External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G19760 52 / 4e-09
Mitochondrial substrate carrier family protein (.1)
AT2G22500 41 / 3e-05
UCP5, ATPUMP5, DIC1
DICARBOXYLATE CARRIER 1, PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 5, uncoupling protein 5 (.1)
AT5G46800 40 / 6e-05
BOU
A BOUT DE SOUFFLE, Mitochondrial substrate carrier family protein (.1)
AT4G24570 39 / 0.0003
DIC2
dicarboxylate carrier 2 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.003G220400
52 / 5e-09
AT5G19760 498 / 6e-180
Mitochondrial substrate carrier family protein (.1)
Potri.001G004500
50 / 1e-08
AT5G19760 519 / 0.0
Mitochondrial substrate carrier family protein (.1)
Potri.005G157300
43 / 8e-06
AT2G22500 412 / 2e-145
DICARBOXYLATE CARRIER 1, PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 5, uncoupling protein 5 (.1)
Potri.002G104400
42 / 2e-05
AT2G22500 427 / 1e-151
DICARBOXYLATE CARRIER 1, PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 5, uncoupling protein 5 (.1)
Potri.017G045200
41 / 2e-05
AT2G22500 286 / 3e-98
DICARBOXYLATE CARRIER 1, PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 5, uncoupling protein 5 (.1)
Potri.017G045100
41 / 4e-05
AT2G22500 451 / 6e-161
DICARBOXYLATE CARRIER 1, PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 5, uncoupling protein 5 (.1)
Potri.017G045150
41 / 4e-05
AT2G22500 457 / 3e-163
DICARBOXYLATE CARRIER 1, PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 5, uncoupling protein 5 (.1)
Potri.017G047800
39 / 0.0001
AT4G24570 435 / 2e-154
dicarboxylate carrier 2 (.1)
Potri.001G142600
39 / 0.0002
AT5G46800 434 / 1e-154
A BOUT DE SOUFFLE, Mitochondrial substrate carrier family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10007883
54 / 1e-09
AT5G19760 536 / 0.0
Mitochondrial substrate carrier family protein (.1)
Lus10030361
52 / 3e-09
AT5G19760 489 / 9e-177
Mitochondrial substrate carrier family protein (.1)
Lus10009270
42 / 5e-06
AT2G22500 204 / 8e-67
DICARBOXYLATE CARRIER 1, PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 5, uncoupling protein 5 (.1)
Lus10015897
41 / 3e-05
AT4G24570 184 / 2e-56
dicarboxylate carrier 2 (.1)
Lus10025725
41 / 4e-05
AT2G22500 310 / 2e-106
DICARBOXYLATE CARRIER 1, PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 5, uncoupling protein 5 (.1)
Lus10035936
41 / 5e-05
AT4G24570 209 / 5e-67
dicarboxylate carrier 2 (.1)
Lus10024769
39 / 0.0003
AT4G03115 401 / 3e-141
Mitochondrial substrate carrier family protein (.1)
PFAM info
Representative CDS sequence
>Potri.001G004366.1 pacid=42793614 polypeptide=Potri.001G004366.1.p locus=Potri.001G004366 ID=Potri.001G004366.1.v4.1 annot-version=v4.1
ATGAGACCCAATGCTGATGGAAAGTCCCCGTTCACAAGCTCCCTGGACTGTGCCCTGAAAGTCCTGAAAGCCAGAGGGCTTGTGGGACTCTACAAAGGAT
TTCCACTGTTTCTTTGCAGGAGTTTACCTGCTACCACGATCACATGCATGGATGATGTTAGAAGATATCCGGGACTTCGAGGAATTTGCTGGATTGTAGT
AGCTCAAGAGTATCCAAGGAGGAGCTGCTCCCTTGATCCTACTCATAATTGTAGTATTTGCAAGCAAGGTTTTAACTTGTTAAGAAAACTGTGTTGCCAG
TAA
AA sequence
>Potri.001G004366.1 pacid=42793614 polypeptide=Potri.001G004366.1.p locus=Potri.001G004366 ID=Potri.001G004366.1.v4.1 annot-version=v4.1
MRPNADGKSPFTSSLDCALKVLKARGLVGLYKGFPLFLCRSLPATTITCMDDVRRYPGLRGICWIVVAQEYPRRSCSLDPTHNCSICKQGFNLLRKLCCQ
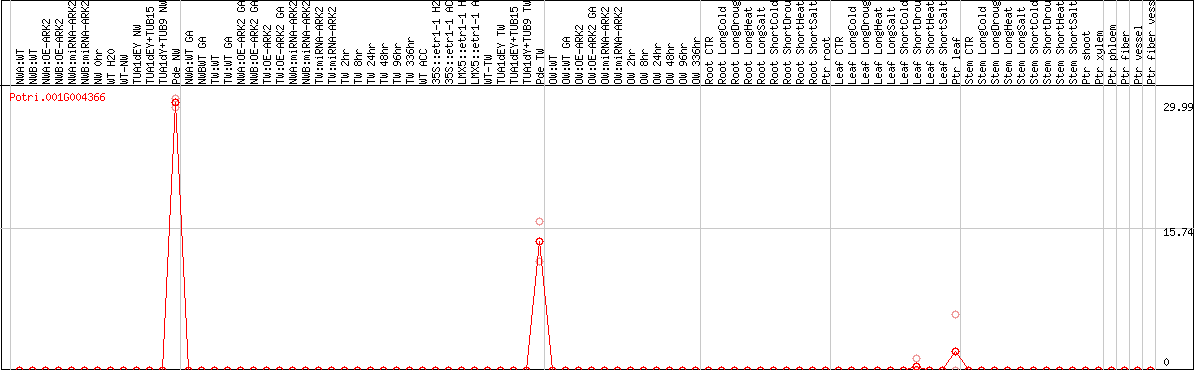
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G004366 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.