External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G35785 97 / 1e-25
RNA-binding (RRM/RBD/RNP motifs) family protein
AT1G07350 89 / 5e-23
SR45a
serine/arginine rich-like protein 45a, RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
AT2G21440 59 / 1e-10
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT3G46020 53 / 1e-09
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT3G53460 53 / 1e-08
CP29
chloroplast RNA-binding protein 29 (.1.2.3.4)
AT2G37510 49 / 6e-08
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT3G10400 50 / 1e-07
U11/U12-31K
U11/U12-31K, RNA recognition motif and CCHC-type zinc finger domains containing protein (.1)
AT1G60650 50 / 1e-07
AtRZ-1b
AtRZ-1b, RNA-binding (RRM/RBD/RNP motifs) family protein with retrovirus zinc finger-like domain (.1), RNA-binding (RRM/RBD/RNP motifs) family protein with retrovirus zinc finger-like domain (.2)
AT2G37220 50 / 1e-07
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
AT3G52380 49 / 2e-07
CP33, PDE322
PIGMENT DEFECTIVE 322, chloroplast RNA-binding protein 33 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.003G130301
128 / 1e-37
AT4G35785 145 / 3e-43
RNA-binding (RRM/RBD/RNP motifs) family protein
Potri.009G041700
100 / 1e-25
AT4G35785 139 / 6e-39
RNA-binding (RRM/RBD/RNP motifs) family protein
Potri.001G248100
98 / 6e-25
AT1G07350 125 / 2e-35
serine/arginine rich-like protein 45a, RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Potri.005G105700
96 / 7e-25
AT4G35785 158 / 4e-48
RNA-binding (RRM/RBD/RNP motifs) family protein
Potri.012G038200
56 / 6e-10
AT1G73530 141 / 6e-43
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.011G130300
52 / 6e-09
AT5G54580 161 / 2e-51
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.001G409800
51 / 2e-08
AT5G54580 174 / 2e-56
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Potri.016G010200
52 / 3e-08
AT3G07810 413 / 2e-140
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Potri.006G208500
50 / 3e-08
AT5G06210 160 / 1e-51
RNA binding (RRM/RBD/RNP motifs) family protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10017507
112 / 3e-31
AT4G35785 123 / 4e-35
RNA-binding (RRM/RBD/RNP motifs) family protein
Lus10028776
101 / 5e-27
AT4G35785 127 / 5e-36
RNA-binding (RRM/RBD/RNP motifs) family protein
Lus10041854
100 / 1e-26
AT4G35785 179 / 6e-56
RNA-binding (RRM/RBD/RNP motifs) family protein
Lus10028400
99 / 4e-26
AT4G35785 181 / 2e-57
RNA-binding (RRM/RBD/RNP motifs) family protein
Lus10040705
98 / 1e-24
AT1G07350 163 / 2e-46
serine/arginine rich-like protein 45a, RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Lus10018202
95 / 4e-23
AT1G07350 145 / 1e-41
serine/arginine rich-like protein 45a, RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Lus10040854
65 / 3e-14
AT3G46020 112 / 1e-33
RNA-binding (RRM/RBD/RNP motifs) family protein (.1)
Lus10005891
64 / 2e-12
AT3G45980 230 / 1e-76
HISTONE H2B, Histone superfamily protein (.1)
Lus10021035
57 / 1e-09
AT3G15010 182 / 4e-51
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
Lus10023817
56 / 2e-09
AT3G15010 188 / 5e-56
RNA-binding (RRM/RBD/RNP motifs) family protein (.1), RNA-binding (RRM/RBD/RNP motifs) family protein (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0221
RRM
PF00076
RRM_1
RNA recognition motif. (a.k.a. RRM, RBD, or RNP domain)
Representative CDS sequence
>Potri.001G101500.2 pacid=42788262 polypeptide=Potri.001G101500.2.p locus=Potri.001G101500 ID=Potri.001G101500.2.v4.1 annot-version=v4.1
ATGACCATCCTTAGGCCTGATTCGCCACGCAATTCAGGTTCACCAGGCAGAGGCCGGCGGTCAAGATCGTTGTCAAGGCCCCGTCGAAGTCGCTCCAGGA
GTATCAATGGTTCTGGTATAACCGGATTATCAACTAGGATCACAAGCAGTGATCTTGAGAAGTATTTTAACAGTGAAGGGAAGGTTTTGGAGTGTCATCT
GGTAACAGATCCTGACACCAGAGAATTTCGTGGATTTGCTTTTGTTACCGTGGAAACCACTGAGGATGCTGACCGTTGCATCAAGTATTTGAATTGTTCA
GTGCTTGAAGGACGACTGATCACCATGGAAAAGATGGCAGACCCAAACAAATGCCTTTCACGCTGTATAAACTGGCGGAATTTCGGAGGGGGAATGAATG
ACCCACCATGGAAGAAGCATTCATTTATGAAAATGAACCTTGTATCAATCTTGGCTCATATCCAATTGAATGCAGCCTGGCTGTAA
AA sequence
>Potri.001G101500.2 pacid=42788262 polypeptide=Potri.001G101500.2.p locus=Potri.001G101500 ID=Potri.001G101500.2.v4.1 annot-version=v4.1
MTILRPDSPRNSGSPGRGRRSRSLSRPRRSRSRSINGSGITGLSTRITSSDLEKYFNSEGKVLECHLVTDPDTREFRGFAFVTVETTEDADRCIKYLNCS
VLEGRLITMEKMADPNKCLSRCINWRNFGGGMNDPPWKKHSFMKMNLVSILAHIQLNAAWL
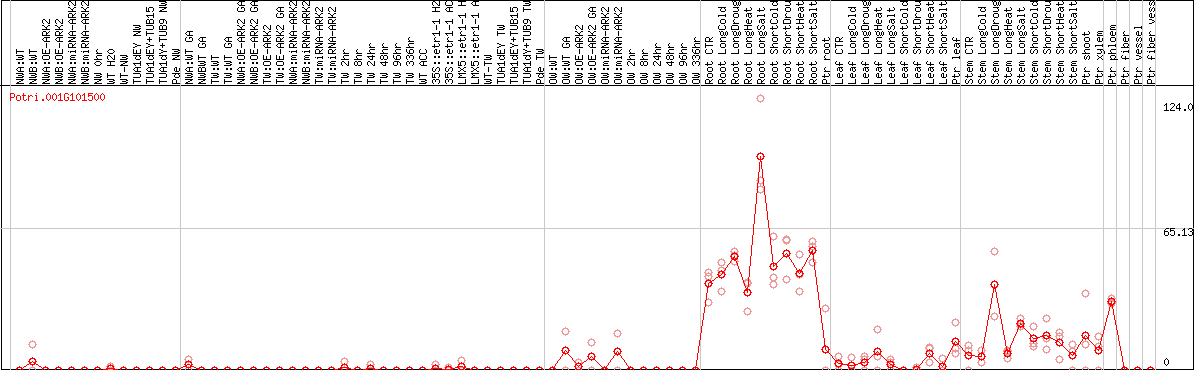
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G101500 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.