 / 3e-40
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
PFAM info
Representative CDS sequence
>Lus10043373 pacid=23168242 polypeptide=Lus10043373 locus=Lus10043373.g ID=Lus10043373.BGIv1.0 annot-version=v1.0
ATGGATAGCTTGTTCAACTTAAGGTTCTTGAACTTGTTTGGCACTCAAGTGAAAAGACTACCAAACTCTATAGGTAAGTTACAAAACCTTCAGACTCTAG
ATATCATAAGTACCAAAGTAGCAACTCTTCCCAAGGAAATCATGAAGCTGCAGAAGCTACGTCACCTAGTTACAGGCTATGTGAAACGTACGACGACATC
GAAGGAGTACAATGTAATGTATGGTATCTGTGTTTTGTCCGGATCGGATATCAGTTGTTTAAAGAACTTACAATCTCTACGTTTAGTGGAAGCAAGTAGG
GACTTGATAAAGCAACTGGGACAGATGACACAGCTAGTAGTACTGGGAATCACAAACGTTAAAGGGAAAAGTCAAATGGACGACTTGTGCTGCTCAATAA
AGAACATGCCTTTCCTGCAGCAGTTAAGCATGGCAGCTGGGAACGAGGATGAGATCCTTCAGTTCGACGAACTCGGTCAATCTCCTCCGTTGTCACGTCT
TCAATATCTCAGCTTGTATGGGAAGTTAGACAAGTTCCCTAATTGGGTTTCCTCCCTGCCACGTCTCAGTGATTTGCAGATGCACTGGAGCAGACTGCCA
AAGGAAGAGGACCCACTGCAGCATTTGCATAGTTTGCGGAACTTGGGCAATATTGTTCTTAACAATGCCTATCAAGGGGACAGAGTTAAATTGTCCAAGA
GGCTTTGA
AA sequence
>Lus10043373 pacid=23168242 polypeptide=Lus10043373 locus=Lus10043373.g ID=Lus10043373.BGIv1.0 annot-version=v1.0
MDSLFNLRFLNLFGTQVKRLPNSIGKLQNLQTLDIISTKVATLPKEIMKLQKLRHLVTGYVKRTTTSKEYNVMYGICVLSGSDISCLKNLQSLRLVEASR
DLIKQLGQMTQLVVLGITNVKGKSQMDDLCCSIKNMPFLQQLSMAAGNEDEILQFDELGQSPPLSRLQYLSLYGKLDKFPNWVSSLPRLSDLQMHWSRLP
KEEDPLQHLHSLRNLGNIVLNNAYQGDRVKLSKRL
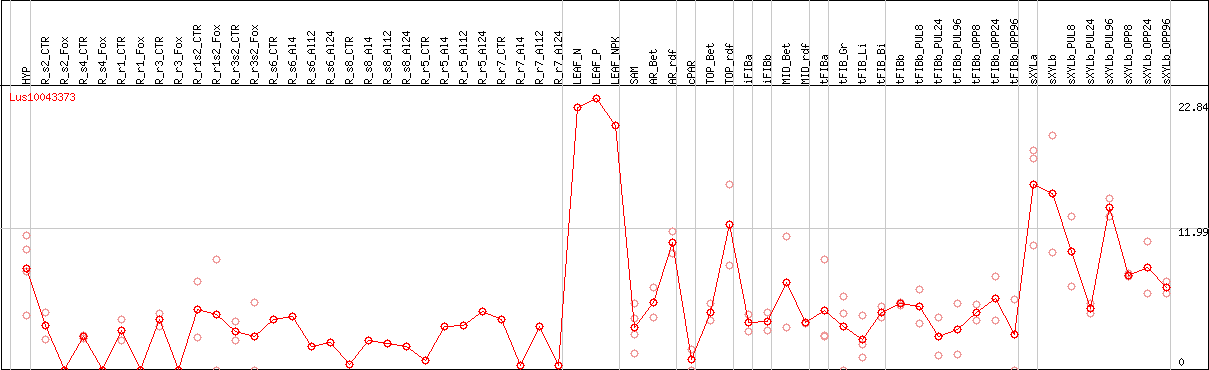
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10043373 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.