External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G13180 309 / 8e-107
NAC
VNDIP2, ANAC083, VNI2
VND-interacting 2, NAC domain containing protein 83 (.1)
AT2G33480 260 / 4e-87
NAC
ANAC041
NAC domain containing protein 41 (.1.2)
AT3G04070 197 / 3e-61
NAC
ANAC047
NAC domain containing protein 47 (.1.2)
AT3G15510 192 / 3e-59
NAC
ATNAC2, ANAC056, NARS1
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
AT1G61110 189 / 7e-59
NAC
ANAC025
NAC domain containing protein 25 (.1)
AT1G77450 186 / 2e-58
NAC
ANAC032
NAC domain containing protein 32 (.1)
AT1G69490 185 / 8e-58
NAC
NAP, ANAC029, ATNAP
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
AT1G01720 179 / 5e-55
NAC
ATAF1, ANAC002
Arabidopsis NAC domain containing protein 2, NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1)
AT4G27410 174 / 2e-53
NAC
RD26, ANAC072
NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.1), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.2), NAC (No Apical Meristem) domain transcriptional regulator superfamily protein (.3)
AT5G61430 172 / 8e-52
NAC
ANAC100, ATNAC5
NAC domain containing protein 100 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10002581
342 / 1e-119
AT5G13180 273 / 6e-93
VND-interacting 2, NAC domain containing protein 83 (.1)
Lus10001809
338 / 4e-118
AT5G13180 271 / 6e-92
VND-interacting 2, NAC domain containing protein 83 (.1)
Lus10018469
198 / 1e-61
AT1G61110 301 / 5e-101
NAC domain containing protein 25 (.1)
Lus10032657
198 / 2e-61
AT3G15510 353 / 1e-120
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
Lus10011215
197 / 2e-61
AT1G61110 305 / 1e-102
NAC domain containing protein 25 (.1)
Lus10043095
197 / 3e-61
AT3G15510 351 / 1e-119
NAC-REGULATED SEED MORPHOLOGY 1, Arabidopsis NAC domain containing protein 56, NAC domain containing protein 2 (.1)
Lus10030446
192 / 2e-60
AT1G69490 302 / 2e-103
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
Lus10026617
189 / 2e-59
AT1G69490 298 / 1e-101
Arabidopsis NAC domain containing protein 29, NAC-like, activated by AP3/PI (.1)
Lus10003269
192 / 3e-59
AT3G04070 324 / 3e-109
NAC domain containing protein 47 (.1.2)
Lus10006547
191 / 1e-58
AT3G04070 337 / 4e-114
NAC domain containing protein 47 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF02365
NAM
No apical meristem (NAM) protein
Representative CDS sequence
>Potri.001G061200.1 pacid=42788658 polypeptide=Potri.001G061200.1.p locus=Potri.001G061200 ID=Potri.001G061200.1.v4.1 annot-version=v4.1
ATGGAGAAGCTTAGTTTTGTTAAGAATGGTGTGCTTAGATTGCCTCCTGGATTTAGGTTCCACCCAACAGATGAGGAGCTTGTTGTCCAGTACTTGAAGA
GAAAGGTGTTTGCTTGCCCCTTGCCTGCTTCCATAATCCCTGAAGTCGATGTTTGCAAGTCTGATCCTTGGGATTTGCCAGGTGATTTGGAGCAAGAACG
GTACTTTTTCAGCACCAGAGAAGCCAAATATCCCAATGGGAATCGATCCAACAGAGCCACAGGCTCTGGCTACTGGAAGGCAACTGGAATAGACAAGCAA
ATTGTGACTTCTAAGGGCCACCAAGTTGTGGGGATGAAGAAAACTCTGGTTTTTTACAGAGGAAAGCCCCCCCATGGCACTAGGACTGATTGGATCATGC
ATGAATACCGCCTTGCAAGCACTGAAACCACAGCCTGCAATACCCTGAAAAATAAAAATTCAACTCAGGGCCCTGTTGTGGTGCCAATGGAAAATTGGGT
TCTATGCCGCATATTTTTGAAGAAGAGAGGCACAAAAAATGAGGAGGAAAACATTCAAGTTGGCAATGATAATAGACTGCCCAAACTCAGGGCCACTGAG
CCTGTTTTCTATGATTTCATGACAAAGGAGAAGACAACTGATTTGAATCTAGCTCCTTCCTCTTCATCCTCAGGATCCAGTGGAATCACAGAGGAGGTGT
CCTGTAATGAATCAGATGATCACGAAGAGAGTAGTAGTTGCAATAGTTTTCCTTACGTTAGAAGAAAACCATAG
AA sequence
>Potri.001G061200.1 pacid=42788658 polypeptide=Potri.001G061200.1.p locus=Potri.001G061200 ID=Potri.001G061200.1.v4.1 annot-version=v4.1
MEKLSFVKNGVLRLPPGFRFHPTDEELVVQYLKRKVFACPLPASIIPEVDVCKSDPWDLPGDLEQERYFFSTREAKYPNGNRSNRATGSGYWKATGIDKQ
IVTSKGHQVVGMKKTLVFYRGKPPHGTRTDWIMHEYRLASTETTACNTLKNKNSTQGPVVVPMENWVLCRIFLKKRGTKNEEENIQVGNDNRLPKLRATE
PVFYDFMTKEKTTDLNLAPSSSSSGSSGITEEVSCNESDDHEESSSCNSFPYVRRKP
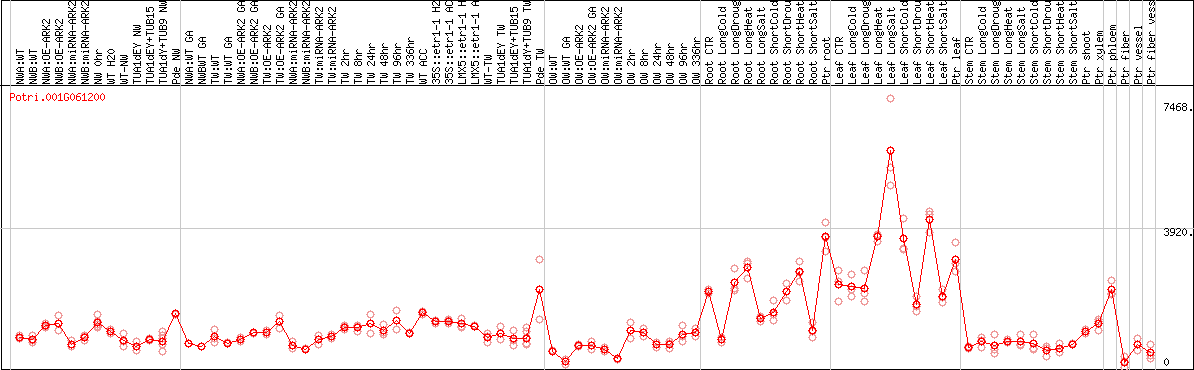
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G061200 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.