Potri.001G004200 [POPLAR]

| External link |
|
|||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G004200.1 pacid=42788139 polypeptide=Potri.001G004200.1.p locus=Potri.001G004200 ID=Potri.001G004200.1.v4.1 annot-version=v4.1
ATGTGGACTAATGTATTCAAAATTGGTGGTCTTCATCATATATCATGGTTCCAGTTTCTCCCAAATGAATCTGATTTGAACTCTTTACCTGATAAGAGTG
TGAAGGTAGAGCAGAAAGATGTGGCTACATGTCTCGTGATTTTGGCTCATTTGCAGTTGCAGAAAGAGGGATTTCTAAGCACTTGGACCAATTCCTTTGT
TGGACCTTGGGATCCATCACAAGGTCTACATAATCCTGATGAGAAAATAAAGCTCTGGCTTTTTCTTCCTGGACGTTATTCATCTGTTGTTGAGAAGGCT
CAAGCTGCAGTGTCTAGATTAAGAGTTGTTGCATCTGGAATTTGGGTTGCTCCTGGGGACTCTGAAGAGGTTGCAGCAGCCCTTTCTCAATCTTTGAGGA
ATTACATAGAGAGAGCATTGGCCAGGCATTCATACATGCGGTTTGGAGATGTGTTTTTAAAGTACCATCCATCTCAAAGTGAAGGACTTTCCAGTAAAGG
ACAGCCCACAGTCGAGTTCATCTTTTCTGCCTCTGAAGAGACGATCTTTGTGCACGTGATAATATCTGCAAAGCATATCCGAGCACTTTCAAATGGTGAT
GTGGAAAGAGTGTTAAAACATTCTTCTACAAATTCCAGTTTTAGGCTTCCAGTGATTGCTTCTCCTCATGGAATTCGTGGTTCGCTGACAGGATGTTGTC
CAAGTGATCTTGTCAAGCAAGTGTATTTGAGTTCTGGCAAGTTTAGGACATTAAATGGATATATTGGCCTGCCATATCATGTTTCTCAAGGACCTGGCTG
TCAGCTGAGGGGGCAAAATTGCTATGTTGAAATTACACTAGGTTGCCCAAGGTCTGATAGTGACAAGGCATTGCAAACAAACTCACAGTCAACTAAGAAT
TCATCCAAGAATTATATTGTAGAATCTATTGCAGCAAGGAGAGGTGATCAGAAGGTATCGCCAGATCACTTATCAGCTCATGAGAAAGCATTTATTTATC
CTGCTGAAGCAGTTGTTGTTCCAGTCTTACAAAAATCTTTTGCTAGGTCTTCTTTGAAAAGATTTTGGCTGCAAAACTGGATAGGGCCATCACTACCTGG
TTCATCATTCTTTATGCATTGTGGTGGTGATACAGATTTTCTGGAAGGATCTTGGATTGAATCTAATGGAGTACGTATGCAGCACGGATATAACAGCAGT
AGCAATAGTAATAGTAGCAGCATCAGCTCCATAAGCAGCGGCTCTAGTGACAGTGATTACAAGATGACCACTGGTGAGCTTGAGGCTGATGCTGATTCAT
TGTCATGCAGACAATCTGGTTTATCTTCCAATGATCAGATGGAAAATGATGGCCTCAAATTGGGTTCTAAGCGTCCTAGAACAGGGATGACTGAGCCATT
TGGTCAAGCGGGCATGGTTAAAAATGTTCACATGCAAGATTTTGGTTCAGTAGAAGTTAATGATTCAGCTATCACAGGAATTGCAAATGATCAGATTGGA
TCTCGTTGGGATTGGGACGATGACAGAGGTGCGGGCATGGATATCCAAGCTCTTCTCTCTGAATTTGGTGATTTTGGGGACTTCTTTGAGAATGATGATC
TGCCTTTTGGAGAGCCTCCTGGAACAGCAGAGTCGCAGGCTCTTATGTTTTCTGGTCCTGATTGTGGTGAAGTTGCTAACAGTCCTATTGGGGTAATGGA
TGTTGTGGATCAGATGCTTCCGCCAGTAGCTTTTCCATCTTTTGAAAGTTTTAATCCGTCTCCTGCAGTAGTCATAGATGAAAGTGCCAGCAAAAGCCAA
GAAGACACACATGGTACTCTGGCATTAATTCCAGTGAACTGCACTCCATCATCTTCTTCAGGCGAGTTTGATCATTTAATCAAAGCCGAAGCTTTGATGA
CCTTTGCCCCAGAATATGGGGCAGTTGAAACCCCTACAAGTGAATTCTCCTCATCGATTTTCAGGAGTCCATATTGTCCAAAATCTCGCCAAGTGGAGAG
CTCAAATTCGAGTTCAAATAATTATGCATATGGTGCAACACCACCATCTTCTCCTTGCTTTGAAGGATCAAATGAGAAAACTGGCATACAGGTGAATCTG
AAAACAGGTCCTGGAAGGAATGACACAAAGAAATATTACACTCTTGTGGAAGGTGGGAAAGTACCACTTGATAGAAGAACATTAACATCTAATGAAAGCC
GTCCTACAAGTGAGGCGATGATGCCATCTCCATTATTGAATTCTAATTCATCAAATACTGTCAAATCTGCCCAGAGGAAAATGAGTGATGGCACACTAGG
AGCAGAAAGTTTCTTAGTTTCTATGAAGACTGTTCTGGCAACTGAAGTAGAATGCATTATGTTTCAAGCTTCCATGTGTAGAATGCGACACACACTGTTG
TCTCCTGGTAACCCTACATCTGTAAATTTGCATAGGTTAAGTGGAAGCACTGGTTTGAATCAGGTGCATGGTGATGCAAGTACCATGACAGACAACATAT
CTGGCAGGCATGAAGTGAAGAAAAAAGAATCTATACCAGTTAGAATAGCTGGGGATATGGATGGTGGAGTACTAGATGGGCATCTTAGTTCACCTGTAGG
TGTTTGGCGCTCTGTTGGAGTTCCTAAATTAACAAAACATACTAGTTCATCGAATATTGAAGTTAGTGTGTCCTTGCCACACCATTCATTCAGCGAGGAA
GGAGTTCTCTCATATAGGCAAAGGCAGCAGCCACTGCAGGAGCTTTTGGATGGCATGGCATTGCTTGTCCAACAAGCTACTTCCTTTGTTGATGTGGCTC
TGGATGCTGATTGTGGTGATGGTCCTTATGGTTGGCTTGCATTACAAGAACATTGGAGGCGTGGATTTTCTTGCGGGCCTTCCATGATCCATGCAGGTTG
TGGAGGGACTTTGGCTTCTTGTCATTTTCTGGACATTGCTGGCGTGGAACTAGTAGACCCACTTTCTGCTGATATTCATTCTTCTGCAGTGACCACCTTG
CTGCAGTCTGAAATCAAAATAGCCTTAAAATCTGCTTTTGGGAATTTGGAAGGTCCTTTATGTGTCACTGATTGGTGCAAAGGCCACATTCAGTCAGGTG
ATGGAGCAACCACATGCGATGGATCCTCTGGGGAGTCCACTTTAAGTGAATGCAAAGATTCAAGTACTGTAACACTGTCTGTTGGAGAACCAATGAGCCC
AGCCCTCTCTTCAGCTGCTGGATCATCTTCCCTCAAAGCATCCAGTACACCAGATGGAGCTAAAGTGGATGAGACATCCCAAAGGAAATCAAATCAAGAA
ATAGAGCCAGAGCTACTTCCCAGGATAAAGCCAACAGTTTTTGTTCTTCCTTTGCCTGCAATTCTTGTTGGGTACCAGGATGATTGGCTTAAGACATCGG
CAAGCTCTTTACAACTCTGGGAAAAGGCTCCTTTTGAACCATATGCTTCACCAAAACCAATAACTTATTGTGTTGTATGTCCAGACATTGATCCCCTTAC
TTCTGCAGCTGCAGATTTTTTCCAACAACTTGGAACTGTCTATGAAACGTGCAAACTGGGAACTCATTCACCTCAGAGTTTGGGAAACCAAATGGAGATG
GATGCAGGGAAATCTTTGTCTTCTGGTTTTGTACTACTTGATTGCCCTCAGTCAATGAAGATAGAAAGCAGTAATGCTTCCCTTGTGGGATCGATAAGCG
ATTATTTTCTCTCGCTCTCCAATGGTTGGGACCTGGCAAGCTATCTTAAGTCTCTTTCGAAGGCTGTCAAGGCTTTGAAAATTGGTCCATCCTTATCAAC
AAACCCAAAAGAAGGCAATAGCAGCCCTTGTATGGTTATCTATGTAGTGTGCCCCTTTCCTGAACCTGCTGCAGTTCTACAAACTGTCATTGAATCCTCT
GTTTCTATTGGGTCAATCATTCCCCCAGCAAATAGAGAGAGGAGGTCTATGTTGCTGGCTCAGGTCGGCAAAGCATTAAGCAGCTTGGCAGCTGTGGATG
AAGCGTCAGCATCAAATGTTCTAGTACTTTCTGGGTTTAGCATTCCTAAATTGGTATTGCAAATTGTGACAGTTGATGCCATTTTTAGGGTCACTAGTCC
AGCTCTTAATGAGCTCATCATTCTTAAGGAGACTGCTTTCACTGTATACAATAAGGCCCGGCGAATTTCAAAAGGCTCTTCAAATGATGTTCAGCCATCA
ACATCAAGTAGATCTCATTTAACACAAATGAGTTCTGTTCCAGCAATGTGGAATTCGTTACCAAGAGAGACTGACATTGATTCTAGATTGAGATCTGGCA
CTTGGGATAACTCCTGGCAAACAGCAAGGGCTGGAGGCTTGACCTGTGATCCAAACCGAAATGGAGATTTTTCTCTTCAAGATGAGATCCGCTACATGTT
TGAACCACTTTTTATTCTTTCAGAGCCCGGATCTCTGGACCATGCAGTTGCACCTACAATTATTGGTAATATGGTCTCTGAATCTTCAAAACTGCAATCT
GATGATACTGGTGGAAGCTTCATGCAAAGTGCAAGCTCAGCAGGGAGTGTGGATTCTGGATCTAGCTCACAACATGATGGATCTGAGCCCGATGGCTATG
GATCTAGTTATCAAAAGACTCTTCCAAGCCTGCATTGCTGCTATGGATGGACAGAGGACTGGCGTTGGCTTGTTTGCATCTGGACAGATGCAAGAGGAGA
ATTGCTTGACAGCCATATATTCCCTTTTGGTGGAATTAGTAGTCGCCAGGATACAAAAGGTTTGCAATGCCTTTTTGTACAAGTTTTGCAGCAAGGATGT
CAGATACTTCAGGCTTGCTCCTCTCCTGACACTGGTGGTGTAAAGCCTAGAGATTTTGTGATTACTCGAATTGGAAACTTCTTTGAGCTTGAGTACATAG
AGTGGCAGAAAGCCATTTATTCAGTAGGGGGTTCTGAGGTGAAGAAATGGCCACTGCAACTACGGCGATCAATGCCTGATGGGATGGCTGCAGGCGCTAA
TGGTGCTTCCTTGCAGCAACAGGAGATGGGTTTAATTCAAGAAAGAACCTTGCCTTCCTCACCTAGTCCGTTGTATAGCCCTCACTCAAAAGCATCTGGC
TATATGAAGGGTGGTTTGGGGCAAACTTCTTCAAGAAAACAGCTTATCGGTGGGCACGCAGCAGTTGACAACTCAAGGGGGATGCTTCAGTGGATGCTGA
GCATCAGTTTTGTTACAATTTCAATTGACCATTCTTTACATCTGTTGCTTCAGGCTGATACACCATCTCCTGGAGGAAGTGGCAGTGGTGTTGGCACGTC
AATCTATCTGGAAGGTATCACTCCTGTAAAGTCTCTCGGTTCAACATCAGCATCTTACATTTTAATTCCATCACCCAGCATGCGCTTCCTTCCGTCAACA
CCCCTGCAGCACCCCACATGCCTCACTGCTGAATCCCCACCACTTGCACATCTCCTTCATAGTAAGGGATCTGCAATTCCGCTCTCCACTGGTTTCGTGG
TTTCAAAGGCTGTACCTTCCGTGAGGAGAGATTATAAGAGCAATTCAAGAGAAGAATGGCCTTCTGTTCTTTCTGTGAGTCTCATTGATTATTATGGAGG
CAATAATATGACCCAAGATAAAATGTTTCGAAGGATTATGAAGCAGGGGGGACGGACTTTAGGTGTAGATGGTAAAGATTTTGAAATCAGGACCCAGGTA
ATTCTTGAGAGTGTCGCAGCAGAACTTCAAGCCCTATCATGGATGACTGTGAGCCCAGCATATTTGGAGCGACGAACCGCACTTCCATTTCATTGTGACA
TGGTTTTAAGACTCAGAAGACTGCTTCATTTTGCTGATAAAGAGCTCTCTTCACAGCCTGGTCGGTCACAGGTATAA
|
|||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G004200.1 pacid=42788139 polypeptide=Potri.001G004200.1.p locus=Potri.001G004200 ID=Potri.001G004200.1.v4.1 annot-version=v4.1
MWTNVFKIGGLHHISWFQFLPNESDLNSLPDKSVKVEQKDVATCLVILAHLQLQKEGFLSTWTNSFVGPWDPSQGLHNPDEKIKLWLFLPGRYSSVVEKA
QAAVSRLRVVASGIWVAPGDSEEVAAALSQSLRNYIERALARHSYMRFGDVFLKYHPSQSEGLSSKGQPTVEFIFSASEETIFVHVIISAKHIRALSNGD
VERVLKHSSTNSSFRLPVIASPHGIRGSLTGCCPSDLVKQVYLSSGKFRTLNGYIGLPYHVSQGPGCQLRGQNCYVEITLGCPRSDSDKALQTNSQSTKN
SSKNYIVESIAARRGDQKVSPDHLSAHEKAFIYPAEAVVVPVLQKSFARSSLKRFWLQNWIGPSLPGSSFFMHCGGDTDFLEGSWIESNGVRMQHGYNSS
SNSNSSSISSISSGSSDSDYKMTTGELEADADSLSCRQSGLSSNDQMENDGLKLGSKRPRTGMTEPFGQAGMVKNVHMQDFGSVEVNDSAITGIANDQIG
SRWDWDDDRGAGMDIQALLSEFGDFGDFFENDDLPFGEPPGTAESQALMFSGPDCGEVANSPIGVMDVVDQMLPPVAFPSFESFNPSPAVVIDESASKSQ
EDTHGTLALIPVNCTPSSSSGEFDHLIKAEALMTFAPEYGAVETPTSEFSSSIFRSPYCPKSRQVESSNSSSNNYAYGATPPSSPCFEGSNEKTGIQVNL
KTGPGRNDTKKYYTLVEGGKVPLDRRTLTSNESRPTSEAMMPSPLLNSNSSNTVKSAQRKMSDGTLGAESFLVSMKTVLATEVECIMFQASMCRMRHTLL
SPGNPTSVNLHRLSGSTGLNQVHGDASTMTDNISGRHEVKKKESIPVRIAGDMDGGVLDGHLSSPVGVWRSVGVPKLTKHTSSSNIEVSVSLPHHSFSEE
GVLSYRQRQQPLQELLDGMALLVQQATSFVDVALDADCGDGPYGWLALQEHWRRGFSCGPSMIHAGCGGTLASCHFLDIAGVELVDPLSADIHSSAVTTL
LQSEIKIALKSAFGNLEGPLCVTDWCKGHIQSGDGATTCDGSSGESTLSECKDSSTVTLSVGEPMSPALSSAAGSSSLKASSTPDGAKVDETSQRKSNQE
IEPELLPRIKPTVFVLPLPAILVGYQDDWLKTSASSLQLWEKAPFEPYASPKPITYCVVCPDIDPLTSAAADFFQQLGTVYETCKLGTHSPQSLGNQMEM
DAGKSLSSGFVLLDCPQSMKIESSNASLVGSISDYFLSLSNGWDLASYLKSLSKAVKALKIGPSLSTNPKEGNSSPCMVIYVVCPFPEPAAVLQTVIESS
VSIGSIIPPANRERRSMLLAQVGKALSSLAAVDEASASNVLVLSGFSIPKLVLQIVTVDAIFRVTSPALNELIILKETAFTVYNKARRISKGSSNDVQPS
TSSRSHLTQMSSVPAMWNSLPRETDIDSRLRSGTWDNSWQTARAGGLTCDPNRNGDFSLQDEIRYMFEPLFILSEPGSLDHAVAPTIIGNMVSESSKLQS
DDTGGSFMQSASSAGSVDSGSSSQHDGSEPDGYGSSYQKTLPSLHCCYGWTEDWRWLVCIWTDARGELLDSHIFPFGGISSRQDTKGLQCLFVQVLQQGC
QILQACSSPDTGGVKPRDFVITRIGNFFELEYIEWQKAIYSVGGSEVKKWPLQLRRSMPDGMAAGANGASLQQQEMGLIQERTLPSSPSPLYSPHSKASG
YMKGGLGQTSSRKQLIGGHAAVDNSRGMLQWMLSISFVTISIDHSLHLLLQADTPSPGGSGSGVGTSIYLEGITPVKSLGSTSASYILIPSPSMRFLPST
PLQHPTCLTAESPPLAHLLHSKGSAIPLSTGFVVSKAVPSVRRDYKSNSREEWPSVLSVSLIDYYGGNNMTQDKMFRRIMKQGGRTLGVDGKDFEIRTQV
ILESVAAELQALSWMTVSPAYLERRTALPFHCDMVLRLRRLLHFADKELSSQPGRSQV
|
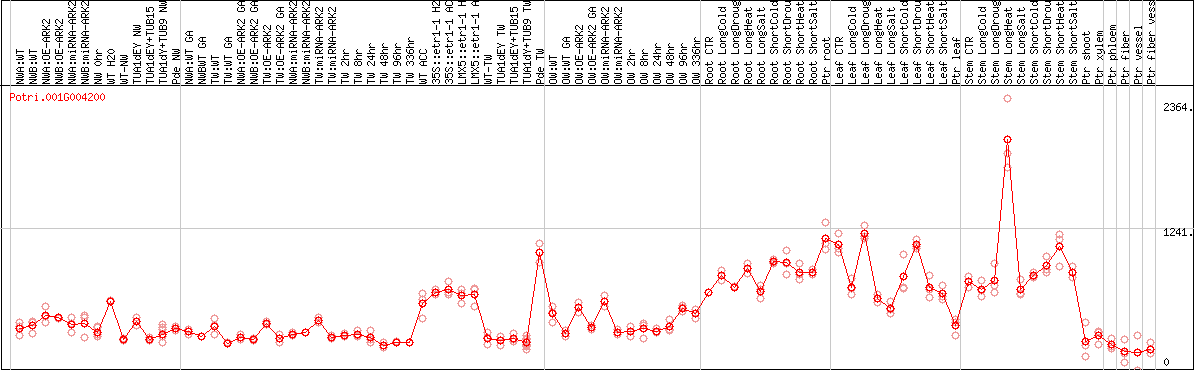
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G004200 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
