External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G58170 238 / 8e-81
Disease resistance-responsive (dirigent-like protein) family protein (.1)
AT1G55210 228 / 1e-76
Disease resistance-responsive (dirigent-like protein) family protein (.1), Disease resistance-responsive (dirigent-like protein) family protein (.2)
AT5G49040 223 / 7e-75
Disease resistance-responsive (dirigent-like protein) family protein (.1)
AT3G13650 218 / 6e-73
Disease resistance-responsive (dirigent-like protein) family protein (.1)
AT5G49030 197 / 3e-58
OVA2
ovule abortion 2, tRNA synthetase class I (I, L, M and V) family protein (.1), tRNA synthetase class I (I, L, M and V) family protein (.2), tRNA synthetase class I (I, L, M and V) family protein (.3)
AT1G65870 179 / 2e-57
Disease resistance-responsive (dirigent-like protein) family protein (.1)
AT3G13662 173 / 3e-55
Disease resistance-responsive (dirigent-like protein) family protein (.1)
AT5G42500 166 / 2e-52
Disease resistance-responsive (dirigent-like protein) family protein (.1)
AT5G42510 163 / 3e-51
Disease resistance-responsive (dirigent-like protein) family protein (.1)
AT3G13660 160 / 7e-51
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.003G216200
323 / 3e-114
AT1G55210 239 / 2e-81
Disease resistance-responsive (dirigent-like protein) family protein (.1), Disease resistance-responsive (dirigent-like protein) family protein (.2)
Potri.003G216300
263 / 1e-90
AT1G55210 221 / 5e-74
Disease resistance-responsive (dirigent-like protein) family protein (.1), Disease resistance-responsive (dirigent-like protein) family protein (.2)
Potri.003G216400
262 / 5e-90
AT1G58170 225 / 1e-75
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Potri.016G061000
254 / 5e-87
AT1G58170 214 / 2e-71
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Potri.006G195300
235 / 2e-79
AT5G49040 202 / 2e-66
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Potri.016G060900
226 / 5e-76
AT1G58170 197 / 1e-64
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Potri.016G060700
221 / 8e-74
AT1G65870 186 / 2e-60
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Potri.016G060801
194 / 1e-63
AT1G58170 164 / 5e-52
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Potri.009G131000
184 / 3e-59
AT2G21100 236 / 4e-80
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10026176
239 / 6e-81
AT1G58170 207 / 1e-68
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Lus10042488
234 / 4e-79
AT1G58170 202 / 1e-66
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Lus10029306
225 / 2e-75
AT1G65870 208 / 5e-69
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Lus10017231
216 / 4e-72
AT1G58170 191 / 2e-62
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Lus10030483
195 / 2e-63
AT1G65870 164 / 1e-51
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Lus10021084
174 / 3e-55
AT5G42500 164 / 2e-51
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Lus10039561
170 / 1e-53
AT3G13650 157 / 5e-49
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Lus10016237
165 / 1e-52
AT1G58170 149 / 6e-47
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Lus10034476
159 / 2e-49
AT2G21100 220 / 1e-73
Disease resistance-responsive (dirigent-like protein) family protein (.1)
Lus10027834
156 / 6e-48
AT2G21100 182 / 5e-58
Disease resistance-responsive (dirigent-like protein) family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0650
AOC_barrel
PF03018
Dirigent
Dirigent-like protein
Representative CDS sequence
>Potri.001G009100.1 pacid=42790846 polypeptide=Potri.001G009100.1.p locus=Potri.001G009100 ID=Potri.001G009100.1.v4.1 annot-version=v4.1
ATGGCTAGATCAGTTCCCATCCTTGCTTCCAAATTCATCACTCTCTTACTCCTCTCTTCCTTTGCCACAATCTTGGTCACTGGAGACCAGGATCATGAAT
TTGTGAGAAGTTTGGACAGGAAGCTACTAGGGCTCAAGAAAGAAAAGCTAAGCCACTTCAAGTTATATTGGCATGACATCCTTACTGGCCAGAACCCCAG
CGCCGTCCAAGTTGTGCCACCACCATCGAACACATCAAGAACAGCTTTTGGGTTAGTGAGAATGATCGATAACCCATTAACCTTAGGGCCTGAAATGAGC
TCAAAGTTGGTAGGAAGGGCACAAGGGTTGTATGCACAAGCGTCGCAACAAGATATTGGGTTATTGATGGCCATGAACTTTGCTTTTATTGAAGGTAAGT
ATAATGGTAGCACTATTACTGTTCTAGGGAAGAACGCAGTGTTCTCGACGGTGAGAGAGATGCCGGTGATCGGAGGAAGCGGACTTTTCCGGTTTGCTAG
AGGTTATGTTCAGGCGAGAACTCACAAGCTCGACATGGCCACAGGAGACGCTACAGTTGAGTATAATGTCTACGTTTTTCATTATTGA
AA sequence
>Potri.001G009100.1 pacid=42790846 polypeptide=Potri.001G009100.1.p locus=Potri.001G009100 ID=Potri.001G009100.1.v4.1 annot-version=v4.1
MARSVPILASKFITLLLLSSFATILVTGDQDHEFVRSLDRKLLGLKKEKLSHFKLYWHDILTGQNPSAVQVVPPPSNTSRTAFGLVRMIDNPLTLGPEMS
SKLVGRAQGLYAQASQQDIGLLMAMNFAFIEGKYNGSTITVLGKNAVFSTVREMPVIGGSGLFRFARGYVQARTHKLDMATGDATVEYNVYVFHY
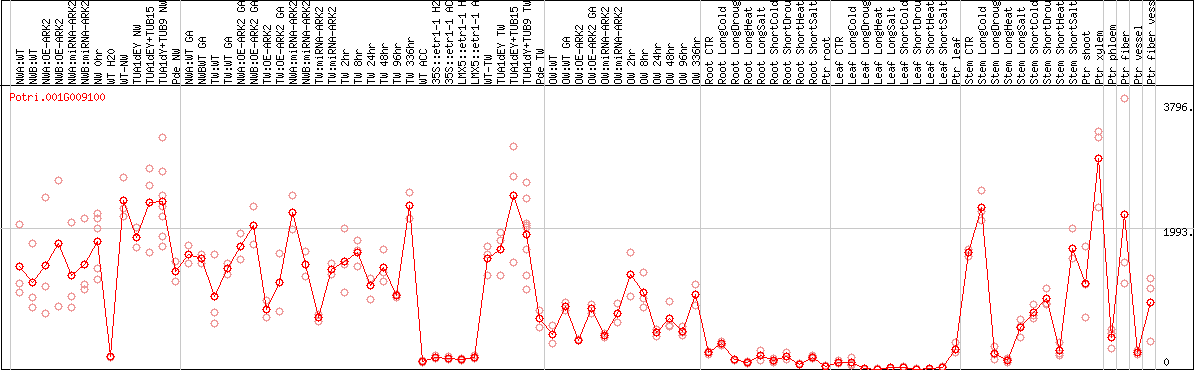
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G009100 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.