Potri.001G074000 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G074000.3 pacid=42791840 polypeptide=Potri.001G074000.3.p locus=Potri.001G074000 ID=Potri.001G074000.3.v4.1 annot-version=v4.1
ATGGATGCATTCGGCCGCTACTTCGTCGACGAGAAAGCTGTCCGTGTTGAGAACATTTTCCTCGACTTCCTCAAAAGTTTTAGATTAGATGGACAAAATC
GTAATATAGGAGAACCGTACTACGATGCTGAGATTGAAGCAATGAAAGCGAATGAATCAACAACGATGTTCATTGATTTCTCTCATGTCATGCTTTTTAA
TGATGTTCTTCAAAAAGCCATTGCCGACGAGTACTTTAGGTTTGAGCCGTATTTGAAGAATGCGTGTAAGAGGTTTGTGATGGAGCTGAGCTCAACGTTT
ATATCTGATGATAATCCTAACAAGGACATCAATGTTGCTTTCTTTAATATCCCTTTTTCCATGAGATTGAGAGAGCTAACAACAGCGGAAATTGGGAAAC
TTGTGTCAGTAACCGGAGTGGTGACTCGCACAAGTGAGGTAAGGCCGGAGCTTTTGCAAGGAACTTTTAGGTGTTTGGAATGTGGTGGTGTTGTCAAGAA
TGTTGAACAGCAATTCAAATATACTGAGCCAACAATTTGTGCCAATGCGACATGTTCCAATAAAATGAGATGGGCATTGCTTAGGCAAGAGAGCAAATTT
GCTGACTGGCAGAGGGTGAGAATGCAGGAAACCTCTAAAGAGATTCCTGCCGGTTCCCTTCCCAGATCCTTGGATGTTATTGTTCGCCATGATATTGTTG
AAAAGGCCAGAGCTGGTGACACGGTCATTTTCACAGGCACTGTGGTTGTTGTACCTGATATATTGGCATTGGCATCCCCTGGAGAGAGAGCAGAATGCCG
GCGGGAATCTTCTCAGCTTAAAAATTCTGCTGTTGGTGGTGAAGGTGTCAGAGGGCTCCGAGCATTGGGAGTGAGAGACCTCTCTTATCGCCTTGCATTT
ATAGCCAATTCAGTGCAGGTTTGTGATGGCAGAAGGGATACTGACATCAGGAATAGAAAAAAGGCTGTTGATGAAGATGATAACCAGGAATTCACGACAG
AGGAGCTAGATGAAATCCAAAGAATGAGAAACACTCCTGATTTCTTCAATAAGATTGTTGATAGCATTGCTCCAACTGTTTTTGGTCACCAAGATATCAA
ACGTGCAATCCTGCTCATGCTCTTGGGTGGCGTTCACAAGTTCACTCATGAGGGTATTAACCTTAGGGGGGACATCAATGTTTGTATTGTTGGTGATCCC
AGCTGTGCAAAGTCTCAGTTCCTCAAGTATGCTTCAGGTATAGTTCCAAGATCTGTCTACACATCTGGAAAATCATCATCTGCTGCTGGGCTGACAGCAT
CTGTGGCCAAAGAACCAGAAACTGGGGAATTTTGTATTGAGGCTGGTGCATTGATGCTTGCTGACAATGGCATTTGTTGCATTGATGAATTCGACAAGAT
GGATATTAGAGATCAGGTTGCAATTCATGAGGCCATGGAACAGCAGACAATAAGCATCACAAAAGCTGGGATACAAGCGACACTCAATGCTCGAACATCA
ATTCTAGCTGCAGCTAACCCCGCTGGAGGACGCTATGATAAATCTAAACCGCTCAAGTATAATGTGGCTCTCCCTCCTGCAATTCTTTCAAGGTTTGATC
TGGTATATGTGATGATTGATGATCCAGATGATCAAACTGATTACCACATTGCGCACCACATCGTAAGAGTTCATCAAAAGCGTGAAGAAGCTCTTTCTCC
TGCATTCACAACTGCTCAGATAAAACGATACATTACATATGCGAAAACCTTGAAACCAAAGCTTAATTCCGAAGCAAGAAAACTGTTGGTGGACTCTTAT
GTTGCTCTTCGCAAAGGTGATACAACTCCAGGGAGTAGAGTTGCTTATCGAATGACTGTTAGACAGCTGGAGGCACTGATCAGACTGTCTGAGGCCATTG
CTCGAAGTCATTTGGAAACTCAGGTACAACCACGACATGTTCGTGTAGCAGTCAAATTGCTGAAAACATCCATAATCAGTGTGGAGTCAAGCGAGATTGA
TCTATCTGAGTTTCAAGAAGCATATGGTGATGGTGGTGATGGTGGAAATGATGGTCCCAGTCAAGGTGATGCCCAGCCAAGCAATGCTGACGCTAACCCA
GTTTCAGAAAACACAGAAAATGGAGCGGCTTCTGCTAGCCGACAGGGGAAAAAACTTGTAATAAGTGAAGAATACTTTCAGAGGGTTACACAAGCACTGG
TCATGCGCCTTAGACAGCATGAAGAAGCTGTGATGCGAGATGGGACGGGGCTGGCTGGAATGAGGCAGGGAGAGTTGATAAGATGGTATGTGGATCAACA
AAATCAGAAAAATAGCTACAGTTCCTTGGAGGAAGCAAAGAATGAAGCTTCCAAAATCAAAGCCATTATAGAGAGTTTGATTCGGAGAGAAGGCTTTTTG
ATTGTGGTAGATGATGGCAGCCGACCAGAAGCAGAGGGTGATGGTGCTAGGCAGTCATCATCAAGAGATGACAGGATATTGGTTGTGGCTCCTAATTATT
TGGTAGAATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G074000.3 pacid=42791840 polypeptide=Potri.001G074000.3.p locus=Potri.001G074000 ID=Potri.001G074000.3.v4.1 annot-version=v4.1
MDAFGRYFVDEKAVRVENIFLDFLKSFRLDGQNRNIGEPYYDAEIEAMKANESTTMFIDFSHVMLFNDVLQKAIADEYFRFEPYLKNACKRFVMELSSTF
ISDDNPNKDINVAFFNIPFSMRLRELTTAEIGKLVSVTGVVTRTSEVRPELLQGTFRCLECGGVVKNVEQQFKYTEPTICANATCSNKMRWALLRQESKF
ADWQRVRMQETSKEIPAGSLPRSLDVIVRHDIVEKARAGDTVIFTGTVVVVPDILALASPGERAECRRESSQLKNSAVGGEGVRGLRALGVRDLSYRLAF
IANSVQVCDGRRDTDIRNRKKAVDEDDNQEFTTEELDEIQRMRNTPDFFNKIVDSIAPTVFGHQDIKRAILLMLLGGVHKFTHEGINLRGDINVCIVGDP
SCAKSQFLKYASGIVPRSVYTSGKSSSAAGLTASVAKEPETGEFCIEAGALMLADNGICCIDEFDKMDIRDQVAIHEAMEQQTISITKAGIQATLNARTS
ILAAANPAGGRYDKSKPLKYNVALPPAILSRFDLVYVMIDDPDDQTDYHIAHHIVRVHQKREEALSPAFTTAQIKRYITYAKTLKPKLNSEARKLLVDSY
VALRKGDTTPGSRVAYRMTVRQLEALIRLSEAIARSHLETQVQPRHVRVAVKLLKTSIISVESSEIDLSEFQEAYGDGGDGGNDGPSQGDAQPSNADANP
VSENTENGAASASRQGKKLVISEEYFQRVTQALVMRLRQHEEAVMRDGTGLAGMRQGELIRWYVDQQNQKNSYSSLEEAKNEASKIKAIIESLIRREGFL
IVVDDGSRPEAEGDGARQSSSRDDRILVVAPNYLVE
|
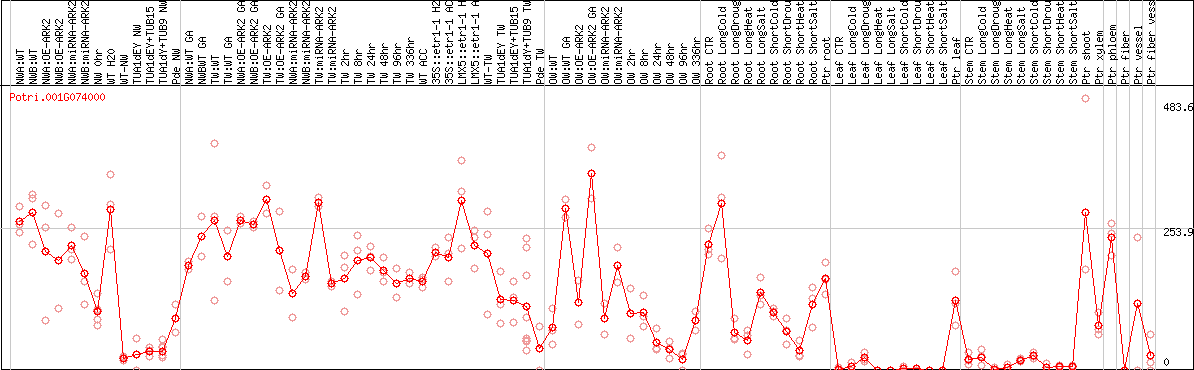
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G074000 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
