External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G44460 75 / 2e-15
bZIP
DPBF2, ATBZIP67
basic leucine zipper transcription factor 67, Basic-leucine zipper (bZIP) transcription factor family protein (.1)
AT3G56850 60 / 3e-10
bZIP
DPBF3, AREB3
ABA-responsive element binding protein 3 (.1)
AT4G34000 60 / 4e-10
bZIP
AtABF3, ABF3, DPBF5
DC3 PROMOTER-BINDING FACTOR 5, abscisic acid responsive elements-binding factor 3 (.1.2.3)
AT1G49720 53 / 9e-08
bZIP
ABF1
abscisic acid responsive element-binding factor 1 (.1.2)
AT2G36270 50 / 7e-07
bZIP
EEL, GIA1, ABI5
GROWTH-INSENSITIVITY TO ABA 1, ABA INSENSITIVE 5, Basic-leucine zipper (bZIP) transcription factor family protein (.1)
AT1G45249 50 / 1e-06
bZIP
AtABF2, ATAREB1, AREB1, ABF2
ABSCISIC ACID RESPONSIVE ELEMENTS-BINDING PROTEIN 2, abscisic acid responsive elements-binding factor 2 (.1.2.3)
AT2G41070 48 / 3e-06
bZIP
DPBF4, ATBZIP12, EEL
ENHANCED EM LEVEL, Basic-leucine zipper (bZIP) transcription factor family protein (.1), Basic-leucine zipper (bZIP) transcription factor family protein (.2), Basic-leucine zipper (bZIP) transcription factor family protein (.3)
AT5G42910 42 / 0.0003
bZIP
Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10019542
254 / 3e-84
AT3G44460 107 / 2e-27
basic leucine zipper transcription factor 67, Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Lus10017049
72 / 4e-14
AT2G36270 321 / 1e-106
GROWTH-INSENSITIVITY TO ABA 1, ABA INSENSITIVE 5, Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Lus10021369
68 / 9e-13
AT2G36270 322 / 2e-107
GROWTH-INSENSITIVITY TO ABA 1, ABA INSENSITIVE 5, Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Lus10022720
66 / 7e-12
AT2G36270 312 / 2e-103
GROWTH-INSENSITIVITY TO ABA 1, ABA INSENSITIVE 5, Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Lus10014194
59 / 1e-09
AT2G36270 342 / 2e-114
GROWTH-INSENSITIVITY TO ABA 1, ABA INSENSITIVE 5, Basic-leucine zipper (bZIP) transcription factor family protein (.1)
Lus10028889
53 / 9e-08
AT3G56850 273 / 7e-91
ABA-responsive element binding protein 3 (.1)
Lus10008929
52 / 1e-07
AT3G56850 214 / 3e-68
ABA-responsive element binding protein 3 (.1)
Lus10014066
51 / 5e-07
AT3G19290 345 / 1e-115
ABA-RESPONSIVE ELEMENT BINDING PROTEIN 2, ABRE binding factor 4 (.1.2.3)
Lus10008928
49 / 7e-07
AT3G56850 137 / 4e-40
ABA-responsive element binding protein 3 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0018
bZIP
PF00170
bZIP_1
bZIP transcription factor
Representative CDS sequence
>Lus10043386 pacid=23168037 polypeptide=Lus10043386 locus=Lus10043386.g ID=Lus10043386.BGIv1.0 annot-version=v1.0
ATGATGAGTGCTGTTCCTAAAGACAAGTCTTTCTGGACTCTGACGCTGAATGAAATTGAACGCGAGAATGGGAGGAACTTAGGGTCCATGAACATGGATG
ATTTTTTGACTCACTTGTTGAATATCAATAATCTTGATGACGAAAATCGTCAGGTTCAACAGCCTCAACTACCACATTACAACATTTATAATAATAATAG
TGGTTATGATCTGCAGCTGCCCAGAAGACCCTTAATGGCCGGTGCTCCTCCATTACTGCAGCAGCAACAACCTTGTTTTCATATACCTTCCCCTTTCTGT
AAGAAGACAGTGGGCGACGTGTGGTTCGAGATCCAGAGGGAGCAGAATCCGGCGAACAATGTTGCCGTTGTTCCTCCTCCCGAGAGGCAGCAGTCTTTAG
GGGAAATGACGCTGGAGGATTTCCTGGTGCTAGCTGGTGTGGTTCGGAAGAAAGCAGAACTACAGCCAACATTAGATCAGGTGCAAATCGAAACAGTACC
GTCGACGCCATTAGAAGAAGTAATGATCCCGGTAGAAAATGTTGCGCCCAAGGCTAGATGTTTCGCGGTGGAGAGAAGGCATCGCAGGATGATTAAGAAC
AGAGAATCTGCTGCCAGATCCAGGGCTAGAAGACAGGCATACACATTAGAGTTGGAACTTGAGTTGGATTCTCTCAAGGAAGAGAATATTAGACTCAAGC
AAACATTGGTATGCATTTATCTCTCTCTATATCATTCATGGTGCACGCTTCAGTACCCCGGATTAGAGGTACGGATACCTCCTTACGTGGCCGTCATATC
GTTGATATCAGTAAGAAGGGTATTGAATTTCGATATCCCCATCGTGGTTGCCACCTAG
AA sequence
>Lus10043386 pacid=23168037 polypeptide=Lus10043386 locus=Lus10043386.g ID=Lus10043386.BGIv1.0 annot-version=v1.0
MMSAVPKDKSFWTLTLNEIERENGRNLGSMNMDDFLTHLLNINNLDDENRQVQQPQLPHYNIYNNNSGYDLQLPRRPLMAGAPPLLQQQQPCFHIPSPFC
KKTVGDVWFEIQREQNPANNVAVVPPPERQQSLGEMTLEDFLVLAGVVRKKAELQPTLDQVQIETVPSTPLEEVMIPVENVAPKARCFAVERRHRRMIKN
RESAARSRARRQAYTLELELELDSLKEENIRLKQTLVCIYLSLYHSWCTLQYPGLEVRIPPYVAVISLISVRRVLNFDIPIVVAT
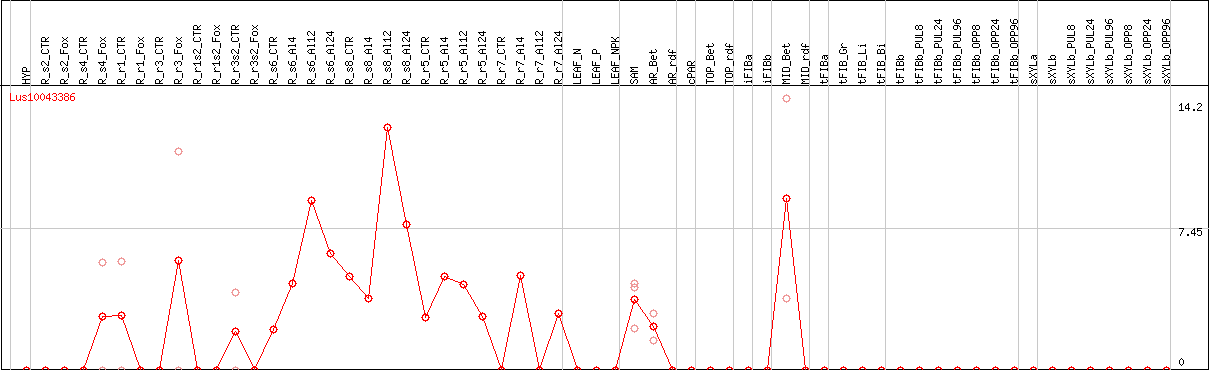
DESeq2's median of ratios [FLAX]
Coexpressed genes
Lus10043386 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.