External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G38720 135 / 7e-38
MAP65-5
microtubule-associated protein 65-5 (.1)
AT5G55230 132 / 1e-36
ATMAP65-1
microtubule-associated proteins 65-1 (.1.2)
AT4G26760 124 / 6e-34
MAP65-2
microtubule-associated protein 65-2 (.1)
AT3G60840 123 / 2e-33
MAP65-4
microtubule-associated protein 65-4 (.1)
AT1G27920 115 / 5e-31
MAP65-8
microtubule-associated protein 65-8 (.1)
AT5G51600 110 / 4e-29
ATMAP65-3, PLE
PLEIADE, ARABIDOPSIS THALIANA MICROTUBULE-ASSOCIATED PROTEIN 65-3, Microtubule associated protein (MAP65/ASE1) family protein (.1)
AT2G01910 106 / 1e-27
ATMAP65-6
Microtubule associated protein (MAP65/ASE1) family protein (.1), Microtubule associated protein (MAP65/ASE1) family protein (.2)
AT1G14690 106 / 2e-27
MAP65-7
microtubule-associated protein 65-7 (.1.2)
AT5G62250 102 / 2e-26
MAP65-9
microtubule-associated protein 65-9 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.003G192400
187 / 2e-57
AT2G38720 555 / 0.0
microtubule-associated protein 65-5 (.1)
Potri.001G356500
130 / 2e-36
AT5G55230 853 / 0.0
microtubule-associated proteins 65-1 (.1.2)
Potri.011G092500
130 / 2e-36
AT5G55230 863 / 0.0
microtubule-associated proteins 65-1 (.1.2)
Potri.001G055100
121 / 7e-33
AT1G27920 623 / 0.0
microtubule-associated protein 65-8 (.1)
Potri.003G173300
120 / 1e-32
AT1G27920 655 / 0.0
microtubule-associated protein 65-8 (.1)
Potri.006G269800
109 / 1e-31
AT5G51600 186 / 2e-56
PLEIADE, ARABIDOPSIS THALIANA MICROTUBULE-ASSOCIATED PROTEIN 65-3, Microtubule associated protein (MAP65/ASE1) family protein (.1)
Potri.014G070100
116 / 5e-31
AT5G51600 451 / 5e-149
PLEIADE, ARABIDOPSIS THALIANA MICROTUBULE-ASSOCIATED PROTEIN 65-3, Microtubule associated protein (MAP65/ASE1) family protein (.1)
Potri.015G131400
113 / 4e-30
AT5G51600 728 / 0.0
PLEIADE, ARABIDOPSIS THALIANA MICROTUBULE-ASSOCIATED PROTEIN 65-3, Microtubule associated protein (MAP65/ASE1) family protein (.1)
Potri.005G057150
104 / 1e-29
AT5G51600 181 / 1e-54
PLEIADE, ARABIDOPSIS THALIANA MICROTUBULE-ASSOCIATED PROTEIN 65-3, Microtubule associated protein (MAP65/ASE1) family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10008705
164 / 1e-48
AT2G38720 565 / 0.0
microtubule-associated protein 65-5 (.1)
Lus10026114
163 / 2e-48
AT2G38720 553 / 0.0
microtubule-associated protein 65-5 (.1)
Lus10041635
132 / 7e-37
AT2G38720 550 / 0.0
microtubule-associated protein 65-5 (.1)
Lus10043178
132 / 2e-36
AT5G55230 847 / 0.0
microtubule-associated proteins 65-1 (.1.2)
Lus10032567
131 / 2e-36
AT5G55230 850 / 0.0
microtubule-associated proteins 65-1 (.1.2)
Lus10015789
124 / 6e-34
AT1G27920 673 / 0.0
microtubule-associated protein 65-8 (.1)
Lus10037691
120 / 2e-32
AT5G51600 768 / 0.0
PLEIADE, ARABIDOPSIS THALIANA MICROTUBULE-ASSOCIATED PROTEIN 65-3, Microtubule associated protein (MAP65/ASE1) family protein (.1)
Lus10015684
120 / 3e-32
AT5G51600 759 / 0.0
PLEIADE, ARABIDOPSIS THALIANA MICROTUBULE-ASSOCIATED PROTEIN 65-3, Microtubule associated protein (MAP65/ASE1) family protein (.1)
Lus10035099
117 / 2e-31
AT2G01910 822 / 0.0
Microtubule associated protein (MAP65/ASE1) family protein (.1), Microtubule associated protein (MAP65/ASE1) family protein (.2)
Lus10031939
117 / 2e-31
AT2G01910 838 / 0.0
Microtubule associated protein (MAP65/ASE1) family protein (.1), Microtubule associated protein (MAP65/ASE1) family protein (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF03999
MAP65_ASE1
Microtubule associated protein (MAP65/ASE1 family)
Representative CDS sequence
>Potri.001G032566.3 pacid=42789226 polypeptide=Potri.001G032566.3.p locus=Potri.001G032566 ID=Potri.001G032566.3.v4.1 annot-version=v4.1
ATGTCTAATCTGCTTGCGAGCATGGATGATCAAATCACAAAAGCCAACGAGCAAGCTCATAGTCGGAAGGATATCTTGGACAAAGTACAGAAATGGAAAT
TTGCATCCGAGGAAGAACAGTGGCTTGATGAATATGAAAAGGATGATAATCGGTATGGTGCAGGAAGAGGAGCACACAAAAATCTGAAACGTGCTGAGAA
AGCACGGATTCTGGCCAGCAAAACACCATCTGTGGTTGAAAATTTGACTTCTATAGTTAAAGCCTGGGAAATGGAGAGAAATATCCACTTCTTATGTGAC
AAGGCTCCCCTCTTACATACCTTAAGAGGAATATACCGTGCTAGGGCAGGAGAGGGAAGAGGAGAAGCGTCGATCTCGGGAGAAGAAACGTTTGCAAGAA
CGGTTTGCTGCTGA
AA sequence
>Potri.001G032566.3 pacid=42789226 polypeptide=Potri.001G032566.3.p locus=Potri.001G032566 ID=Potri.001G032566.3.v4.1 annot-version=v4.1
MSNLLASMDDQITKANEQAHSRKDILDKVQKWKFASEEEQWLDEYEKDDNRYGAGRGAHKNLKRAEKARILASKTPSVVENLTSIVKAWEMERNIHFLCD
KAPLLHTLRGIYRARAGEGRGEASISGEETFARTVCC
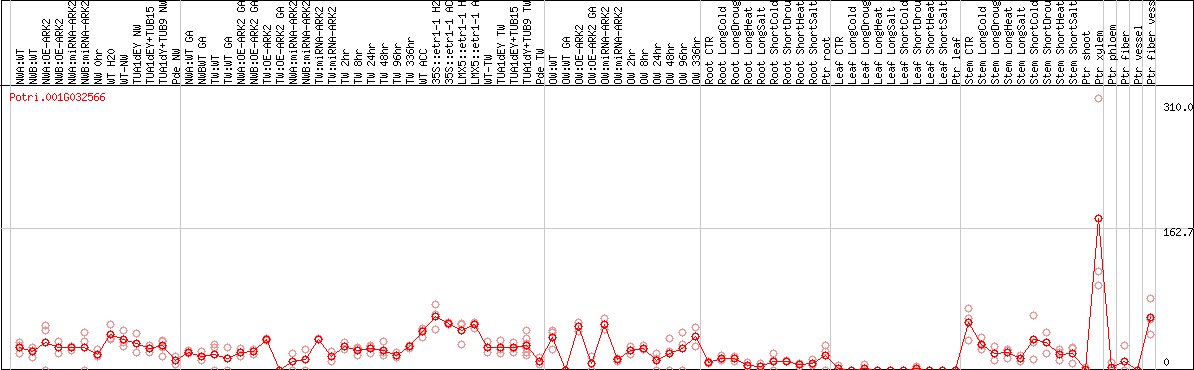
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G032566 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.