Potri.001G053000 [POPLAR]

| External link |
|
|||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||
|
Paralogs
|
|
|||||||||||||||
|
Flax homologues |
|
|||||||||||||||
| PFAM info |
| |||||||||||||||
|
Representative CDS sequence |
>Potri.001G053000.4 pacid=42791191 polypeptide=Potri.001G053000.4.p locus=Potri.001G053000 ID=Potri.001G053000.4.v4.1 annot-version=v4.1
ATGGAGAGAAAAAGCTCCAGCAAGGAGCTGCAGAATCTCCTTCAAGCCATCAAATCTTCAGATGTTGTTGAGAGCCGGATCGAACTGTTTGATAAGCTTC
GGGATTTTGATTTACTGGAGAAATCTGATTTGGCTCCTGTTATCGAATGTCTCACAGTATTTTGGGAAGACTTCACTTGCTTGGATATATCTCAGTGCTT
GTTAAACAAGTCTATTTTGAGTGTGGCTACGAAATATGTAGATTCAGGATTGTTTGGATGTTTAGTACAGTTTCTCGTTCTTGGGACAAAGGCAAGTGGG
TGGTGCGGAAAACATCTGAAGATGACTGCTATGTCAACTGAGGAATCTCAAGAAGAAGAACATTCCAACCTATTCTTTCAGCTTCTTTTAGATTTATTGA
GTCTCTCTTCTGCCAGTACGGTTGCTTTGACAAGACACCCTGTTTTTATTGACAATGCATCAGCAGCTATTGTTGAGAGATTCATTTTGGAACAGCTAAA
TTTGATAAAAGACGTAGTATCAGAGTTTAAGACAATCAGCTCATTTGGCTCAGAAATACTAAAGGCTGCACAGACTGTGATTGATACTGTAATGAGACTA
TGCAAAGGGTATTTTGATGCTGTGAATTGGGATCTTTTTGATTCAAGACCTGAGAAGGATGAAAACAATATAGATTCAGAAAGAGCCAATATAATGAATC
ATGTTACAAACATAACAAAACGTACAACAGAGAAATTGTGTGAACTGGGGATCCTTGCAGGAAATGATGGTGGAAGTCTGGTAACCATTCTCAATGTGTC
ATGGAAAGGTGTGGTTACCTTGCTTCAGCAAGGGAAGAGGGTGTCAAAAGAGATGTTAAGCGTTCAGGATATTATTGTAACCCTAATTTCACTGGTGAAT
GAACCTCTTAGGTGTGCAGCCGAGGCTTGGTCTTCTTCACTGAGGGAAACCATTTCTTTAACTGAAGCCAGAAGGGCATTTCTTCCATCCAAGTTTTACC
TTACCACTGTTGTGAAAATATCTTCCCTCTATCCATGCCAAGCATATTTGGTGTACAAGGAGGTGACACTCTGTGTCCTAATGATCTCAACCTTTAAAGT
TTTACTGAGCTATGAAAAACTTTTGAACACTGCTAGTGAAGTGTTTTCAGAACTTTTGGAGAAGACATCAATGGATTTGCTGAATTCTTTACTCAATTCA
ACTGAAGTGAAACAGGAGCACAAGTTTAAACTTCTGGACTGGTTGTTCAGTGATGAGTCCTGCTCAAATTCTATGCATGAAGGTTCAAGTATTTTTTCTC
GCATGACTTCGATGGTTGAAATCTTTTCTGTGAGCTGTGAAGCTATGTCTGAAGCAAGATTATTGTTGCTTGGCAGGGTTGCATTGTTTCATGATTTGTT
GAGATACTCCATGGTTCTTGAAGAAGACATAAGGATCAAGATCACTGGAAAATTTGGATGGTTTCTGGATATGCTTGTTGATGAAGATGTATACTCTTTC
GTTCTTGATTTGCAGATTCCTGTGCCATATGGTTCAGGGAAAGCACAGGAATTGGTTTGGCAGCCTATGTTTTCTGCTCTTTTGCATGCACTAAAAACCT
TCATGATTGTGGTGTATTCAAGTTGTGCCTGGGAAGAATTAGAGGCTTTCCTGCTTGAGAATCTTTTCCATCCACATTTTCTTTGCCGGGAGATTGTAAT
GGAACTTTGGTGCTTTTTGGTGCGTTATGCTGAAATGGACATGGTGAATAGCATTATTGATAAACTTTGCTCATTACTGAAGCTCCTGGAATCTCCAGAG
TCATTTCTTGTTCCAGGTTCTCCTCTGAGGAAAGTAGCAAGAATAATTTGCTTGCTTGCTAATGGTACAACACCTATGGCAGACCGTGTTTACAGCTCTG
TAGTTGGTGATGGAAGATCTCAATTGTCATCAGTTATGTACGTAGCATTGCTCTTGGAAGGGTTTCCTCTAAATTCACTATCCGATAGCATCAGAAGTAC
TGCAAAAGAGAAGATCATTACAGACTATTTTGGTTTCATAGGGAGTTTTGATGATAAAATGTTGACTACATGTAGTTCTGGTGCATTTGGGATCCCAGTT
CATGCTCTATCTGCTTCCTTGCGGGCACAGCAGGTTAGCATATCTGATGTTGACATGAAGACCCTGAAGTTTTTAGTTGCTATAATTCGTAACTTCAGAA
ACCCTGTGGAAAAAATAAGGAAGGAACACTATTATAAGCTTTTGAGTGGAACGTTGGGGATTGTCTCAAATATGAAGCATCTATACAAATCTGATGAGAT
GGAGGGAGTCATATTGGAACTTCAAACCCTGTTTGTTTCAGCGCCAGCAGCATCAAGTACCCAATTGCATCAGTGCAAACCATATCTTGCTCTTTTTATG
GGAGGACTTGGAGACATGGAGATGACAGAGAGTGATGATTGTGCAAAAAGCTCTGCTGTGTGGGAGTTGTACCACATGCTATTCAGGGAGCGGCATTGGG
CACTTGTTCATCTTGCAATTGAAGCATTTGGATATTTTGCTGCTCGTACTTCTTGCAATCAGCTATGGAGATTTGTGCCACAAAATGCATCGCTTTCTTA
TGATCTGATGTCAGGAAATGAAGCAAGCGAGAAGAGGTTTATGTCCGATTTGAAGGCATTTCTGGAGAAGGAAACAGCTCTTCTTAACACTACGCCTAGC
ATGGAGCAGCTTGAGTTGCTTGTGACGGAAGGTATGACACTAAAAGAAATGGTTCAGAAGATCTCAAGGATACATATAGATGCTACGGAGTGTGAAAGTA
TGGAGATTAATGTTGACATTGTGTCAAAAAAGAGACGGAAACTCCCTGATGGAATCAGCAGGGGAATGGAATTACTCCAGAGTGGTCTGAAGTTAATTGG
TGGTGGTATTTCCCAGTGGCAGGAAAATCACTTCGAATCCTCTGAACTTCATGACAAGTTCTTGAGTCACCTTTCTTGCCTTGAAGACGTCGTTTCTCAC
TTAACTGGCTTGGCTGGTAATGATTAA
|
|||||||||||||||
|
AA sequence
|
>Potri.001G053000.4 pacid=42791191 polypeptide=Potri.001G053000.4.p locus=Potri.001G053000 ID=Potri.001G053000.4.v4.1 annot-version=v4.1
MERKSSSKELQNLLQAIKSSDVVESRIELFDKLRDFDLLEKSDLAPVIECLTVFWEDFTCLDISQCLLNKSILSVATKYVDSGLFGCLVQFLVLGTKASG
WCGKHLKMTAMSTEESQEEEHSNLFFQLLLDLLSLSSASTVALTRHPVFIDNASAAIVERFILEQLNLIKDVVSEFKTISSFGSEILKAAQTVIDTVMRL
CKGYFDAVNWDLFDSRPEKDENNIDSERANIMNHVTNITKRTTEKLCELGILAGNDGGSLVTILNVSWKGVVTLLQQGKRVSKEMLSVQDIIVTLISLVN
EPLRCAAEAWSSSLRETISLTEARRAFLPSKFYLTTVVKISSLYPCQAYLVYKEVTLCVLMISTFKVLLSYEKLLNTASEVFSELLEKTSMDLLNSLLNS
TEVKQEHKFKLLDWLFSDESCSNSMHEGSSIFSRMTSMVEIFSVSCEAMSEARLLLLGRVALFHDLLRYSMVLEEDIRIKITGKFGWFLDMLVDEDVYSF
VLDLQIPVPYGSGKAQELVWQPMFSALLHALKTFMIVVYSSCAWEELEAFLLENLFHPHFLCREIVMELWCFLVRYAEMDMVNSIIDKLCSLLKLLESPE
SFLVPGSPLRKVARIICLLANGTTPMADRVYSSVVGDGRSQLSSVMYVALLLEGFPLNSLSDSIRSTAKEKIITDYFGFIGSFDDKMLTTCSSGAFGIPV
HALSASLRAQQVSISDVDMKTLKFLVAIIRNFRNPVEKIRKEHYYKLLSGTLGIVSNMKHLYKSDEMEGVILELQTLFVSAPAASSTQLHQCKPYLALFM
GGLGDMEMTESDDCAKSSAVWELYHMLFRERHWALVHLAIEAFGYFAARTSCNQLWRFVPQNASLSYDLMSGNEASEKRFMSDLKAFLEKETALLNTTPS
MEQLELLVTEGMTLKEMVQKISRIHIDATECESMEINVDIVSKKRRKLPDGISRGMELLQSGLKLIGGGISQWQENHFESSELHDKFLSHLSCLEDVVSH
LTGLAGND
|
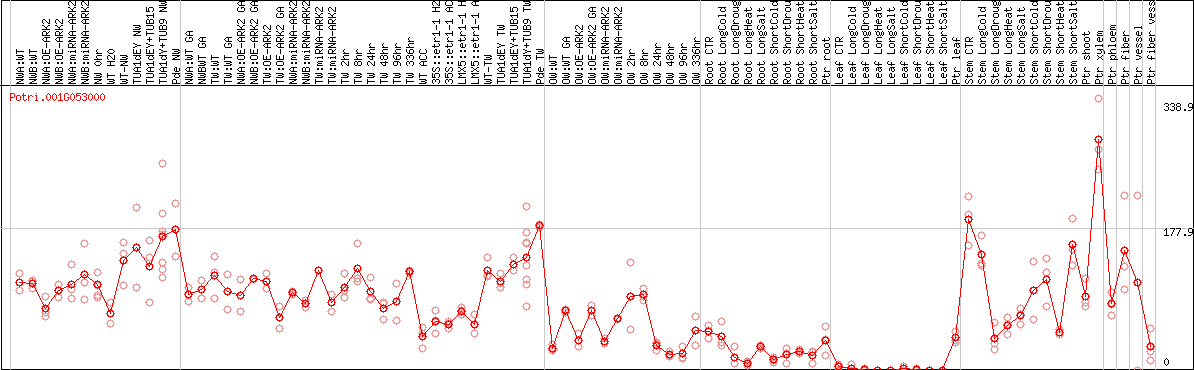
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G053000 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
