Potri.001G082900 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G082900.1 pacid=42788450 polypeptide=Potri.001G082900.1.p locus=Potri.001G082900 ID=Potri.001G082900.1.v4.1 annot-version=v4.1
ATGGGATGGAGTGTAAAGCTGCTCCATTCTTCAGTCTTCGCTGTCTGCTCTTGCTGTCTTCACTTGTTCATCCTTCTTTCAGTCTTTCACTTGTCCAACG
CTCAAAACGCCACTACTGATCCATCTGAAGTGAGTGCATTGAACTTATTATTTGAACAATGGGACACAAAAGCTGTGGGATTATGGAACTTAAGTGGAGA
GCCATGCAGTGGATCTGCCATTAACGGCACCGATTTTGAAGACACTGCCAACAATCCTGCCATCAAATGTGTCTGTACCTACAATAATAGTGCTACATGT
CACATTACTCAACTGAGAGTCTATGCTTTGAACAAAAGAGGGGAGATTCCAGAAGTAATTACGGCTTTGAAGTATCTTACCCTTCTGAAAATTGATCAAA
ATTACTTCACCGGTCCATTGCCTGCATTTATTGGAAATTTAACTGCATTGCAGAGTTTGTCAATTGCGCATAATGCGTTTTCTGGGACCATTCCTACGGA
GCTTGGAAATCTCAAGGAGCTAACTTTGCTGTCTATTGGTATAAATAATTTCTCAGGAACACTCCCTCCAGAACTGGGTCAACTTGTCAATCTTGAGCAA
CTTTATGTGAATAGTTGTGGATTGGGTGGTGAAATTCCCTCAACATTTGTTAACCTTAAAAAAATGACAATCTTTTCGGCTTCAGATGCTGCTTTCACAG
GCAACATACCTGACTTTATTGGCAACTGGACAAGGCTGACATCATTGAGATTTCAAGGGAATTCTTTTGAAGGTCCTATACCATCCAGTTTCTCCAACTT
GACCTCACTGGAATCTCTGCGAATCAGCGATTTAAGCAATGTGAGCTCCACTCTTGATTTTATCAAGAACTTGAAGAGCTTGACCGACTTGACTCTGAGG
AATGCATTGATTAGTGGTAGTATTCCATCTGATATTGGAGAAATATTCCAGACTCTAGATAGACTGGATTTAAGTTTCAACAATTTAACAGGCCAAGTTC
CAAGTGCTTTATTCAATATGAGTTCTCTTCAATACTTGTTTCTTGGAAACAATAGTCTAATCGGAACCCTTCCTAACCAGAAGAGCAGTAAACTTCAGAC
AATAGATTTGTCTTACAATTATCTGTCAGGGACTTTTCCTTCATGGGTAACCTCAAATATACAATTGAATTTAGTGGCCAACAACTTCACTTTTGACAGC
TCAAATATAAGTGTTCTGCCAGGATTGAACTGTCTACAGAGAAATTTCCCATGCAATCGGAATCCTCCACTCTATGCAAATTTCTCAATCAAGTGTGGTG
GTCCAATGATGAGAACTGCTGACGGCACAGTTTATGAAGCTGAAAATTCATCTATCAGTGCAGCATCATTCACTGTAACTAGCACTGAGAAATGGGCGGT
GAGCAACGCAGGCTTGTACGCTGATAGAGAAAATCCTAGTTATGTTGAAAATAACTTGAAACAAGTTACCGGCACCAACACCCCAGAGCTCTATCAGACT
TCAAGGATATCTCCTGGTTCACTTAGGTACTATGGCCTAGGTCTCCAGAATGGGCCTTACACTATAAATCTGTTATTTGCAGAAACTAGATTTGCAGCTC
GAAGCTCACAAACCTGGGATAGTTTAGCACGACGTGTTTTTGATATCTATATTCAGGGAAACCGACAATTGAAGGACTTTGACATATCAATGGAGGCAGG
TGGGGTCGACAGAGCAATTACAAAGACTTTTAACGTTACTGTGTCAGAAAATCATCTTGAAATTCACCTGTTTTGGGCTGGTAAGGGTACGTGCTGTAAC
CCTGTACAAGGCTACTATGGACCAATCATTTCAGCCCTCAATGTTGTTCCAGATTTTACACCAAATGTTAGTGGGATACCATCCAGTACTCGAAAAGAGA
AGAGCAGGACGGGGGTGATTGTTGGTGTTTCCATTTCTGTTGGAGTTGTGAGCTTGATCCTGATATCTGTACTTCTTTACATCAGGTTGAAAAAAGACAG
CGAAGATGAGGAAGTGTTGCTTGGGATGGGCCCTAGACCAAACACTTTTAGTTATTCCCAGTTGAGGACTGCCACCGAAGACTTCAGCCCTTCAAATAAG
CTAGGGGAAGGAGGATATGGACCTGTGTACAAGGGTATGCTTTCTGATGGGAGAGAAGTTGCTGTAAAGAAGCTTTCAGTAGCATCTAACCAGGGAACAA
ATCAATTTGTGACGGAGATCGCAACCATATCGGCAGTTCAACATCGCAATCTTGTGAAATTGTATGGATGCTGCATTGAAGGAAATAGACGCCTCCTCGT
TTATGAATACCTCGAAAACAAAAGCCTTGATAAGACACTCTTTGAAAAAGATGGCATGCATCTTGATTGGCCAACCCGCCTCAATATATGCCTGGGAACT
GCTAGAGGGTTAGCTTATCTCCATGAGGAGTCAAGGCCAAGGATTGTACATAGAGATGTCAAGGCCAGTAACATTTTGCTTGATGCAAACCTCTTCCCCA
AAATATCAGATTTTGGATTGGCAATACTCTATGATGACAAGAAAACTCACATCAGCACTCGAGTTGCAGGAACCATAGGCTATTTGGCACCGGAGTATGC
AATGCGTGGACATCTGACAGAGAAGGCTGATGTTTTTGGTTTTGGTGTGGTTGCTCTGGAGATCCTGAGTGGGAGAGCAAATTCTGATTCTAGTTTGGAT
GACGAAAGGGTCTATCTTCTCGAATGGGCGTGGAAATTGCATGAAAGTGGGCGAAGTTTAGAGCTGATGGATCCGAGTGTAACAGAATTCGATGAAAATG
AAGCTCTTCGAGTTGTTGGAGTAGCTCTCTTGTGCACTCAAGGGTCACCAGCTATGAGGCCAACAATGTCAAGAGTTGTAGCAATGCTTACTGGAGATAT
TGAAGTGAGTGCTGTTACATCAAAGCCAAGCTACTTGACTGATTGGGATTTTAAGGATATAACTGGCACCTTTTCGACTGAAAATACTCAAGCATCCACT
TCATCCGAGGCTAGCAAGAGCAAGAATCACAATCCAATTGATCTTATTCCACGAGGTGATCAAATGCACTCTCCACTAAATGTCACTGAACCAAGGCTCA
GTGACCTCATTGGGGATGGAAGGTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G082900.1 pacid=42788450 polypeptide=Potri.001G082900.1.p locus=Potri.001G082900 ID=Potri.001G082900.1.v4.1 annot-version=v4.1
MGWSVKLLHSSVFAVCSCCLHLFILLSVFHLSNAQNATTDPSEVSALNLLFEQWDTKAVGLWNLSGEPCSGSAINGTDFEDTANNPAIKCVCTYNNSATC
HITQLRVYALNKRGEIPEVITALKYLTLLKIDQNYFTGPLPAFIGNLTALQSLSIAHNAFSGTIPTELGNLKELTLLSIGINNFSGTLPPELGQLVNLEQ
LYVNSCGLGGEIPSTFVNLKKMTIFSASDAAFTGNIPDFIGNWTRLTSLRFQGNSFEGPIPSSFSNLTSLESLRISDLSNVSSTLDFIKNLKSLTDLTLR
NALISGSIPSDIGEIFQTLDRLDLSFNNLTGQVPSALFNMSSLQYLFLGNNSLIGTLPNQKSSKLQTIDLSYNYLSGTFPSWVTSNIQLNLVANNFTFDS
SNISVLPGLNCLQRNFPCNRNPPLYANFSIKCGGPMMRTADGTVYEAENSSISAASFTVTSTEKWAVSNAGLYADRENPSYVENNLKQVTGTNTPELYQT
SRISPGSLRYYGLGLQNGPYTINLLFAETRFAARSSQTWDSLARRVFDIYIQGNRQLKDFDISMEAGGVDRAITKTFNVTVSENHLEIHLFWAGKGTCCN
PVQGYYGPIISALNVVPDFTPNVSGIPSSTRKEKSRTGVIVGVSISVGVVSLILISVLLYIRLKKDSEDEEVLLGMGPRPNTFSYSQLRTATEDFSPSNK
LGEGGYGPVYKGMLSDGREVAVKKLSVASNQGTNQFVTEIATISAVQHRNLVKLYGCCIEGNRRLLVYEYLENKSLDKTLFEKDGMHLDWPTRLNICLGT
ARGLAYLHEESRPRIVHRDVKASNILLDANLFPKISDFGLAILYDDKKTHISTRVAGTIGYLAPEYAMRGHLTEKADVFGFGVVALEILSGRANSDSSLD
DERVYLLEWAWKLHESGRSLELMDPSVTEFDENEALRVVGVALLCTQGSPAMRPTMSRVVAMLTGDIEVSAVTSKPSYLTDWDFKDITGTFSTENTQAST
SSEASKSKNHNPIDLIPRGDQMHSPLNVTEPRLSDLIGDGR
|
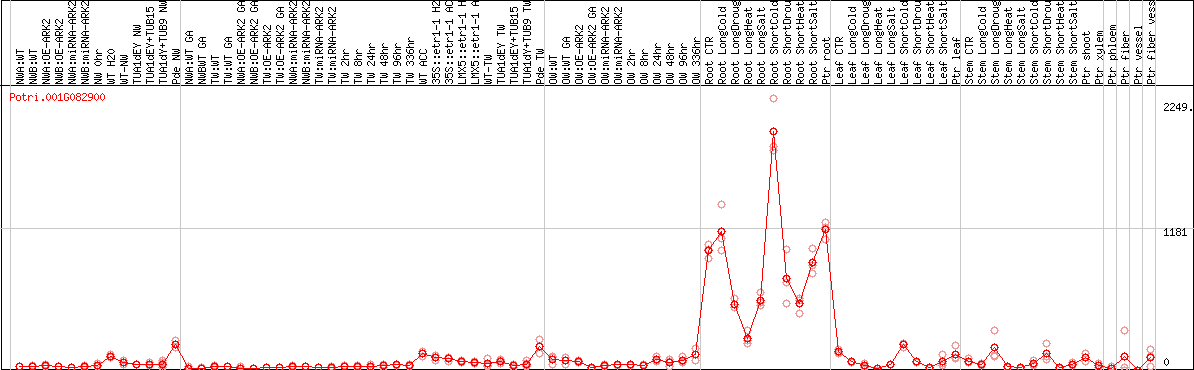
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G082900 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
