External link
Symbol
J8.1
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G80920 129 / 2e-38
AtToc12, AtJ8, J8
translocon at the outer envelope membrane of chloroplasts 12, Chaperone DnaJ-domain superfamily protein (.1)
AT5G59610 54 / 4e-09
Chaperone DnaJ-domain superfamily protein (.1.2)
AT2G41000 52 / 1e-08
Chaperone DnaJ-domain superfamily protein (.1.2)
AT2G33735 49 / 7e-08
Chaperone DnaJ-domain superfamily protein (.1)
AT3G08970 50 / 1e-07
TMS1, ATERDJ3A
THERMOSENSITIVE MALE STERILE 1, DNAJ heat shock N-terminal domain-containing protein (.1)
AT5G16650 48 / 1e-07
Chaperone DnaJ-domain superfamily protein (.1)
AT1G72070 48 / 2e-07
Chaperone DnaJ-domain superfamily protein (.1)
AT1G28210 48 / 9e-07
ATJ1
DNAJ heat shock family protein (.1.2)
AT4G39960 47 / 1e-06
Molecular chaperone Hsp40/DnaJ family protein (.1)
AT1G79940 47 / 2e-06
ATERDJ2A
DnaJ / Sec63 Brl domains-containing protein (.1.2.3.4)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10014352
132 / 5e-40
AT1G80920 135 / 2e-41
translocon at the outer envelope membrane of chloroplasts 12, Chaperone DnaJ-domain superfamily protein (.1)
Lus10026061
124 / 9e-37
AT1G80920 136 / 1e-41
translocon at the outer envelope membrane of chloroplasts 12, Chaperone DnaJ-domain superfamily protein (.1)
Lus10041671
124 / 2e-36
AT1G80920 138 / 2e-42
translocon at the outer envelope membrane of chloroplasts 12, Chaperone DnaJ-domain superfamily protein (.1)
Lus10024067
117 / 9e-34
AT1G80920 130 / 4e-39
translocon at the outer envelope membrane of chloroplasts 12, Chaperone DnaJ-domain superfamily protein (.1)
Lus10016510
63 / 5e-12
AT5G59610 252 / 2e-83
Chaperone DnaJ-domain superfamily protein (.1.2)
Lus10040777
61 / 3e-11
AT5G59610 246 / 6e-81
Chaperone DnaJ-domain superfamily protein (.1.2)
Lus10003380
57 / 7e-10
AT3G08970 615 / 0.0
THERMOSENSITIVE MALE STERILE 1, DNAJ heat shock N-terminal domain-containing protein (.1)
Lus10002852
56 / 2e-09
AT3G08970 600 / 0.0
THERMOSENSITIVE MALE STERILE 1, DNAJ heat shock N-terminal domain-containing protein (.1)
Lus10012702
51 / 7e-08
AT1G77930 368 / 2e-129
Chaperone DnaJ-domain superfamily protein (.1.2)
Lus10001296
51 / 7e-08
AT1G77930 370 / 2e-130
Chaperone DnaJ-domain superfamily protein (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0392
Chaperone-J
PF00226
DnaJ
DnaJ domain
Representative CDS sequence
>Potri.001G043100.1 pacid=42789356 polypeptide=Potri.001G043100.1.p locus=Potri.001G043100 ID=Potri.001G043100.1.v4.1 annot-version=v4.1
ATGGCCAGTGCAGCAGCTGTTGGGATGGTGGGAGGAAATGGGTGTGCGGGGTCTTCAGCTTCTTGGTTTCAGATCAAGAATAGGAGAAAGAAGAATAGTA
ATGATAAGATGGGGAGAGATGGAGTCAGGTTTTTTACTGCTGCTTCCTATTCTTCTCCTTCTGTCATGGACCCTTACAAGACTCTTAGGATCCAACCTGA
TGCTTCTGAATCTGAGGTCAAGAAAGCTTTTAGACAGCTTGCTTTGCAGTATCATCCAGATGTCTGCAGAGGAAGCAATTGTGGTGTGCAGTTTAGCCTG
ATCAATGAAGCCTATGATACTGTGATGAGTAATTTGAGGGAAGAACCAGATGAATCATCACAAATGTACATGTCTTATGAGCCGTCGGAACAGGGAATTG
ATGAGCCCATGAGAGGAATGAACGACACAGATTGGGATATGTGGGAGGAATGGATGGGATGGGAAGGGGCTGGAATTCGTGACTACTCATCCCATGTTAA
TCCTTACATTTAA
AA sequence
>Potri.001G043100.1 pacid=42789356 polypeptide=Potri.001G043100.1.p locus=Potri.001G043100 ID=Potri.001G043100.1.v4.1 annot-version=v4.1
MASAAAVGMVGGNGCAGSSASWFQIKNRRKKNSNDKMGRDGVRFFTAASYSSPSVMDPYKTLRIQPDASESEVKKAFRQLALQYHPDVCRGSNCGVQFSL
INEAYDTVMSNLREEPDESSQMYMSYEPSEQGIDEPMRGMNDTDWDMWEEWMGWEGAGIRDYSSHVNPYI
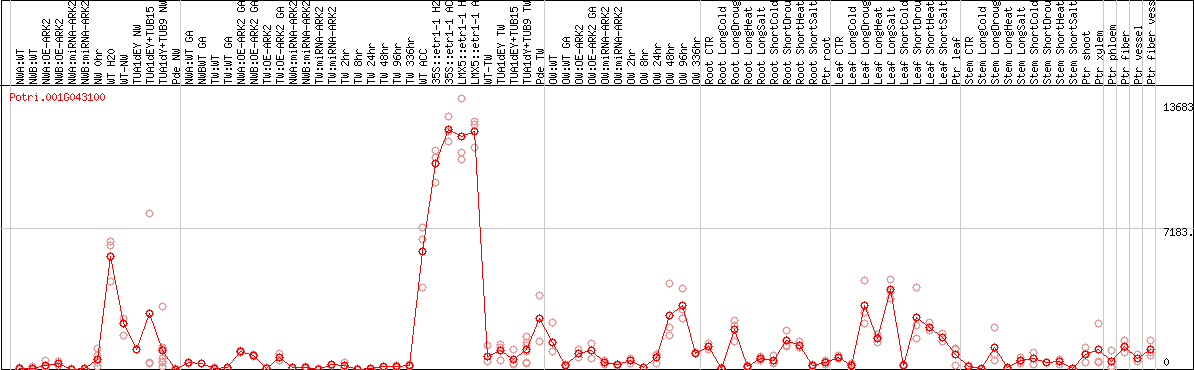
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G043100 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.