Potri.001G121200 [POPLAR]

| External link |
|
||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.001G121200.2 pacid=42792983 polypeptide=Potri.001G121200.2.p locus=Potri.001G121200 ID=Potri.001G121200.2.v4.1 annot-version=v4.1
ATGTGGTGTGAGATGATGGACTTGTTCGATCCTGGCAAGAAGATGTTGATGCAAGGAGATTTCGATCTGAACTCTGTACAGCGACACGCAGATTCATTTA
AAGGAGTTATAAAGCAGACAATTCTCAAGCAGGAGGTTATATTTAGGACTCAGGTCCATGAACTCCATCAGTTATATAGGACGCAAAAAACACTGATGAA
GGATCTTGGTAGTAAGTGTTGTGGTGGATACAATTCATGGGATGCAAATGTACAACCATCTCTACCACCTTTTCCGAACCCTTCAAGAATTGAACCATTG
GCGAAAGAAACAGGAATTTCCTCAATTTCCAAGGTGGGTCCAGCACCTTTCGCAAGCAAGGAGTTGTTACATGGGTGCCAAGATACATATTATAGACTTA
AGCAGATGCCTCTTGATCTTCAACTCTCTGCTAACAAACTCATCAACCATGTCGAGGAAGATCGCCCAAACAGAGGACATGCCTGGAATCATCCCTTAAA
AGAACCAATAGACGTTAAACATCCTCTCTCTGCTAATTACTCTTCAGGTACAGAGGAGTTGAAGCTTTCCCTGACTACTAGAGATGACTATAGAAGAACA
GAAGGTACATTGCGGACTTGGTTTGATAAGAGAACTCATCAGTACTGTTCCGTGATTGATCTGGAATCAGAAGAGAGGATATCAGATGACAGTGCAAAAT
GCACACCTTCTGTTGGCTGTGCTACTCCAGAAACTTATTCCCCAGGGAAGTACAAAAAAGTTTCTGCCTTCTCTAGTTTAATTTTCTTAACTAGTGCAGA
GAAAGATCCATCTGTCGAGATTTCAGAAAGCAGCTATTTTCAAGAACATAGTGAATGCTGCCAAGAGCAGACCTCTTCTAATGAAGGAATTATGGAGTGC
CATGATGATAACCTGTGTAATAATCTTTCCACCAAAATGAAACAGTCCTCTTCACATAAAGGAACTGATTTGGACCTCAACAAAGCTCACTTTGATGACC
CGTCATGTTTCTCAAATGATCCTCTAGTGGCCTGTCCTTCACCAGCTAGATCAGCTGGTGACTCTGCGGCAGTTATTAGGAGCATGCAGGAAGAAACCTG
TCCAACCACATTTTGGGACAAACAGGTCAATAGCTGCTCAAATGAAATTTCTGACATGCTTGCAGAAAATAATTTCATAAATGCTACTGAAGTAGATTTA
AACAGCACAACTAGGAGTACAGATGTTTGGACCGGAAATTCTGAGCAGGATGGAATAAGTGGTACTTTACCAAACCCTACAGGCCCGGAGCCCATACCCA
GCCCCCCTGTAGACATTTCTGAAGACATCGACAGGTACACTGGCGACCACAAGAATGATAATGTTGTGATGAAGGCGAAACTCGCGAATTGTCTTCTACA
TGATTTAAATCAAATGCCTTTAGATACTAGAGAACTGAGCTCTGAGAAGAGCCAAGTGGAAGATGCTGTCTTTTCATGTATTGATCAGTATCAGAATGAT
GGACATGGCAATCAATCTCCTGTTTCATGCAAGTCTGGTATTTATGATAATGATTCAAACAGTGGAAAGACAGTGCACTTTGGCAGTATGTCTGGTGATG
TGAACACTGATTTGAAAACCCTCTCAGGGTCACAAGTTGCCGGTGCCTCATCAGATGAGCATGACCTAAGAACTTCTGACAGTGGTGATTTGAAAAATGA
ATGCTACGACAAGAAAGAAGAATCAGCTGAAGTTGATGTTTTGATGAAAAGAGCAGCTGAATCACTTGTAGATATGTCTTTGAAGAATTTAATTTGCTTT
CAAGATTCTTTTGCCAAAGAAGGGTCCAAGGAGATGAGAAAAGAGACGAGGGAGCAGCCACAGTATACTTGCGATTCTTTTGAATTAATGGTTATGGAAC
TAACGGAAAGCAATGTGGATGAAAATTCAGTGACATCAAAGCCATATGAAGTAAATGATATGGAAACAAAAGATTTTGGTTCTAAACTGAGACGAGGAAG
GAGAATGAAAGATTTCCAGAAGGAGATACTGCCTGCTCTGGCATCTCTTTCTAGACACGAAATTCATGAGGATATAAACATTATCGAGGGAGTTTTAAGA
TCAAGAGAATACCGCAAAATCAGAGGCAAGATGGCAACAAATGGAGAGAACTGGTCTCCACTGGTGAGAAGTAGACGGTCAAGACTCAATAATGTTGGGA
GGAGAAGAAGTTCATCGAAGTTCAACTAG
|
||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.001G121200.2 pacid=42792983 polypeptide=Potri.001G121200.2.p locus=Potri.001G121200 ID=Potri.001G121200.2.v4.1 annot-version=v4.1
MWCEMMDLFDPGKKMLMQGDFDLNSVQRHADSFKGVIKQTILKQEVIFRTQVHELHQLYRTQKTLMKDLGSKCCGGYNSWDANVQPSLPPFPNPSRIEPL
AKETGISSISKVGPAPFASKELLHGCQDTYYRLKQMPLDLQLSANKLINHVEEDRPNRGHAWNHPLKEPIDVKHPLSANYSSGTEELKLSLTTRDDYRRT
EGTLRTWFDKRTHQYCSVIDLESEERISDDSAKCTPSVGCATPETYSPGKYKKVSAFSSLIFLTSAEKDPSVEISESSYFQEHSECCQEQTSSNEGIMEC
HDDNLCNNLSTKMKQSSSHKGTDLDLNKAHFDDPSCFSNDPLVACPSPARSAGDSAAVIRSMQEETCPTTFWDKQVNSCSNEISDMLAENNFINATEVDL
NSTTRSTDVWTGNSEQDGISGTLPNPTGPEPIPSPPVDISEDIDRYTGDHKNDNVVMKAKLANCLLHDLNQMPLDTRELSSEKSQVEDAVFSCIDQYQND
GHGNQSPVSCKSGIYDNDSNSGKTVHFGSMSGDVNTDLKTLSGSQVAGASSDEHDLRTSDSGDLKNECYDKKEESAEVDVLMKRAAESLVDMSLKNLICF
QDSFAKEGSKEMRKETREQPQYTCDSFELMVMELTESNVDENSVTSKPYEVNDMETKDFGSKLRRGRRMKDFQKEILPALASLSRHEIHEDINIIEGVLR
SREYRKIRGKMATNGENWSPLVRSRRSRLNNVGRRRSSSKFN
|
DESeq2's median of ratios [POPLAR]
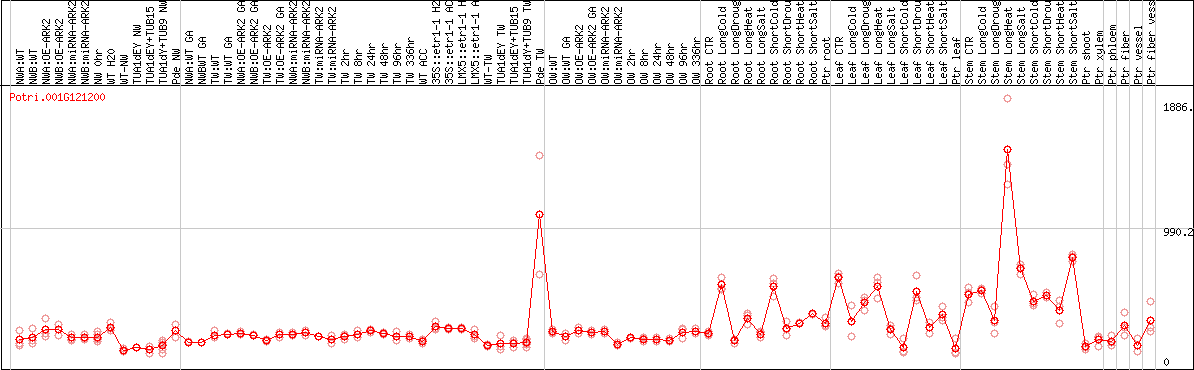
Coexpressed genes
Potri.001G121200 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
