Potri.001G089900 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G089900.2 pacid=42788110 polypeptide=Potri.001G089900.2.p locus=Potri.001G089900 ID=Potri.001G089900.2.v4.1 annot-version=v4.1
ATGATAATGGAGGTGGATTTTGTTACAAGTGGAATATTAGAAGAAGAATTGCCTCTTGAAGTTGACGAGAATATTCATGATAATTATAATTTTGAGGGCG
AAAGATGTGGGATATGCATGGATATTGTTATTGACCGGGGCGTTCTCGATTGTTGCCATCACTGGTTTTGTTTTGGTTGCATTGACAATTGGGCCACAAT
CACGAACCTTTGTCCACTCTGTCAGAATGAATTTCAGTCTATTACATGTGTTCCGGTATATGATACCATTGGAAACAACAAAGTTGATGAGGACTCACTT
TCCAGAGATGATGATTGGTCTATTGAAGGGAAGAACAACACTCTCTCCTTTCCATCATACTATATCGATGAAAATGCAGTTATCTGCTTGGATGGAGATG
GCTGCAAGATTCGAAGTGGATCAGCAAATATTGAGGAAGAATCTAATCTCGATACCTCTATTGCTTGCGATTCTTGTGACATTTGGTATCATGCTTTTTG
TGTGGGCTTTGATGCTGAAGGGACATCAGAGGATACATGGTTATGCCCGAGATGCACAGTTGGTGAAGTGCCACAGAAATCTGATGTTGCTTCGCTACAG
AAGCCAAACAACCAGTGTTATTCAGAGAATTCTCACAGTAGTAGTTTTGCTGAGGCTGAGGCTGCTTTTTCTGGAAAGATGTCTGTATCTATCGCTGATG
CTGGAGAAACAGCTGTTGTTGTCTCAATGGTTGGAGGCACTAAATGGACTGAAGAACCAAGTAAACCAACCCTTGAAGTTGATGAGGACCTGATGGATGA
TGCAGTTAAACCTGATGGCAATAGCTACAAGGTAGAAAGACAATCTAGCAAAAAGACTGATGTACAACCAACCGTGGAGGCACCGGAACTTGAACTGTCC
TTATCGTGTGATGCATCCTTCAGTCACCTATCTACTTCCTTGGTGCTTGCTGAGTTGAAGACCATTTGTGATGATGGAACAGTGAATGAACCAATTATTG
GTGATGGTGTTAAAAATTCTTTGAGAAAGTTGTTTAATGATTCCCTTGCCAGAAACAAGCTGTCTGGAAAGGAATCTAGTGAAGGTCTTCATCTTGGTTT
ATCTTTGGGCTGTTCTTCATCTGGTTACATTAAGACCAATGAAACTGAAGATCAAGGGACCATTGAAGTACAACAGCAAAGTCTTTCTGAAGAATCTTTG
CTAAGAAGAGATGAGAAGATTCTACCAGTTGCTAATGAGGAGGCCATGAAAATTATTGGTGTGAAGAGAAAGCATGCCACTTGCAGTGATGATGCTGTTA
AGACTGCTGATGATAACGAAGACAATGCTAAGAATGAAGCTGCCGTTCTTGCAAAGAAGACCAGAGTCAGCAGGAAGTTGCAAATAACTCCTAAGGGCCA
GGACAGTGCATTTCTTCCAGAAGATTCCCAAAAGTGCCCCGCCAAAATAGCAGTTCTAAAAAATGTCAAATTAAAACGATCTCTTGAAAAGCAAGATGTC
ACTTCTGATATAATGAGTGTGGTTAAAGGGACTGGTCGTAGGACTTTGAAGGGGCTTGCTCACCAGAGTCCTCCAGATAAATCATCAAAAGAAGGGGAAA
ATGCAGCGGGCTTGAGAGTGAAAAAGATCATGAGGAGAGCTGTGGAGGACAAGGAATCATCTGTTGTAGTTCAAAACCTTAGAAAAGAAATAAGAGAAGC
TGTGCGTAACCGATCTTCTGATGAAATTGGGGAAAACCTTTTTGACCCAAAGCTTCTTGCTGCCTTCCGAACTGCTGTAGCAGGATCCACAGCGGAACCA
GTGAAGAAGTTGCCCCCTTCTTCTCTGAAGGCAAAGAAGTCCTTGCTGCAGAAGGGTAAAGTTCGTGAGAATCTGACAAAGAAAATATATGGAGATTCTA
ATGGAAGAAGAAAACGTGCATGGGATCGAGACTGTGATGTTGAATTCTGGAAGTATCGCTGTATGAGGGTTACAAAGCCTGAAAAAATTGCAACTTTGAA
GTCAGTTCTTACTCTCCTTAGAAAGAATCCAGAAGGCTCAGAGATGGATCAGGGATATGAATTCCAGGAGACCAATCCCATTCTTTCAAGATTATATTTA
GCAGACACATCTGTTTTTCCAAGAAAGGATGATATCAAGCCTCTCTTAGCTTCTACAACTACAAGTAACACTGAACAGAATAAAGCTCAGGAAATTTCAA
TGGATAAAGTTCGGAAGCTATCTCCTGATGATCATACTCTTAAATCTGCAGGAGCAAATAAAGTATCATCAAAGCTTGTTGTGCCTTCAATTCATGATAA
AGGGCTTAAGGATAAAGTTCTCAGCACAAATTGTCAGCCTGCTTCTAGCAAAGCACAACCGGGTGGATTTTCCAAAGTCAATTCACAGAAGGAAAAAGGT
GCTCAATCTGATGATAAAAGGATGGATAAAAGAAAATGGGCCTTGGAGGTGCTTGCAAGAAAAAAGGCAGTTTCAGGTAAAACTGCAGCAGATGAAAAAC
AGGAAGACAATGCAGTGCTTAAAGGAAATTATCCTTTACTCGCTCAACTGCCAATTGACATGAGACCGGTGTTAGCATCCTGTCGCCACAATAAAATCCC
CATATCGGTTAGACAGACACAACTTTACCGACTGACAGAGCACTTCTTGAGGAAAGTAAATCTGCCAGAGATTCGCAAAACTGCAGAAACAGAGTTGGCT
GTTGCAGATGCAATTAATATTGAGAAGGAGGTTGCCGACAAGGCAAACAGCAAAATAGTATATTTGAATCTTTGTTCTCAGGAAATAATGCGTCACTCAG
ATGACAGAAAGTCCAATAGAGCCACGGTATCAAATTCTTCACCATCTGCAGTCACTGTTGATAGATTAGAACAAGATATCGATGAACTTCCAACGGATCC
TGCAGTTTTAGATGCATTGAGAAATGCAGGACTCTTATCTGATTCTCCTCCAAGTAGTCCCCATCATAAAATGGAGGTTTCCAATGAAGTAGATGATTCC
TCAATGCAGATCAAAGAAGAAGGGCCTGATAATGTATTTGAAATGGATTCTCATCCAGATGTTGATATATATGGTGATTTTGAGTATGATTTAGAAGATG
AAGACTACATTGGTGCTACTAATTTGACAGTCCCCAAGCTAATAGTGGAAGAAGGCGAGTCAAGGATGAAAGTAGTCTTCTCCACTCTTAAATCTGAAAT
GCCGAACAATTTTCAGGATCTTGAAGGTTGTTTGACTTTGGGGAATAATGAAGAACTCAAGGACTCTGCCTCCTCGCCAAAGATTCATGTTGATGCAGGC
ATCATCAGCACAACCATGGAGGGTGGAACTAATAGGTCTTGTGCTGATTCAGAACCTCTACCTGGCGAAGAAGGTGAAGAACCTTCTCTTGCAGAATGTG
ATGAACTATATGGTCCAGACAAGGAGCCTCTGATCAACAAATTTCCAGAAGAAGCATCTAGAAATCTACATGAGCTGACTGACCCTGAGGCTTCAACTAA
GCACAAAGGTTCGGGAGAAAATGAAAATAATTCATCAAGACAAGATGGGAATACCAATGCGACCTCTGCTGGTCACACTTGTGATGGAGAAACCACGTGT
GACCATTCTCAAACAGCTGAAAGTGGCCGTAAAAAGGATAGTTCTAAGACTAACACTAACAAGCAGGGTGACATTATCAATTCTGTTTCTAAGAAGGTTG
AAGCGTACATCAAAGAGCACGTCAGGCCACTGTGTAAGAGTGGCATAATCACTGCTGAACAGTATAGATGGGCTGTGGCAAAAACTACTGATAAGGTTAT
GAAATACCACTTGAATGCGAAAAATGCAAATTTCCTAATCAAGGAAGGTGAGAAGGTGAAGAAACTTGCTGAGCAGTATGTTGAGGCAGCACAGCAAAAG
GAAAGAAGTGATTCTGTATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G089900.2 pacid=42788110 polypeptide=Potri.001G089900.2.p locus=Potri.001G089900 ID=Potri.001G089900.2.v4.1 annot-version=v4.1
MIMEVDFVTSGILEEELPLEVDENIHDNYNFEGERCGICMDIVIDRGVLDCCHHWFCFGCIDNWATITNLCPLCQNEFQSITCVPVYDTIGNNKVDEDSL
SRDDDWSIEGKNNTLSFPSYYIDENAVICLDGDGCKIRSGSANIEEESNLDTSIACDSCDIWYHAFCVGFDAEGTSEDTWLCPRCTVGEVPQKSDVASLQ
KPNNQCYSENSHSSSFAEAEAAFSGKMSVSIADAGETAVVVSMVGGTKWTEEPSKPTLEVDEDLMDDAVKPDGNSYKVERQSSKKTDVQPTVEAPELELS
LSCDASFSHLSTSLVLAELKTICDDGTVNEPIIGDGVKNSLRKLFNDSLARNKLSGKESSEGLHLGLSLGCSSSGYIKTNETEDQGTIEVQQQSLSEESL
LRRDEKILPVANEEAMKIIGVKRKHATCSDDAVKTADDNEDNAKNEAAVLAKKTRVSRKLQITPKGQDSAFLPEDSQKCPAKIAVLKNVKLKRSLEKQDV
TSDIMSVVKGTGRRTLKGLAHQSPPDKSSKEGENAAGLRVKKIMRRAVEDKESSVVVQNLRKEIREAVRNRSSDEIGENLFDPKLLAAFRTAVAGSTAEP
VKKLPPSSLKAKKSLLQKGKVRENLTKKIYGDSNGRRKRAWDRDCDVEFWKYRCMRVTKPEKIATLKSVLTLLRKNPEGSEMDQGYEFQETNPILSRLYL
ADTSVFPRKDDIKPLLASTTTSNTEQNKAQEISMDKVRKLSPDDHTLKSAGANKVSSKLVVPSIHDKGLKDKVLSTNCQPASSKAQPGGFSKVNSQKEKG
AQSDDKRMDKRKWALEVLARKKAVSGKTAADEKQEDNAVLKGNYPLLAQLPIDMRPVLASCRHNKIPISVRQTQLYRLTEHFLRKVNLPEIRKTAETELA
VADAINIEKEVADKANSKIVYLNLCSQEIMRHSDDRKSNRATVSNSSPSAVTVDRLEQDIDELPTDPAVLDALRNAGLLSDSPPSSPHHKMEVSNEVDDS
SMQIKEEGPDNVFEMDSHPDVDIYGDFEYDLEDEDYIGATNLTVPKLIVEEGESRMKVVFSTLKSEMPNNFQDLEGCLTLGNNEELKDSASSPKIHVDAG
IISTTMEGGTNRSCADSEPLPGEEGEEPSLAECDELYGPDKEPLINKFPEEASRNLHELTDPEASTKHKGSGENENNSSRQDGNTNATSAGHTCDGETTC
DHSQTAESGRKKDSSKTNTNKQGDIINSVSKKVEAYIKEHVRPLCKSGIITAEQYRWAVAKTTDKVMKYHLNAKNANFLIKEGEKVKKLAEQYVEAAQQK
ERSDSV
|
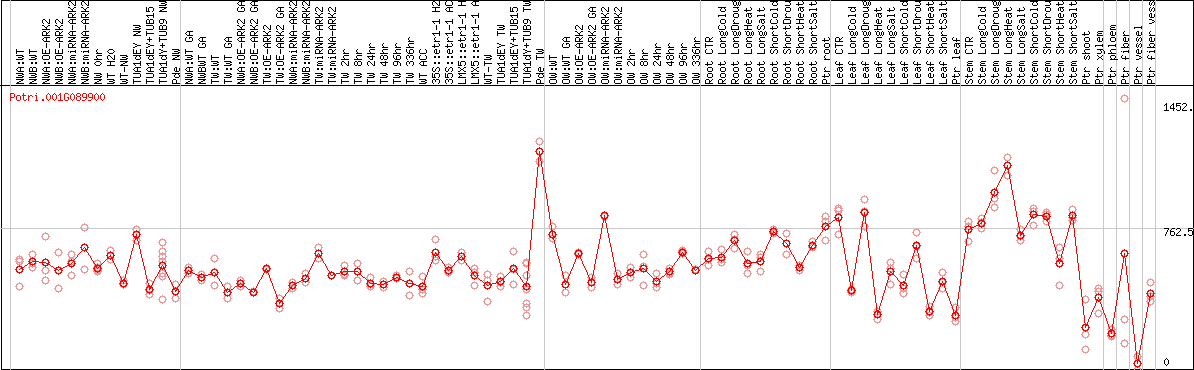
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G089900 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
