External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G63000 535 / 0
UER1, NRS/ER
"UDP-4-KETO-6-DEOXY-D-GLUCOSE-3,5-EPIMERASE-4-REDUCTASE 1", nucleotide-rhamnose synthase/epimerase-reductase (.1)
AT1G78570 503 / 2e-176
ATRHM1, RHM1, ROL1
REPRESSOR OF LRX1 1, ARABIDOPSIS THALIANA RHAMNOSE BIOSYNTHESIS 1, rhamnose biosynthesis 1 (.1)
AT3G14790 457 / 2e-158
ATRHM3, RHM3
ARABIDOPSIS THALIANA RHAMNOSE BIOSYNTHESIS 3, rhamnose biosynthesis 3 (.1)
AT1G53500 455 / 1e-157
ATRHM2, ATMUM4, RHM2, MUM4
ARABIDOPSIS THALIANA RHAMNOSE BIOSYNTHESIS 2, ARABIDOPSIS THALIANA MUCILAGE-MODIFIED 4, NAD-dependent epimerase/dehydratase family protein (.1)
AT2G28760 49 / 2e-06
UXS6
UDP-XYL synthase 6 (.1.2.3)
AT5G59290 49 / 3e-06
ATUXS3, UXS3
UDP-glucuronic acid decarboxylase 3 (.1.2)
AT3G46440 45 / 3e-05
UXS5
UDP-XYL synthase 5 (.1.2)
AT3G62830 44 / 6e-05
ATUXS2, UXS2, AUD1
UDP-GLUCURONIC ACID DECARBOXYLASE 2, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
AT2G47650 44 / 6e-05
UXS4
UDP-xylose synthase 4 (.1.2)
AT3G53520 43 / 0.0001
ATUXS1, UXS1
UDP-glucuronic acid decarboxylase 1 (.1.2.3.4)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.003G120000
605 / 0
AT1G63000 532 / 0.0
"UDP-4-KETO-6-DEOXY-D-GLUCOSE-3,5-EPIMERASE-4-REDUCTASE 1", nucleotide-rhamnose synthase/epimerase-reductase (.1)
Potri.011G103700
506 / 2e-177
AT1G78570 1211 / 0.0
REPRESSOR OF LRX1 1, ARABIDOPSIS THALIANA RHAMNOSE BIOSYNTHESIS 1, rhamnose biosynthesis 1 (.1)
Potri.001G383500
501 / 9e-176
AT1G78570 1190 / 0.0
REPRESSOR OF LRX1 1, ARABIDOPSIS THALIANA RHAMNOSE BIOSYNTHESIS 1, rhamnose biosynthesis 1 (.1)
Potri.006G272700
495 / 2e-173
AT1G78570 1078 / 0.0
REPRESSOR OF LRX1 1, ARABIDOPSIS THALIANA RHAMNOSE BIOSYNTHESIS 1, rhamnose biosynthesis 1 (.1)
Potri.008G053100
48 / 5e-06
AT2G28760 623 / 0.0
UDP-XYL synthase 6 (.1.2.3)
Potri.001G237200
47 / 1e-05
AT3G46440 637 / 0.0
UDP-XYL synthase 5 (.1.2)
Potri.010G207200
47 / 1e-05
AT2G28760 625 / 0.0
UDP-XYL synthase 6 (.1.2.3)
Potri.014G129200
45 / 6e-05
AT3G62830 720 / 0.0
UDP-GLUCURONIC ACID DECARBOXYLASE 2, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Potri.002G204400
44 / 6e-05
AT3G62830 723 / 0.0
UDP-GLUCURONIC ACID DECARBOXYLASE 2, NAD(P)-binding Rossmann-fold superfamily protein (.1), NAD(P)-binding Rossmann-fold superfamily protein (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10006719
549 / 0
AT1G63000 531 / 0.0
"UDP-4-KETO-6-DEOXY-D-GLUCOSE-3,5-EPIMERASE-4-REDUCTASE 1", nucleotide-rhamnose synthase/epimerase-reductase (.1)
Lus10014147
548 / 0
AT1G63000 530 / 0.0
"UDP-4-KETO-6-DEOXY-D-GLUCOSE-3,5-EPIMERASE-4-REDUCTASE 1", nucleotide-rhamnose synthase/epimerase-reductase (.1)
Lus10005560
501 / 5e-176
AT1G78570 1123 / 0.0
REPRESSOR OF LRX1 1, ARABIDOPSIS THALIANA RHAMNOSE BIOSYNTHESIS 1, rhamnose biosynthesis 1 (.1)
Lus10042497
501 / 2e-175
AT1G78570 1227 / 0.0
REPRESSOR OF LRX1 1, ARABIDOPSIS THALIANA RHAMNOSE BIOSYNTHESIS 1, rhamnose biosynthesis 1 (.1)
Lus10013695
499 / 2e-175
AT1G78570 1005 / 0.0
REPRESSOR OF LRX1 1, ARABIDOPSIS THALIANA RHAMNOSE BIOSYNTHESIS 1, rhamnose biosynthesis 1 (.1)
Lus10038146
499 / 5e-175
AT1G78570 1228 / 0.0
REPRESSOR OF LRX1 1, ARABIDOPSIS THALIANA RHAMNOSE BIOSYNTHESIS 1, rhamnose biosynthesis 1 (.1)
Lus10010942
475 / 1e-165
AT1G78570 1130 / 0.0
REPRESSOR OF LRX1 1, ARABIDOPSIS THALIANA RHAMNOSE BIOSYNTHESIS 1, rhamnose biosynthesis 1 (.1)
Lus10007355
324 / 8e-107
AT1G78570 982 / 0.0
REPRESSOR OF LRX1 1, ARABIDOPSIS THALIANA RHAMNOSE BIOSYNTHESIS 1, rhamnose biosynthesis 1 (.1)
Lus10020776
313 / 1e-102
AT1G78570 964 / 0.0
REPRESSOR OF LRX1 1, ARABIDOPSIS THALIANA RHAMNOSE BIOSYNTHESIS 1, rhamnose biosynthesis 1 (.1)
Lus10031389
72 / 9e-16
AT1G78570 84 / 2e-20
REPRESSOR OF LRX1 1, ARABIDOPSIS THALIANA RHAMNOSE BIOSYNTHESIS 1, rhamnose biosynthesis 1 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0063
NADP_Rossmann
PF04321
RmlD_sub_bind
RmlD substrate binding domain
Representative CDS sequence
>Potri.001G112000.1 pacid=42790646 polypeptide=Potri.001G112000.1.p locus=Potri.001G112000 ID=Potri.001G112000.1.v4.1 annot-version=v4.1
ATGGGATTCGAATCTTCGAACGGCACAGCATCACCAAAACTGCTCAAATTCCTCATCTACGGACGTACTGGCTGGATCGGTGGACTCTTAGGCAAGCTTT
GCCAATCCCAGGGCATCGACTTCACCTATGGTTCCGGTCGCCTTGAAAACCGACCCTCCCTCGAAGCCGACCTGGTTGCCGTTAATCCCACTCATGTTTT
CAACGCTGCCGGCGTTACTGGCAGGCCTAACGTCGATTGGTGCGAATCCCATAAAGTCGAGACCATACGAACCAATGTGGTCGGCACACTGACGTTGGCT
GACCTTTGCAGGGAGAAAGGTTTGGTCTTGATTAATTATGCTACGGGCTGTATCTTTGAGTATGATTCGAGCCATCCTCTCGGGTCGGGTATTGGGTTCA
AGGAGGAGGATACTCCGAACTTTATCGGTTCTTTCTATTCCAAGACTAAGGCCATGGTAGAGGATTTGCTCAGGAATTATGAAAATGTTTGCACATTGCG
TGTCAGAATGCCCATTTCATCTGATTTAGCCAACCCTCGCAATTTCATCACAAAGATCACTCGATATGAGAAGGTAGTGGACATTCCAAACTCGATGACA
ATCTTGGATGAACTCCTTCCCATCTCCATCGAGATGGCAAAGAGAAACCTTACTGGGATCTATAACTTTACAAATCCTGGAGTGGTTAGTCACAATGAGA
TTTTAGAAATGTATCGGGATTACATTGACCCCGATTTCACATGGAAAAACTTCACTCTTGAGGAGCAAGCAAAGGTAATAGTTGCCCCAAGGAGCAACAA
TGAGCTTGATACTGCCAAATTGAAGCAGGAATTCCCTGAGCTCTTGCCAATAAAAGAGTCCCTTATCAAGTATGTCTTCAAGCCAAATCAGAAGACAGCT
GCTGCTTGA
AA sequence
>Potri.001G112000.1 pacid=42790646 polypeptide=Potri.001G112000.1.p locus=Potri.001G112000 ID=Potri.001G112000.1.v4.1 annot-version=v4.1
MGFESSNGTASPKLLKFLIYGRTGWIGGLLGKLCQSQGIDFTYGSGRLENRPSLEADLVAVNPTHVFNAAGVTGRPNVDWCESHKVETIRTNVVGTLTLA
DLCREKGLVLINYATGCIFEYDSSHPLGSGIGFKEEDTPNFIGSFYSKTKAMVEDLLRNYENVCTLRVRMPISSDLANPRNFITKITRYEKVVDIPNSMT
ILDELLPISIEMAKRNLTGIYNFTNPGVVSHNEILEMYRDYIDPDFTWKNFTLEEQAKVIVAPRSNNELDTAKLKQEFPELLPIKESLIKYVFKPNQKTA
AA
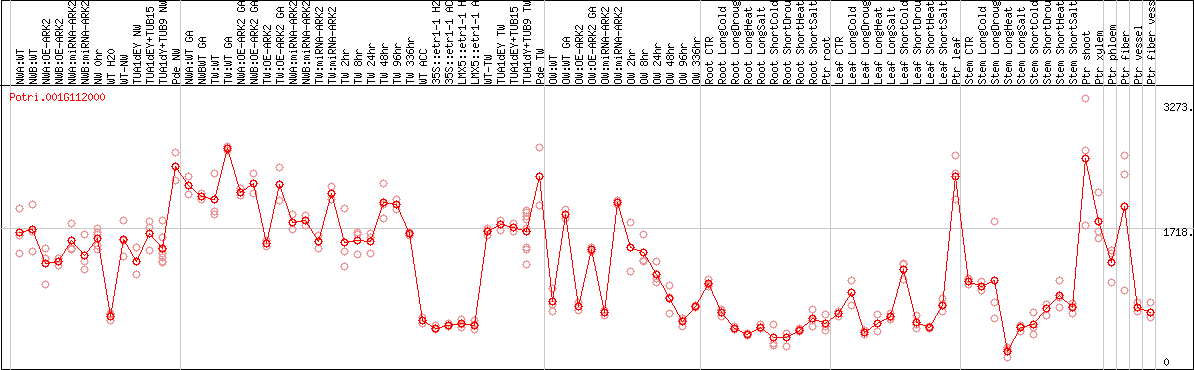
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.001G112000 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.