External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G64320 65 / 1e-12
myosin heavy chain-related (.1)
AT5G41780 60 / 3e-11
myosin heavy chain-related (.1)
AT5G41790 56 / 8e-10
CIP1
COP1-interactive protein 1 (.1)
AT1G64330 50 / 2e-07
myosin heavy chain-related (.1)
AT5G58320 40 / 0.0003
Kinase interacting (KIP1-like) family protein (.1), Kinase interacting (KIP1-like) family protein (.2), Kinase interacting (KIP1-like) family protein (.3)
AT3G10880 39 / 0.001
unknown protein
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.003G137400
78 / 3e-17
AT5G41790 384 / 6e-109
COP1-interactive protein 1 (.1)
Potri.001G094200
71 / 7e-15
AT5G41790 407 / 9e-121
COP1-interactive protein 1 (.1)
Potri.019G061600
47 / 2e-06
AT5G58320 180 / 8e-50
Kinase interacting (KIP1-like) family protein (.1), Kinase interacting (KIP1-like) family protein (.2), Kinase interacting (KIP1-like) family protein (.3)
Potri.013G158100
43 / 4e-05
AT2G30500 384 / 7e-127
Kinase interacting (KIP1-like) family protein (.1), Kinase interacting (KIP1-like) family protein (.2)
Potri.019G131000
40 / 0.0002
AT2G30500 389 / 4e-129
Kinase interacting (KIP1-like) family protein (.1), Kinase interacting (KIP1-like) family protein (.2)
Potri.003G137300
0 / 1
AT1G64320 237 / 1e-69
myosin heavy chain-related (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10033944
73 / 1e-15
AT1G64320 169 / 6e-46
myosin heavy chain-related (.1)
Lus10032360
70 / 1e-14
AT1G64320 164 / 5e-44
myosin heavy chain-related (.1)
Lus10032362
59 / 1e-10
AT5G41790 275 / 7e-79
COP1-interactive protein 1 (.1)
Lus10033945
58 / 2e-10
AT5G41790 309 / 8e-86
COP1-interactive protein 1 (.1)
Lus10030507
50 / 1e-07
AT5G05180 120 / 3e-30
unknown protein
Lus10012861
49 / 3e-07
AT5G05180 124 / 2e-31
unknown protein
Lus10034251
44 / 2e-05
AT5G05180 149 / 4e-40
unknown protein
Lus10029012
41 / 0.0002
AT5G05180 146 / 5e-39
unknown protein
Lus10034448
40 / 0.0004
AT5G58320 350 / 2e-114
Kinase interacting (KIP1-like) family protein (.1), Kinase interacting (KIP1-like) family protein (.2), Kinase interacting (KIP1-like) family protein (.3)
Lus10019111
39 / 0.0008
AT5G58320 347 / 3e-113
Kinase interacting (KIP1-like) family protein (.1), Kinase interacting (KIP1-like) family protein (.2), Kinase interacting (KIP1-like) family protein (.3)
PFAM info
Representative CDS sequence
>Potri.001G094266.1 pacid=42789134 polypeptide=Potri.001G094266.1.p locus=Potri.001G094266 ID=Potri.001G094266.1.v4.1 annot-version=v4.1
ATGCATTTTCAGGAGCAGAAGCAGTGCTCAAGAAACCCAACGAGGACAATGAAAATCTCTCGGACCCCATTAGCCGAATCTCAAACGAGCTTCAGTTTGC
GAAGAACTGTAACAAAAGATGACATAAAAAAGCTGAAACATAATGTGGATGGTTTGAAGCTCCAGCTCAAGGGCAAGGAAGAGAAGGAGCTTCTGTTAGG
AGAAGAGGTGGGGAAACTAAAAGCAAAGCTGTGCAAGGAAGGGGGAGACAAGCTGAATTCTGTTAGCCAGCTTGAGATAAAGGTGGTATATTTAGAGCAG
CAGGTAAAAGATAAGGAAGAGGTTTTGCTAGGTCTCAGTGAGGAAAGGAGGGAGGCCATAAGGCAGCTATGCATCTTGATTGATTATCATCGCGGCCGCT
ATGATCATCTCAGAGAAGCAATATCAAAGAAGACTGTTCATATCAAGAGAATGGCTTAG
AA sequence
>Potri.001G094266.1 pacid=42789134 polypeptide=Potri.001G094266.1.p locus=Potri.001G094266 ID=Potri.001G094266.1.v4.1 annot-version=v4.1
MHFQEQKQCSRNPTRTMKISRTPLAESQTSFSLRRTVTKDDIKKLKHNVDGLKLQLKGKEEKELLLGEEVGKLKAKLCKEGGDKLNSVSQLEIKVVYLEQ
QVKDKEEVLLGLSEERREAIRQLCILIDYHRGRYDHLREAISKKTVHIKRMA
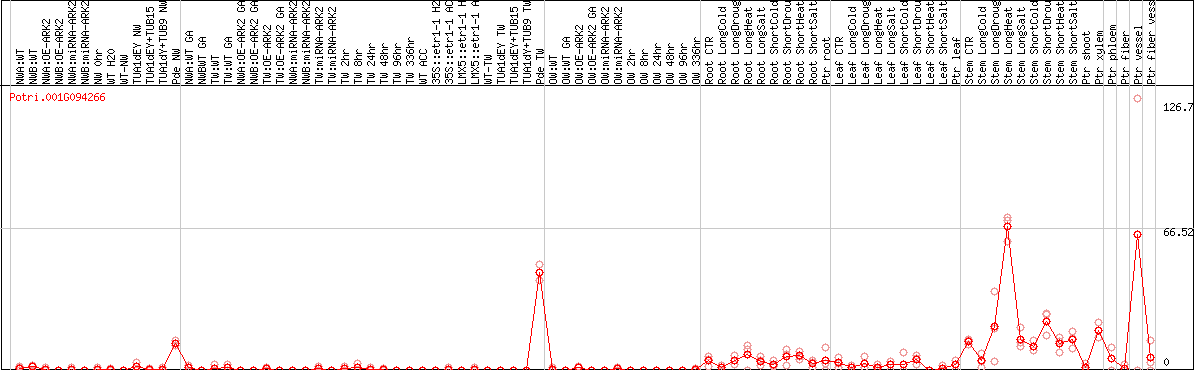
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G094266 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.