External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G02050 63 / 2e-13
ATKT4, ATKUP3, KUP3
K+ uptake transporter 3, ARABIDOPSIS THALIANA POTASSIUM TRANSPORTER 4, K+ uptake transporter 3 (.1)
AT4G13420 61 / 9e-13
HAK5, ATHAK5
high affinity K+ transporter 5, ARABIDOPSIS THALIANA HIGH AFFINITY K+ TRANSPORTER 5, high affinity K+ transporter 5 (.1)
AT4G23640 57 / 1e-11
ATKT3, KUP4, TRH1
TINY ROOT HAIR 1, Potassium transporter family protein (.1)
AT2G30070 56 / 4e-11
ATKUP1, ATKT1P, ATKT1
POTASSIUM UPTAKE TRANSPORTER 1, potassium transporter 1 (.1)
AT2G40540 54 / 3e-10
ATKUP2, ATKT2, TRK2, SHY3, KT2
potassium transporter 2 (.1.2)
AT1G70300 48 / 3e-08
KUP6
K+ uptake permease 6, K+ uptake permease 6 (.1)
AT5G09400 45 / 2e-07
KUP7
K+ uptake permease 7, K+ uptake permease 7 (.1)
AT1G60160 44 / 5e-07
Potassium transporter family protein (.1)
AT4G33530 44 / 6e-07
KUP5
K+ uptake permease 5, K+ uptake permease 5 (.1)
AT2G35060 42 / 2e-06
KUP11
K+ uptake permease 11, K+ uptake permease 11 (.1), K+ uptake permease 11 (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.003G148200
86 / 7e-22
AT2G30070 831 / 0.0
POTASSIUM UPTAKE TRANSPORTER 1, potassium transporter 1 (.1)
Potri.014G144900
65 / 3e-14
AT3G02050 1175 / 0.0
K+ uptake transporter 3, ARABIDOPSIS THALIANA POTASSIUM TRANSPORTER 4, K+ uptake transporter 3 (.1)
Potri.002G237500
65 / 3e-14
AT3G02050 1179 / 0.0
K+ uptake transporter 3, ARABIDOPSIS THALIANA POTASSIUM TRANSPORTER 4, K+ uptake transporter 3 (.1)
Potri.007G068200
63 / 5e-14
AT2G30070 198 / 3e-59
POTASSIUM UPTAKE TRANSPORTER 1, potassium transporter 1 (.1)
Potri.005G095900
62 / 3e-13
AT2G30070 769 / 0.0
POTASSIUM UPTAKE TRANSPORTER 1, potassium transporter 1 (.1)
Potri.003G133900
56 / 4e-11
AT4G23640 947 / 0.0
TINY ROOT HAIR 1, Potassium transporter family protein (.1)
Potri.008G147400
55 / 1e-10
AT1G70300 1221 / 0.0
K+ uptake permease 6, K+ uptake permease 6 (.1)
Potri.013G083400
54 / 3e-10
AT2G40540 1225 / 0.0
potassium transporter 2 (.1.2)
Potri.010G094300
52 / 1e-09
AT1G70300 1228 / 0.0
K+ uptake permease 6, K+ uptake permease 6 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10018270
76 / 6e-18
AT2G40540 652 / 0.0
potassium transporter 2 (.1.2)
Lus10024709
60 / 2e-12
AT3G02050 636 / 0.0
K+ uptake transporter 3, ARABIDOPSIS THALIANA POTASSIUM TRANSPORTER 4, K+ uptake transporter 3 (.1)
Lus10032327
59 / 4e-12
AT2G30070 635 / 0.0
POTASSIUM UPTAKE TRANSPORTER 1, potassium transporter 1 (.1)
Lus10030539
55 / 8e-11
AT2G40540 1245 / 0.0
potassium transporter 2 (.1.2)
Lus10012888
54 / 2e-10
AT5G14880 566 / 0.0
Potassium transporter family protein (.1)
Lus10030857
52 / 2e-09
AT1G70300 1249 / 0.0
K+ uptake permease 6, K+ uptake permease 6 (.1)
Lus10013304
51 / 2e-09
AT4G13420 950 / 0.0
high affinity K+ transporter 5, ARABIDOPSIS THALIANA HIGH AFFINITY K+ TRANSPORTER 5, high affinity K+ transporter 5 (.1)
Lus10030632
50 / 6e-09
AT1G70300 1238 / 0.0
K+ uptake permease 6, K+ uptake permease 6 (.1)
Lus10029050
50 / 7e-09
AT2G40540 563 / 0.0
potassium transporter 2 (.1.2)
Lus10032154
48 / 3e-08
AT2G30070 562 / 0.0
POTASSIUM UPTAKE TRANSPORTER 1, potassium transporter 1 (.1)
PFAM info
Representative CDS sequence
>Potri.001G082267.1 pacid=42789509 polypeptide=Potri.001G082267.1.p locus=Potri.001G082267 ID=Potri.001G082267.1.v4.1 annot-version=v4.1
ATGGCATATATGATGGGAAATACTCAGTGTCTTGGCGATGAAACATCATCACTTGTGAAGAAATCTGTCATTAACATTGCACATGGTTTCTCGAGACAGA
ACTGTCGAAGCCCATCCACAGCACAAGGGGTTCCTCGTACATCTTTGATTGAAGTAGGAATGTCATATTATGTTTAG
AA sequence
>Potri.001G082267.1 pacid=42789509 polypeptide=Potri.001G082267.1.p locus=Potri.001G082267 ID=Potri.001G082267.1.v4.1 annot-version=v4.1
MAYMMGNTQCLGDETSSLVKKSVINIAHGFSRQNCRSPSTAQGVPRTSLIEVGMSYYV
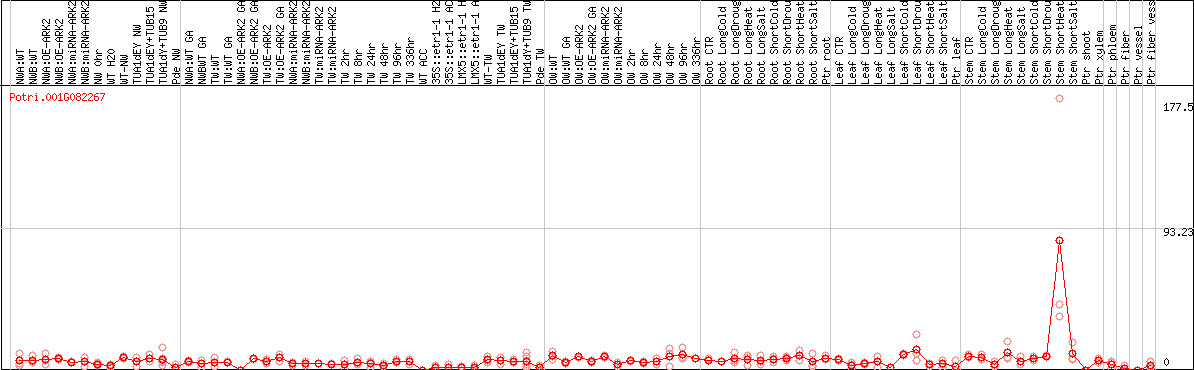
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G082267 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.